Molecular dynamics simulation of Aquaporin-1 · Tertiary structure = 3D fold of one polypeptide...
Transcript of Molecular dynamics simulation of Aquaporin-1 · Tertiary structure = 3D fold of one polypeptide...
-
4 nm
Molecular dynamics simulation of Aquaporin-1
-
i~@t (r, R) = H (r, R)
He e(r;R) = Ee(R) e(r;R)
Molecular Dynamics Simulations
Schrödinger equation
Born-Oppenheimer approximation
Nucleic motion described classically
Empirical Force field
1
-
Molecular Dynamics Simulations
Interatomic interactions
-
„Force-Field“
-
Molecular Dynamics SimulationMolecule: (classical) N-particle system
Newtonian equations of motion:
with
Integrate numerically via the „leapfrog“ scheme:
(equivalent to the Verlet algorithm)
with
Δt ≈ 1fs!
-
“Aquaporin” water channel
-
Human hemoglobin
-
Lipid membranes
-
Today’s lecture
• Protein structures • Notes on force calculations • Setup of a simulation • Organize force field parameters • Algorithms used during simulation • Energy minimization and equilibration of
initial structure
• Analysis of a simulation
-
Protein structures: primary structure
• 20 different amino acids encoded in the DNA
• 3-letter and 1-letter codes
www2.chemistry.msu.edu
Primary structure = amino acid sequence
KVFGRCELAAAMKRHGLDNYRGYSLGNWVCAAKFESNFNTQATNRNTDGSTDYGILQINSRWWCNDGRTPGSRNLCNIPCSALLSSDITASVNCAKKIVSDGNGMNAWVAWRNRCKGTDVQAWIRGCRL
Lysozyme
• From N- to C-terminus
-
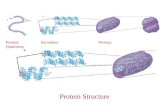
Protein structures: secondary structure
Secondary structure = 3D fold of local AA segments
Lysozyme:
alpha-helices, beta sheets, connected by loops
• alpha helix
• beta sheet
• Turns, 310-helix,…
-
Protein structures: tertiary structure
Tertiary structure = 3D fold of one polypeptide chain
Mainly alpha-helical
-
Protein structures: tertiary structure
Tertiary structure = 3D fold of one polypeptide chain
Mainly beta sheets
-
Protein structures: tertiary structure
Tertiary structure = 3D fold of one polypeptide chain
OmpX (pdb 2M06)
-
Protein structures: ter-ary structure
Alpha helices and beta sheets
-
Protein structures: ter-ary structure
Alpha helices and beta sheets
-
Protein structures: quaternary structure
Arrangement of multiple folded polypeptides
Example: Haemoglobin• four subunits
Interesting: Cooperative oxygen binding
through quaternary transitions
-
Multiple Time Stepping
H. Grubmüller, H. Heller, A. Windemuth, K. Schulten; Mol. Sim. 6 (1991) 121
-
1. Taylor expansion
Multipole Methods
Exact for infinite multipole series
O(N2)
i
i j
j
-
Fast Multipole Method (FMM)
+ arbitrary accuracy
- high order expansions required to achieve moderate accuracy
à O(N)
L. Greengard and V. Rokhlin, J. Comp. Phys. 73 (1987) 325
-
Fast structure-adapted multipole methods: O(N)
M. Eichinger, H. Grubmüller, H. Heller, P. Tavan, J. Comp. Chem. 18 (1997) 1729
-
Ewald summationAnother very popular method to efficiently compute Coulomb forces of without simple cutoffs
(applicable for periodic systems)
q
x x x
q q
Idea: Rewrite the charge density as a sum of two terms:
• Quickly varying density: Potential can be computed accurately with cut-offs (“direct space calculation”)
• Slowly varying density: potential can be efficiently computed in reciprocal space using the Fast Fourier Transform (FFT); O(N log(N))
Charge density:
Point charges
Fourier transform of charge densityEwald, Ann. Phys. 64:253-287 (1921)
-
Simulation system setup 1
• Get PDB structure and check for ‣ missing atoms/groups ‣ inaccuracies (flipped histidine ring) ‣ missing ligands ‣ chemical plausibility ‣ mutations (e.g., to facilitate crystallization) ‣ read the paper!!
• Choose force field ‣ “all-atom” or “united-atom”, e.g. CH2, CH3 as one atom ‣ implicit or explicit hydrogen atoms ‣ polarizable force field required? ‣ QM methods required (chemistry?)
• Add hydrogen atoms to protonable (“titratable”) groups (Histidine!)
-
Simulation system setup 2
• Choose periodic boundary conditions or not
-
Role of environment - solvent
explicit or
implicit solvation?
box or droplet?
Typical: box with periodic boundary conditions,
avoid surface artefacts
-
periodic boundary conditions and the minimum image convention
Surface (tension) effects?
-
~xi(t = 0) done!
Simulation system setup 2
• Choose periodic boundary conditions or not • if membrane protein: add lipid membrane atoms • add water molecules • add ions as counter ions (if possible, according to Debye-
Hückel)
-
b(i)0 ,K(i)b for all bonds
�(j)0 ,K(j)� for all angles
Simulation system setup 3
• Define V(x1,...xN) via force field
‣ bond parameters
‣ angle parameters
‣ dihedrals, extraplanars
‣ partial charges
‣ Van-der-Waals parameters (Lennard-Jones potential)
VLJ = 4✏
⇣�r
⌘12�⇣�r
⌘6�
qi for all atoms
�i, ✏i for all atoms
-
Simulation system setup 4• For frequently reoccurring chemical motifs
define atom types, e.g.: ‣ hydrogen HC ‣ carbon CH2
• parameter file: list properties of atom types and their bonds, angles, ...
HC q=+0.2 m=1.0 # charge, massCH2 q=-0.4 m=12.0
HC -CH2 K=200 b=1.1 # bondsCH2-CH2 K=500 b=1.5
HC-CH2-HC K=20 118° # anglesHC-CH2-CH2 ...
-
Simulation system setup 5
‣ Topology file: defines • atoms • bonds • angles • dihedrals etc. of the simulation system
[ atoms ]; nr type name … 1 HC HA1 2 HC HA2 3 HC HB1 4 HC HB2 5 CH2 CA 6 CH2 CB
[ bonds ] 1 5 HC-CH2 2 5 HC-CH2 3 6 HC-CH2 4 6 HC-CH2 5 6 CH2-CH2
[ angles ] 1 5 2 HC-CH2-HC 1 5 6 HC-CH2-CH2...
1
25
3
46
-
Simulation phase - algorithms
‣ Integration of Newton’s equations of motion
Integrate numerically via the „leapfrog“ scheme:
(equivalent to the Verlet algorithm)
with
Δt ≈ 1fs!
where
-
~P =N
atomsX
i=0
~pi
~pi0 = ~pi �
miM
~P
Simulation phase - algorithms
‣ Integration of Newton’s equations of motion ‣ Constrain bond lengths (LINCS, SHAKE)
idea: eliminate fastest vibrations (C-H) to increase the integration time step from 1fs to 2fs side-effect: better descriptions of QM vibrations
‣ Remove overall translation (and rotation): Avoid drift of the molecule: remove translation (and rotation) of the entire simulation system:
Remove overall momentum:
Remove angular momentum analogously
-
Simulation phase - algorithms
‣ Remove overall translation (and rotation): Avoid drift of the molecule: remove translation (and rotation) of the entire simulation system:
0 1000 2000 3000 4000 5000
Time (ps)
0
500
1000
1500
2000
Coord
inate
(nm
)
Center of mass
0 1000 2000 3000 4000 5000
Time (ps)
-10000
-8000
Pote
ntia
l (kJ
/mol)
Numerical instability: Accumulation of kinetic energy in to one degree of freedom. (Flying ice cube problem)
-
~vi ~vi
s
1� �t⌧
✓T
T0� 1
◆
T =2
3
1
NkB
NX
i=1
m
2v2i
Simulation phase - algorithms
‣ Choose thermodynamic ensemble NVE (microcanonical ensemble) NVT (canonical ensemble, isochoric): T-coupling NPT (canonical ensemble, isobaric): T-coupling and P-coupling
‣ T-coupling, e.g. Berendsen thermostat After each step Δt:
‣ P-coupling: analogous, by scaling volume ‣Write out coordinates at some frequency
𝝉 = coupling time constant
T0 = target temperature
-
Mimimization/equilibration: 1) Energy minimization
☞ Reduce the steric strain by a moving along the steepest descent in
V (~x1, . . . , ~xN )
☞ Notes:
• Protein moves in to local minimum
• Attention: proteins don’t tend towards the local minimum in V(x), but towards the global minimum in the free energy! ☞ Entropy/ensembles are important!
-
BPTI: Minimization
-
Mimimization/equilibration: 2) Thermalization
☞ Heat the system to, e.g. 300K by assigning Maxwell-distributed velocities
p(vx
) / e�mv
2x
2kB
T , p(vy
) / · · ·
Trick to avoid distortion of the protein: • assign velocities to to the system• keep protein backbone restrained• equilibrate for ~100ps
-
Mimimization/equilibration: 3) Equilibration
How long? → Multiple checks:
• Convergence of energy contributions (particularly Coulomb and Lennard-Jones) and box dimensions
• Room-mean square deviation (RMSD) from the crystal/NMR structure
RMSD(t) =
✓1
N
XNi=1
[~xi(t)� ~xi(0)]2◆1/2
Typically:
0 1 2 3 4 5 6 7 8 9
Time (ns)
0.00
0.05
0.10
0.15
RM
SD
(n
m)
picosecond jumpconformationalsampling
?
-
Mimimization/equilibration: 3) Equilibration
Reasons for RMSD increase/drift:
• Fast fluctuations → picosecond jump ☞ OK• slow conformational motions
→ nanosecond drift ☞ OK
• Conformational transitions → stairs ☞ OK
• Structural drift due to ☞ NOT OK - bad X-ray structure- inaccurate force field- software bug- …
-
Mimimization/equilibration: 3) Equilibration
Judgement of RMSD:
• RMSD does not converge ⟹ simulation is not OK.• But: RMSD converges ⇏ simulation is OK.
Better check, e.g., PCA projections (see later lecture)
-
Flow chart of MD simulation
Get initial positions of atoms (e.g., from the PDB)
Compute forces using your force field
Update atom positions & velocities (“integration step”)
Update time step t ! t+�t
Repeat up to requested simulation time
Take care of pressure and temperature
e.g.
109
times
Prepare simulation system (add hydrogen atoms, water, ions)
Choose force field
Specify simulation parameters (time step, temperature, …)
Preparation Simulation
Energy minimisation
Set initial velocities
-
Simulation analysis
Available after simulation:
• Positions:
e.g., T = 10ns, N = 100.000, Δt = 2fs
☞ 5·106 × 105 × 3 × 4 Byte = 6 TByte !
• Velocities
• Temperature
• Potential energies:
• Anything you can program…
~x1(ti), . . . , ~xN (ti), ti = 0,�t, 2�t, . . . , T
~v1(ti), . . . ,~vN (ti)
T (ti) =1
(3N � 6)kB
NX
i=1
miv2i (ti)
Vbond
(ti), Vangle(ti), Vdih(ti), VCoul(ti), VLJ(ti),
-
Simulation analysis
Observables that may be interesting: everything that can be measured
• Size of atomic fluctuations
Note: ensemble average ⟨⋯⟩ ≠ time average
• Anything that helps to understand the protein function:
- Movie (!), motion of groups
- interaction energies, hydrogen bonds, radial distribution
functions, transition rates, change in secondary structure
x̄j = M�1
MX
i=1
~xj(ti)
h(~xj � h~xij)2i ⇡1
M
MX
i=1
⇥~xj(ti)� x̄j
⇤2
-
BPTI: Molecular Dynamics (300K)
-
Opening transition of the enzyme ATCase
-
4 nm
Molecular dynamics simulation of Aquaporin-1




![Fundamentals of Protein Structure · PDF fileProtein Structure Thomas Funkhouser ... Tertiary Structure How protein folds: 1atp [pymol] Tertiary Structure ... Root: scop 2. Class:](https://static.fdocuments.us/doc/165x107/5abd21b77f8b9a5d718b532c/fundamentals-of-protein-structure-structure-thomas-funkhouser-tertiary-structure.jpg)