Fig. 11-0a. Fig. 11-0b Fig. 11-0c Fig. 11-1a Fig. 11-1b DNA RNA polymerase cannot attach to promoter...
-
Upload
simon-montgomery -
Category
Documents
-
view
212 -
download
0
Transcript of Fig. 11-0a. Fig. 11-0b Fig. 11-0c Fig. 11-1a Fig. 11-1b DNA RNA polymerase cannot attach to promoter...

Fig. 11-0a

Fig. 11-0b

Fig. 11-0c

Fig. 11-1a

Fig. 11-1b
DNA
DNA
RNA polymerasecannot attach to promoter
Lactose-utilization genesPromoter OperatorRegulatorygene
OPERON
Protein
mRNA
Inactiverepressor
Lactose Enzymes for lactose utilization
RNA polymerasebound to promoter
Operon turned on (lactose inactivates repressor)
mRNA
Activerepressor
Operon turned off (lactose absent)
Protein

Fig. 11-1ba
DNA
RNA polymerasecannot attach to promoter
Lactose-utilization genesPromoter OperatorRegulatorygene
OPERON
mRNA
Activerepressor
Operon turned off (lactose absent)
Protein

Fig. 11-1bb
DNA
Protein
Inactiverepressor
Lactose Enzymes for lactose utilization
RNA polymerasebound to promoter
Operon turned on (lactose inactivates repressor)
mRNA

Fig. 11-1c
DNA
Inactiverepressor
Activerepressor
Inactiverepressor
Activerepressor
Lactose
Promoter
Tryptophan
Operator Gene
lac operon trp operon

Fig. 11-2
Muscle cell Pancreas cells Blood cells

Fig. 11-2a
Muscle cell

Fig. 11-2b
Pancreas cells

Fig. 11-2c
Blood cells

Fig. 11-3
DNA double helix(2-nm diameter)
“Beads ona string”
Linker
Histones
Metaphasechromosome
Tight helical fiber(30-nm diameter)
Nucleosome(10-nm diameter)
Supercoil(300-nm diameter)
700 nm

Fig. 11-4
Two cell populationsin adult
X chromosomes
Early embryo
Allele forblack fur
Inactive X
Black furAllele fororange fur
Orange fur
Cell divisionand random
X chromosomeinactivation Active X
Inactive X
Active X

Fig. 11-5Enhancers
Otherproteins
DNA
Transcriptionfactors
Activatorproteins
RNA polymerase
Promoter
Gene
Bendingof DNA
Transcription

Fig. 11-6
1
or
Exons
DNA
RNA splicing
RNAtranscript
mRNA
2 3 4 5
1 2 3 4 5
1 2 4 51 2 3 5

Fig. 11-7
miRNA 1
Translation blockedORmRNA degraded
Target mRNA
Protein
miRNA-proteincomplex
2
3 4

Fig. 11-8
Folding ofpolypeptide andformation ofS—S linkages
Initial polypeptide(inactive)
Folded polypeptide(inactive)
Active formof insulin
Cleavage

Fig. 11-9NUCLEUS
DNA unpackingOther changes to DNA
Addition of cap and tail
Chromosome
Gene
RNA transcript
GeneTranscription
Intron
Exon
Splicing
CapmRNA in nucleus
Tail
Flow throughnuclear envelope
Broken-downmRNA
CYTOPLASM
Breakdown of mRNA
Translation
mRNA in cytoplasm
Broken-downprotein
Cleavage / modification /activation
Breakdown of protein
Polypeptide
Active protein

Fig. 11-9a
NUCLEUS
DNA unpackingOther changes to DNA
Addition of cap and tail
Chromosome
Gene
RNA transcript
GeneTranscription
Intron
Exon
Splicing
CapmRNA in nucleus
Tail
Flow throughnuclear envelope

Fig. 11-9b
Broken-downmRNA
CYTOPLASM
Breakdown of mRNA
Translation
mRNA in cytoplasm
Broken-downprotein
Cleavage / modification /activation
Breakdown of protein
Polypeptide
Active protein

Fig. 11-10a
Head of a normal fruit fly
Antenna
Eye
Head of a developmental mutant
Leg

Fig. 11-10aa
Head of a normal fruit fly
Antenna
Eye

Fig. 11-10ab
Head of a developmental mutant
Leg

Fig. 11-10bEgg cellwithin ovarianfollicle
Follicle cells
“Head”mRNA
Proteinsignal
Egg cell
Gene expression1
Cascades ofgene expression2
Embryo Bodysegments
Adult fly
Gene expression
3
4

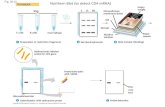
Fig. 11-11
cDNA
Nonfluorescent spotFluorescent
spot
Actual size(6,400 genes)
Each well contains DNAfrom a particular gene
DNA microarray
mRNAisolated
DNA of anexpressed gene
DNA of anunexpressed gene
Reverse transcriptaseand fluorescent DNAnucleotides
1
cDNA madefrom mRNA
2
cDNA appliedto wells
3
UnboundcDNA rinsedaway
4

Fig. 11-12Signaling cell
DNA
Nucleus
Transcriptionfactor(activated)
Signaling molecule Plasma
membraneReceptorprotein
Relayproteins
TranscriptionmRNA
Newprotein
Translation
Target cell
2
1
3
4
5
6

Fig. 11-13
Yeast cell,mating type a factor
factor
Receptor
a
Yeast cell,mating type a
a
a/

Fig. 11-14
Root ofcarrot plant
Root cells culturedin nutrient medium
Cell divisionin culture
Single cell
Plantlet Adult plant

Fig. 11-15
Removenucleusfrom eggcell
Implant blastocyst insurrogate mother
Add somatic cellfrom adult donor
Donorcell
Remove embryonicstem cells fromblastocyst andgrow in culture
Reproductivecloning
Nucleus fromdonor cell
Grow in cultureto produce anearly embryo(blastocyst)
Therapeuticcloning
Clone ofdonor is born
Induce stemcells to formspecialized cells

Fig. 11-15a
Removenucleusfrom eggcell
Add somatic cellfrom adult donor
Donorcell Nucleus from
donor cell
Grow in cultureto produce anearly embryo(blastocyst)

Fig. 11-15b
Implant blastocyst insurrogate mother
Remove embryonicstem cells fromblastocyst andgrow in culture
Reproductivecloning
Therapeuticcloning
Clone ofdonor is born
Induce stemcells to formspecialized cells

Fig. 11-16

Fig. 11-17
Blood cells
Different types ofdifferentiated cells
Nerve cells
Adult stemcells in bone
marrow
Different cultureconditions
Culturedembryonicstem cells
Heart muscle cells

Fig. 11-18a
Mutation withinthe gene
Hyperactivegrowth-stimulatingprotein in normalamount
Proto-oncogene DNA
Multiple copiesof the gene
Gene moved tonew DNA locus,
under new controls
Oncogene New promoter
Normal growth-stimulatingprotein in excess
Normal growth-stimulatingprotein in excess

Fig. 11-18b
Mutated tumor-suppressor geneTumor-suppressor gene
Defective,nonfunctioningprotein
Normalgrowth-inhibitingprotein
Cell divisionunder control
Cell division notunder control

Fig. 11-19a
1
Colon wall
Cellularchanges:
DNAchanges:
Oncogeneactivated
Increasedcell division
Tumor-suppressorgene inactivated
Growth of polyp
Second tumor-suppressor geneinactivated
Growth of malignanttumor (carcinoma)
2 3

Fig. 11-19b
Chromosomes 1mutation
Normalcell
4mutations
3mutations
2mutations
Malignantcell

Fig. 11-20aGrowth factor
Protein thatStimulatescell division
Translation
Nucleus
DNA
Target cell
Normal productof ras gene
Receptor
Relayproteins
Transcriptionfactor(activated)
Hyperactiverelay protein(product ofras oncogene)issues signalson its own
Transcription

Fig. 11-20b
Growth-inhibiting factor
Protein thatinhibitscell division
Translation
Normal productof p53 gene
Receptor
Relayproteins
Transcriptionfactor(activated)
Nonfunctional transcriptionfactor (product of faulty p53tumor-suppressor gene) cannot trigger transcription
Transcription
Protein absent(cell divisionnot inhibited)

Fig. 11-21

Fig. 11-UN1
Regulatorygene
Promoter
DNA
Gene 1Operator
A typical operon
Gene 2Gene 3
Encodes repressorthat in active formattaches to operator
RNApolymerasebinding site
Switches operonon or off

Fig. 11-UN2
Nucleus fromdonor cell
Early embryoresulting fromnuclear trans-plantation
Surrogatemother
Cloneof donor

Fig. 11-UN3
Nucleus fromdonor cell
Early embryoresulting fromnuclear trans-plantation
Embryonicstem cellsin culture
Specializedcells

Fig. 11-UN4Gene
regulation(a)
prokaryoticgenes oftengrouped into
in eukaryotesmay involve when
abnormalmay lead to
can produce
is a normal gene thatcan be mutated to an
oncogene
can cause
(c)
(g)(f)
multiple mRNAsper genetranscription
femalemammals
(e)
(d)
(b)
operons
areswitchedon/off by
in activeform binds to
controlled byprotein called
occurs inare proteinsthat promote