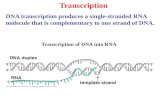
Transcription DNA
Transcript of Transcription DNA
-
8/13/2019 Transcription DNA
1/29
Lecture materials may be downloaded from:
http://www.msi.upd.edu.ph/claroline/
Username: MS272
Password: student
The transcription of genes
-
8/13/2019 Transcription DNA
2/29
DNA
The central dogma of molecular biology
transcription translation
replication
reverse-transcription
replication
RNA protein
DNA
Types of RNA
transcription
translation
mRNA protein
tRNA
rRNA
other types
-
8/13/2019 Transcription DNA
3/29
The transcription of genes:prokaryotes
Key proteins involved in thetranscription of bacterial genes
RNA polymerase, core enzyme
two subunits, one subunit,
one subunit, [one subunit]
sigma factor,
RNA polymerase + sigma factor holoenzyme
synthesizes all 3 classes of RNA
(mRNA, tRNA, rRNA)
-
8/13/2019 Transcription DNA
4/29
Structure ofeubacterial
RNA polymerases
Transcription unit
A sequence of DNA transcribed into a single RNA, startingat the promoter and ending at the terminator
-
8/13/2019 Transcription DNA
5/29
Promoter - region of DNA involved in binding
of RNA polymerase to initiate
transription
Functions of a promoter
The promoter determines:
Which strand will serve as a template.
Transcription starting point.
Strength of polymerase binding.
Frequency of polymerase binding.
-
8/13/2019 Transcription DNA
6/29
Four stages of transcription
1
2
3
4
template recognition
initiation
elongation
termination
Template recognition
forming the closedpromoter complex
-
8/13/2019 Transcription DNA
7/29
Template recognition
RNA polymerase recognizes promoters
based on consensus sequences
consensus sequence an idealizedsequence in which each positionrepresents the base most often foundwhen many actual sequences are
compared
- 10 box- 35 box
-
8/13/2019 Transcription DNA
8/29
Transcription bubble
-
8/13/2019 Transcription DNA
9/29
Contact points between the sigma factorand a typical prokaryotic promoter
HOOC NH2
-
8/13/2019 Transcription DNA
10/29
Initiation of transcription
formation of open complex
initial synthesis
loss of sigma factor
promoter clearance
RNA polymerase moves like an inchwormduring elongation
Back end moves 1bp per nucleotideadded to RNA
Front end movesseveral bpdiscontinuously
-
8/13/2019 Transcription DNA
11/29
RNA synthesis
Termination of transcription
How does the RNA polymeraseknow when to stop?
Types (based on in vitro experiments):
intrinsic terminators - no proteins (factors) required
rho-dependent - rho factor required
TERMINATORS - sequences at which
transcription stops and thecomplex dissociates
-
8/13/2019 Transcription DNA
12/29
Intrinsic terminators: structural features
hairpin-forming (palindromic, inverse-repeat)region
a run of U residues
Intrinsic terminators: structural features
-
8/13/2019 Transcription DNA
13/29
Principal feature: C-rich, G-poor sequence
preceding actual termination site
rho-dependent terminators
Hot-pursuit modelfor rho-dependent
termination
-
8/13/2019 Transcription DNA
14/29
Coupling of transcription and translationin prokaryotes
The transcription of genes:
eukaryotes
-
8/13/2019 Transcription DNA
15/29
Transcription
Pre-mRNA processing
Transport to the cytoplasm
The generation of mature mRNAsinvolves the following major steps
1. transcription
the eukaryotic RNA polymerases
the transcription factors
promoters and enhancers
stages
chromatin remodeling
initiation of transcription
elongation termination
2. pre-mRNA processing
capping
splicing
polyadenylation
3. transport to the cytoplasm
-
8/13/2019 Transcription DNA
16/29
Type of Polymerase Product Location
RNA Polymerase I pre-rRNA (except 5S) nucleolus
RNA Polymerase II pre-mRNA (hnRNA) nucleoplasm
RNA Polymerase III tRNA, 5S rRNA, etc. nucleoplasm
Types of eukaryotic RNA polymerases
-
8/13/2019 Transcription DNA
17/29
Transcription factor - any protein other than RNA Polymerase
that is required for transcription
may bind to RNA Polymerase
may bind to another transcription factor
may bind to cis-acting DNA sequences may bind to effectors
RNA pol II cannot initiate transcription by itself;
requires auxilliary transcription factors to
initiate transcription.
General factors - required for initiating transcription at all
promoters
Upstream factors - ubiquitous factors that increase the
efficiency of transcription initiation; set of
factors unique to each promoter
Inducible factors - act in the same manner as an upstream
factor but their synthesis is regulated in a
temporal or spatial manner
Transcription factorsTypes of transcription factors:(according to their importance to transcription initiation)
Basal transcription apparatus - RNA polymerase + general
factors both needed to initiate
transcription
-
8/13/2019 Transcription DNA
18/29
-
8/13/2019 Transcription DNA
19/29
Major types of DNA-binding domains
(motifs found in the sequences)
zinc finger
helix-turn-helix
amphipathic helix-loop-helix
leucine zippers
-
8/13/2019 Transcription DNA
20/29
helix-turn-helix
zinc finger
leucine zipper
Promoters vs. enhancers
-
8/13/2019 Transcription DNA
21/29
-
8/13/2019 Transcription DNA
22/29
-
8/13/2019 Transcription DNA
23/29
-
8/13/2019 Transcription DNA
24/29
Initiation of transcription
ordered sequence of assembly of
initiation complex
phosphorylation of the C-terminal
domain (CTD) tail of RNA polymerase II
(may be needed for promoter clearance)
Assembly ofinitiationcomplex
-
8/13/2019 Transcription DNA
25/29
-
8/13/2019 Transcription DNA
26/29
Elongation of transcript
phosphorylation ofthe CTD tail ofRNA polymerase II
CPSF
CPSF = cleavage and polyadenylation specificity factor
Termination and polyadenylation
-
8/13/2019 Transcription DNA
27/29
Pre-mRNA processing: capping
Caps may differ in number of methylations
Pre-mRNA
processing:
splicing
Spliceosomes
consist of
snRNPs
-
8/13/2019 Transcription DNA
28/29
Conserved signals at intron-exon junctions
Translation of mRNA in the cytoplasm
-
8/13/2019 Transcription DNA
29/29
Next meeting
Translation of mRNAs