DNA Synthesis and Transcription
-
Upload
julesarojinee -
Category
Documents
-
view
497 -
download
1
description
Transcript of DNA Synthesis and Transcription
-
(318 305)
DNA synthesis & Gene Expression.. Room: 7338, [email protected], http://www.champa.kku.ac.th/thanaset/
: 1. . . 2. : 2540. 2. Lehninger AL, Nelson DL and Cox MM. Principles of Biochemistry. 2nd edition. Irving Place, New York: Worth publishers, 1993. 3. Lewin B. Gene VII. New York; Oxford University Press Inc., 2000. 4. Mathews CK, Van Holde KE, and Ahern KG. Biochemistry. 3rd edition. Sanfrancisco, California: Benjamin/Cummings, an imprint of Addison Wesley Longman, 2000. 5. Nelson DL and Cox MM. Lehninger Principles of Biochemistry. 3rd edition. New York: Worth pub1ishers, 2000. 6. Voet D and Voet JG. Biochemistry. 3rd edition. Wiley International edition. Hoboken, New Jersey: John Wiley & Sons. Inc. , 2004.
-
1. DNA (DNA Synthesis) DNA (DNA replication) 1.1 DNA E. coli (DNA replication in E.coli) 1.2 DNA (DNA replication in Eukaryotic cells) 2. (Gene expression) 2.1 RNA (RNA Synthesis) (Transcription)
2.2 (Protein Synthesis) (Translation)
3. 3.1 3.2 : DNA replication, Transcription, Translation Regulation of Gene Expression
-
RNA DNA
double strand
-
DNA
RNA
Protein
DNA replication
Reverse transcription
RNA replication
Translation
Transcription
Central dogma
-
DNA replication
= DNA Synthesis
=
DNA replication
1. (Semiconservative)- DNA DNA DNA (template)
-
Parental duplex
Two daughterduplexes
Semiconservativereplication
-
2. (Bidirectional)- (Origin of
replication) 2 ORI
ORI
ORI ORI
ORI ORI
ORI
ORI ORI
ORIORI
ORIORI
ORI ORI
(Replication fork)
Bidirectionalreplication
E.coli
Eukaryote
*** origin of replication
oriC
*** origin of replication
1
-
DNA replication in E.coli
: DNA polymerase
DNA polymerase E.coli 3 - DNA polymerase I- DNA polymerase II- DNA polymerase III
DNA replication E.coli DNA polymerase III
-
DNA polymerase E.coli DNA polymerase I ()
polypeptide 3 activities
1. Polymerase activity
2. 3 5 exonuclease activity
3. 5 3 exonuclease activity
-
1. Polymerase activity
DNA 5 3
phosphodiester 5--P dNTP 3-OH DNA
-
NT 3 phosphodiester
NT
proofreading activity
2. 3 5 exonuclease activity
-
3. 5 3 exonuclease activity
NT 5 phosphodiester
RNA primer
DNA replication
DNA
excision-repair
activity
-
DNA polymerase I
protease
Large fragment(Klenow fragment)
Small fragment
+
Polymerase activity 3 5 exonucleaseActivity (proofreading)
5 3 exonucleaseactivity
-
1: DNA Polymerase E.coli DNA Polymerase
I II III (Structural gene)* polA polB polC (dnaE)
(Subunits)
1 > 4 >10
(Molecular weight)
103,000 88,000 830,000
3 5 Exonuclease (proofreading)
5 3 Exonuclease
DNA Polymerization rate (nucleotides / )
16-20 40 250-1000
Processivity (nucleotides added before dissociation)
3-200
1,500
>500,000
(Biological function)
DNA RNA primer
DNA SOS DNA Repair
DNA DNA
-
DNA replication E.coli
(subunits) --->
DNA polymerase III holoenzyme
DNA polymerase III complex
DNA polymerase III
-
2: (subunits) DNA polymerase III E.coli
holo
enzyme
2 132,000 polC (dnaE)
Polymerization Activity Core
2 27,000 dnaQ (mutD)
35Proofreading Polymerase exonuclease
2 10,000 holE
2 71,000 dnaX ; DNA polymerase III dimer
2 52,000 DnaX* Clamp-loading 1 35,000 HolA compex load -subunit 1 33,000 HolB DNA 1 15,000 holC lagging strand 1 12,000 holD
4 37,000 dnaN (DNA clamp)
processivity
-
subunit DNA Polymerase III processivity
Sliding clamp
-
:
DNA 2
DNA replication E.coli
DNA 5 3
1. leading strand = DNA
2. lagging strand = DNA
= Okazaki fragment
DNA replisome DNA replicase system
-
DNA E.coli :
1. Initiation step:
8 ( 3)
DNA E.coli ori C
Consensus sequenceGATCTNTTNTTTT
Consensus sequenceTTATCCACA
Tandem array ofthree 13 bp sequences
Binding sites for DnaA protein,four 9 bp sequences
DNA DnaA
-
3: oriC E.coli
DnaA protein 52,000 1 oriC oriC
DnaB protein (helicase)
300,000 6*
DnaC protein 29,000 1 DnaB oriC
HU 19,000 2 ; ;
Primase (DnaG protein)
60,000 1 RNA primers; primosome
SSB 75,600 4* RNA polymerase
454,000 5 DnaA
DNA topoisomerase II (gyrase)
400,000
4
Dam methylase 32,000 1 5GATC oriC
-
Supercoiledtemplate
Initialcomplex
Opencomplex
1
2
3
4
Preprimingcomplex
DnaAATP+ HU + IHF
(DnaBDnaC)6+ DnaA
DnaC +ADP + PiDnaB
Priming and replication
E.Coli DNA
P1 nuclease = endonuclease Penicillium citrinum
DnaB = helicase
HU = histonelike proteinIHF = integration host factor
-
2. Elongation step: DNA
DNA
DNA 2 leading strand lagging strand
leading strand
primase RNA primer (~10-60 nt)
DNA pol III RNA primer dNTP DNA
RNA primer DNA template Watson & Crick
-
(DnaB)
leading strand
-
lagging strand
Okazaki fragment ~1,000 - 2,000 nt
-
RNA primer
5 3 exonuclease activity
DNA Polymerase I
Polymerase activity DNA Polymerase I
DNA RNA primer
RNA primer
-
Okazaki fragment
DNA Ligase
Enzyme-AMP
AMP--- NAD+ (E.coli)ATP (Eukaryote)
-
DNA polymerase III dimer
-
3. Termination step: DNA
Tus
DNA E.coli Termination sites (Ter sites)
DnaB (Helicase)
DNA 2
-
DNA replication in Eukaryotic cells
S phase interphase cell cycle
Nucleosome(11 nm diameter)
Histone and nonhistoneproteins
Chromatin fiber
-
:
5: E.coli
E.coli
( 1 )
3.9 x 106 ~ 109
(m/)
30 3
(//)
850 60-90
1
1 103-104
0.67 8
0.33 24 : E.coli 37C Hela cells
~10
-
(..)
lagging strand E.coli ~ 135 nt
DNA lagging strand
-
6: DNA polymerase DNA polymerase primase
Lagging strand
Leading strand
4 1 4 (identical) 2 ? , kDa
160-185 40 125 125 210-230 or 125-140
KM dNTPs, M
2-5 10a 0.5 2-4 ?
Processivity ()
Processivity ( PCNA)
35 exonuclease b 2, 3- dideoxy-NTPs
arabiosyl-CTP
?
aphidicolin
(..)
-
(..continued)
DNA : DNA 2 DNA
Telomerase DNA gap 5
gap 5 lagging strand 5
-
Template = double strand DNAPrimer = single strand DNA
-
5 GATCGAACCACGGA 3DNA template:
DNA primer: 5 CCGTG 3
5 GATCGAACCACGGA 3GTGCC 5
2C : 3T : 2G : 1A
CTAGCTTG
-
DNA
RNA
Protein
DNA replication
Translation
Transcription
Gene expression
DNA ( RNA )
DNA ( RNA . ) .
- Translation- Transcription
-
Transcription: DNA RNA
RNA
RNA 3
1. mRNA :
: template
2. tRNA : tRNA
: mRNA
-
RNA 3 RNA
1. RNA RNA polymerase: RNA 5--P NTP 3-OH RNA phosphodiester
(Transcription)
2. RNA primer
3. DNA 3 5 template RNA
3. rRNA : rRNA
:
-
(sense strand)
(nontemplate strand)
(antisense/noncoding strnad)
5
5
3
3
5 3
-
E.coli RNA polymerase 1 mRNA, tRNA rRNA
E.coli
(Dalton)
1
36,500 2 RNA
151,000 1 RNA RNA
155,000 1
70,000a 1
11,000 1 a 70 kDa
7: RNA polymerase E.coli
-
Holoenzyme RNA pol core enzyme
subunit Holoenzyme DNA promoter
-
E.coli 3 initiation step, elongation step termination step
1. Initiation step :
RNA pol DNA promoter subunit Holoenzyme
DNA -10 region Pribnow box
(Pribnow box)
-
DNA template 3 5
5 RNA triphosphate; 5-ppp
riboNT phosphodiester 5 3
subunit core enzyme RNA ~7-10 nt
rifampicin subunit RNA pol
RNA pol DNA template primer
-
2. Elongation step :
DNA RNA pol RNA
RNA 5 3
Actinomycin D G-C DNA
-
2 - () rho factor-dependent- rho factor-independent
1. rho factor-dependent
RNA RNA RNA pol DNA
activity ATP-dependent RNA-DNA helicase
3. Termination step :
DNA-RNA polymerase
3 RNA
RNA RNA polymerase
-
2. rho factor-independent
Stem-loop structure 3 RNA RNA pol DNA
Stem-loop structure hairpin structure symmetrical G-C rich segments 3 15-20 nt
stem-loop structure 3 U ~ 4-8
RNA DNA
-
E.coli
RNA pol 3
RNA polymerase
RNA
RNA polymerase I Nucleus (nucleolus)
Pre-rRNA (except 5S)
RNA polymerase II Nucleus Pre-mRNA, some small nuclear RNAs
RNA polymerase III Nucleus Pre-tRNA, 5S rRNA, other small RNAs
Mitochondrial Mitochondrion Mitochondrial RNA RNA polymerasea
Chloroplast Chloroplast Chloroplast RNA RNA polymerasea
a RNA polymerase
8 : RNA polymerase
-
. promoter promoter RNA pol I Pre-rRNA
(continued)
promoter RNA pol II Pre-mRNA TATA box, regulatory seq.(CCAAT box, GC box) Inr seq.
RNA pol II Initiation & Elongation steps Transcription factors
-
9 Proteins required for transcription at the RNA polymerase II promoters of eukaryotes ( Nelson DL and Cox MM. Lehninger Principles of Biochemistry. 3rd edition. 2000; P. 987) Transcription factors
Number of subunits
Subunit Mr
Functions
Initiation RNA polymerase II TBP (TATA-binding protein) TFIIA TFIIB TFIID TFIIE TFIIF TFIIH Elongation* ELL
P-TEFb SII (TFIIS) Elongin (SIII)
12 1 3 1 12 2 2 12 1 2 1 3
10,000-220,000 38,000 12,000, 19,000, 35,000 35,000 15,000-250,000 34,000, 57,000 30,000, 74,000 35,000-89,000 80,000 43,000, 124,000 38,000 15,000, 18,000, 110,000
RNA TATA box Stabilizes binding of TFIIB and TBP to the promoter Binds to TBP; recruits RNA polymerase-TFIIF complex Interacts with positive and negative regulatory protein Recruits TFIIH; ATPase and helicase activities Binds tightly to RNA polymerase II; binds to TFIIB and prevents binding of RNA polymerase to nonspecific DNA sequences UnwindsDNA at promoter; phosphorylates RNA polymerase; recruits nucleotide-excision repair complex
* All elongation factors suppress the pausing or arrest of transcription by the RNA polymerase II-TFIIF complex. The name is derived from the term Eleven-nineteen Lysine-rich Leukemia. The gene for the factor ELL is the site of chromosomal recombination events frequently associated with the cancerous condition known as acute myeloid leukemia.
-
(Termination step)
-
- RNA (post-transcriptional RNA processing)
mRNA -**
rRNA tRNA -
RNA (mRNA, rRNA, tRNA) -
- mRNA :
1. RNA Splicing :
intron exon template
-
intron = primary transcript
intervening sequence
exon = primary transcript
-
- primary transcript mRNA (heterogeneous nuclear RNA; hnRNA) small nuclear ribonucleoproteinparticle (snRNP snurps)
RNA intron histone
intron ribozyme primary transcript splicing Self Splicing
intron primary transcript tRNA ATP
-
7-methylguanosine 5 nt RNA
5, 5-triphosphate 5-cap
5-cap mRNA 5-cap ribosome mRNA
2. 5 cap :
-
3. poly(A) tail :
A 20-250 3 mRNA
mRNA
5-cap splicing
-
- rRNA : primary transcript rRNA pre-rRNA
- pre-rRNA E.coli
I
-
- pre-rRNA Eukaryote
- nucleolus
*** tRNA
-
- tRNA : tRNA E.coli : - 30S pre-rRNA tRNA tRNA
tRNA : tRNA
: primary transcript 1 tRNA 2-7
Clover leaf
-
DNA
RNA
DNA replication
Protein
Translation
Transcription
-
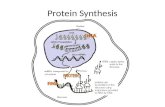
(protein synthesis) peptide
mRNA template mRNA
Translation :
5 3 mRNA Polycistronic mRNA
Monocistronic mRNA
-
cytoplasm
polypeptide (N) (C)
-
(Genetic Code) mRNA = 3 (triplet code codon) 1
AUG = initiation codon
: Met (euk.): fMet (prok.)
1 1
degeneracy of genetic code
-
codon 1 2
frameshift
universal code
-
mitochondria:
Codon Universal code Mitochondrial code
UGA Stop Trp AUA Ile Met and
initiation AGA Arg Stop
AGG Arg Stop
mRNA tRNA tRNA codon mRNA hydrogen
3 tRNA codon mRNA anticodon
-
3 2 anticodon 1 2 codon Watson-Crick
1 anticodon 3 codon Watson-Crick
1 anticodon wobble position
-
cytoplasm
ribosomal protein rRNA
ER
RER
tRNA 3
1. Peptidyl site (P site)
2. Aminoacyl site (A site)
3. Exit site (E site)
-
Polysome Polyribosome:10-100
-
tRNA
tRNA
tRNA
tRNA aminoacyl-tRNA
tRNA aminoacyl-tRNA synthetase
20
2
1. Amino acid + ATP + enzyme enzyme(aminoacyl-AMP) + PPi
2. tRNA + enzyme(aminoacyl-AMP) aminoacyl-tRNA + AMP + enzyme
-
3-OH A 3 tRNA
tRNA
tRNAMet
Met-tRNAMet
Met
(methionyl-tRNA)
***
3
-
3 initiation, elongation, termination steps
1. Initiation step:
E.coli:- 30S ribosome ( 16S rRNA )- 50S ribosome ( 23S rRNA )- mRNA- fMet-tRNAfMet- Initiation factors (IF-1, IF-2 IF-3)- GTP- Mg2+
30S initiation complex
70S initiation complex
-
initiation factors 9
eIF initiation Factor
(IF)
IF-1 tRNA A site
IF-2 fMet-tRNAfMet mRNA P site
IF-3 50S 30S 30S fMet-tRNAfMet P site
(eIF)
eIF2 Met-tRNAfMet 40S
eIF2B, eIF3 2 40S
eIF4A RNA helicase (secondary structure) mRNA mRNA 40S eIF4F (eIF4F complex)
eIF4B mRNA AUG mRNA
eIF4E 5cap mRNA; eIF4FeIF4G eIF4E poly(A) binding protein (PAB);
eIF4F
eIF5 40S 60S 40S 80S initiation complex
eIF6 80S 40S 60S
13 ( 50)
-
2. Elongation step: E.coli:- 70S initiation complex- aminoacyl-tRNA - Elongation factors (EF-Tu, EF-Ts
EF-G)- GTP
70S initiation complex
1 peptide 3
1. Elongation Step 1
-
2. Elongation Step 2
Peptidyltransferase= 23S rRNA***(ribozyme)
3. Elongation Step 3
Translocation
EF-G =Translocase
-
3. Termination step:
E.coli :- (UAG,
UAA UGA)- Termination factors
Release factors (RF-1, RF-2 RF-3)
- GTP
Release factor 1 eRF
-
(posttranslational modification)
polypeptide
:
: insulin ( preproinsulin)
: (glycosylation) glycoprotein ( glycosyltransferase ER)
: (Disulfide bond formation) insulin ( -SH cysteine)
: (attachment of prosthetic group) Hemoglobin cytochrome ( heme prothetic group)
: (Methylation) Cytochrome c ( Lys Glutamic acid )
: (Hydroxylation) Collagen (-OH Lys proline )
-
Regulation of Gene Expression :
(constitutive gene expression)
Glycolysis Crebs cycle
inducible protein
repressible protein
(transcription initiation)
-
transcription initiation
-
1. : regulatory protein, attenuation, antiterminator (regulatory protein):
regulatory protein regulatory gene : mRNA
- activator- repressor
metabolism
Polycistronic gene
DNA operon
-
:lac operon & trp operon
allolactose IPTG = inducer
Z = -galactosidase
lac operon
lactose allolactose Lac repressor
(IPTG)
lactose
Negative regulation
-
glucose lactose lac operon
Activator
Positive regulation
-
trp operonaporepressor
trpB trpA
-
trp operon transcription translation :
Trp
Trp
attenuation :
-
antiterminator :
antiterminator RNA pol DNA template RNA pol mRNA
2. :
:- polycistronic mRNA- translational frameshift- translational repressor- antisense RNA
-
:- metabolism
-polypeptide mRNA -transcription translation - exon
1. :
:-promoter,transcriptional activator protein (TAP), enhancer
-
TAP 4 : Helix-turn-helix, Zinc finger, Leucine Zipper Helix-loop-helix DNA
promoter 1
promoter response element heat shock elements
2. :
, mRNA