Lecture 10 Protein Tertiary (3D)...
Transcript of Lecture 10 Protein Tertiary (3D)...

Introduction to Bioinformaticsfor Medical Research
Gideon [email protected]
Lecture 10Protein Tertiary (3D) Structure

2
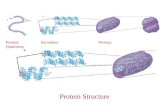
Protein Tertiary Structure
• Defining Structure• Determining experimentally
– PDB• Predicting Structure
– TOPITS– GenTHREADER
• Structural classification– SCOP

3
Defining Structure
1 N MET A 1 -14.830 -2.121 10.034 2 CA MET A 1 -14.608 -1.535 8.679 3 C MET A 1 -15.821 -1.799 7.781 4 O MET A 1 -15.713 -2.464 6.770 5 CB MET A 1 -13.372 -2.254 8.135 6 CG MET A 1 -13.531 -3.764 8.330 7 SD MET A 1 -12.739 -4.636 6.956 8 CE MET A 1 -13.839 -6.072 6.937 9 1H MET A 1 -15.554 -2.865 9.976 10 2H MET A 1 -13.942 -2.531 10.386
Hydrogen number
Residue numberRemotenessAtomic symbol
3D co-ordsChainResidue

4
X-ray Crystallography
• Create repetitive crystal of molecule– Often difficult, especially hydrophobic portions
• X-rays generate diffraction pattern– Pattern represents electron density
• Generate comparison patterns– Add ions or change wavelength
• Obtain electron density map– Fit protein sequence to map

5
Nuclear Magnetic Resonance
• Dissolve molecules in water– Allows free tumbling and vibration
• Detect activity of atoms with quantum spin– 1Hydrogen (natural), 13Carbon, 15Nitrogen
• Defines set of atomicdistance constraints– Ensemble of models
• Can detect motion

6
PDB
• Database of molecular structures– Obtained by crystallography or NMR– Carefully curated and validated
• Founded in 1971– 19375 proteins, 2117 other structures
• Additional protein information– Secondary structure– References, external links

7
PDB: Summary Information
Chains in molecule
Experimentalmethod
Molecule in PDB entry
Link to SCOP

8
PDB: 3D Structure
• Still images at fixed orientation– Generate at any size
• Interactive molecule explorer– Requires Java or Chime plug-in
• Download structure file– Display in RasMol,
Swiss-PDBViewer, etc…• Demonstration

9
Predicting 3D Structure
• Outstanding difficult problem• Based only on protein sequence
– Comparative modeling (homology)– Ab-initio modeling
• Based on secondary structure– Fold recognition– Protein threading

10
Comparative Modeling
• Similar sequence suggests similar structure– Amino acid characteristics determine folding
• Similarity particularly high in core– Alpha helices and beta sheets preserved– But even near-identical sequences vary in loops
• Effectiveness depends on protein length– Longer fi less sequence similarity required

11
Ab Initio Modeling
• Compute molecular structure from laws ofphysics and chemistry alone– Ideal solution (theoretically)
• Simulate process of protein folding– Apply minimum energy considerations
• Practically nearly impossible– Exceptionally complex calculations– Biophysics understanding incomplete

12
Protein Folds
• A combination of secondary structural units– Forms basic level of classification
• Each protein family belongs to a fold– Estimated 700–1500 different folds

13
Fold Recognition / Threading
• Compare sequence against known structures– Try to ‘thread’ sequence along chain
• Score suitability of the threading– Can adjacent amino acids bond?– Are amino acids close to or far from water?– Are secondary structures similar?
• Examine list of most threadable structures– Correct answer is often in top 10 or so

14
Threading Example
Knownstructure
Querysequence
Gaps inthreading

15
TOPITS Output (1)
Alignmentscore
Alignmentlength
Lengthof indels
Numberof indels
Length ofsequence
Alignmentsignificance
Matchedsequence
% sequenceidentity

16
TOPITS Output (2)
Querysequence
Predictedstructure
Buried /Outside
Databasesequence
Amino acidmatches
Database knownsecondary structure

17
GenTHREADER Output
Predictionconfidence
Expectederrors
Score fromneural network
Sequence alignmentscore and length
Energymeasurements
Length ofsequence
Structurefrom PDB

18
Prediction Flowchart
PSI-BLAST
Ab initiomethods
TOPITS,GenTHREADER
PHDsec,PSIpred

19
Structure Classification
• Class– All alpha, all beta, alpha/beta, alpha+beta
• Fold– Strong structural similarity
• Superfamily– Probably common evolutionary origin
• Family– Evolutionary relationship, sequence similarity

20
SCOP
• Structural Classification of Proteins– Based on known protein structures– Manually created by visual inspection
• Hierarchical database structure– Class, fold, superfamily, family– Proteins/domains, species instances
• Founded in 1995– 765 folds, 1232 superfamilies, 2164 families

21
SCOP: Navigation
Nodename
Nodedescription
Path fromroot to node
Childrenof node

22
Other Resources
• CATH (classification of protein domains)– http://www.biochem.ucl.ac.uk/bsm/cath/
• SWISS-MODEL (comparative modeling)– http://www.expasy.ch/swissmod/
• CASP (structure prediction competition)– http://predictioncenter.llnl.gov/
• GTSP (guide to structure prediction)– http://speedy.embl-heidelberg.de/gtsp/









![Protein structure determination & prediction. Tertiary protein structure: protein folding Three main approaches: [1] experimental determination (X-ray.](https://static.fdocuments.us/doc/165x107/56649d605503460f94a41068/protein-structure-determination-prediction-tertiary-protein-structure-protein.jpg)