1 Mechanisms of transcription RNA splicing Translation The genetic code Section III: Expression of...
-
Upload
lynn-lizbeth-wilcox -
Category
Documents
-
view
224 -
download
3
Transcript of 1 Mechanisms of transcription RNA splicing Translation The genetic code Section III: Expression of...
1
Mechanisms of transcription RNA splicingTranslationThe genetic code
Section III: Expression Section III: Expression of the Genomeof the Genome
Section III: Expression Section III: Expression of the Genomeof the Genome
2
Mechanisms of Mechanisms of TranscriptionTranscription
1. RNA polymerase and transcription cycle
2. The transcription cycle in bacteria
3. Transcription in eukaryotes
•Molecular Biology Course
3
The Central DogmaThe Central Dogma
DNA RNA PROTEINTranscription TranslationTranscription Translation
replicationreplication
4
Transcription is very similar to DNA replication but there are some important differences:
1.RNA is made of ribonucleotides2.RNA polymerase catalyzes the
reaction3.The synthesized RNA does not
remain base-paired to the template DNA strand
4.Less accurate (error rate: 10-4)
5
5.Transcription selectively copies only certain parts of the genome and makes one to several hundred, or even thousand, copies of any given section of the genome. (Replication?)
8
RNA polymerasesRNA polymerases come in come in different forms, but share different forms, but share many featuresmany features
RNA polymerases performs essentially the same reaction in all cells
Bacteria have only a single RNA polymerases while in eukaryotic cells there are three: RNA Pol I, II and III
RN
A p
oly
mera
se a
nd
the tra
nscrip
tion
cycle
9
RNA Pol II is the focus of eukaryotic transcription, because it is the most studied polymerase, and is also responsible for transcribing most genes-indeed, essentially all protein-encoding genes
RNA Pol I transcribe the large ribosomal RNA precursor gene
RNA Pol II transcribe tRNA gene, some small nuclear RNA genes and the 5S rRNA genes
12
’
RPB3
RPB11
RPB2
RPB1
RPB6
RNAP Comparison
The same color indicate the homologous of the two enzymes
prokaryotic
eukaryotic
14
There are various channels allowing DNA, RNA and ribonucleotides (rNTPs) into and out of the enzyme’s active center cleft
15
Transcription by RNA Transcription by RNA polymerase proceeds polymerase proceeds in a series of stepsin a series of steps
InitiationElongationTermination
RN
A p
oly
mera
se a
nd
the tra
nscrip
tion
cycle
16
Initiation Promoter: the DNA sequence that
initially binds the RNA polymerase The structure of promoter-
polymerase complex undergoes structural changes to proceed transcription
DNA at the transcription site unwinds and a “bubble” forms
Direction of RNA synthesis occurs in a 5’-3’ direction (3’-end growing)
18
Elongation Once the RNA polymerase has
synthesized a short stretch of RNA (~ 10 nt), transcription shifts into the elongation phase.
This transition requires further conformational change in polymerase that leads it to grip the template more firmly.
Functions: synthesis RNA, unwinds the DNA in front, re-anneals it behind, dissociates the growing RNA chain
19
Termination After the polymerase
transcribes the length of the gene (or genes), it will stop and release the RNA transcript.
In some cells, termination occurs at the specific and well-defined DNA sequences called terminators. Some cells lack such termination sequences.
21
Transcription initiation Transcription initiation involves 3 defined stepsinvolves 3 defined steps
1. Forming closed complex2. Forming open complex3. Promoter escape
RN
A p
oly
mera
se a
nd
the tra
nscrip
tion
cycle
22
The initial binding of polymerase to a promoter
DNA remains double stranded
The enzyme is bound to one face of the helix
Closed complex
23
Open complex
the DNA strand separate over a distance of ~14 bp (-11 to +3 ) around the start site (+1 site)
Replication bubble forms
24
Stable ternary complex
The enzyme escapes from the promoter
The transition to the elongation phase
Stable ternary complex =DNA +RNA + enzyme
RN
A p
oly
mera
se a
nd
the tra
nscrip
tion
cycle
26
Bacterial promotersBacterial promoters varyvary in in strength and sequences, strength and sequences, but have but have certaincertain defining defining featuresfeatures
Th
e tra
nscrip
tion
cycle
in b
acte
ria
27,
Holoenzyme= Holoenzyme= factorfactor + + core enzymecore enzyme
In cell, RNA polymerase initiates transcription only at promoters. Who confers the polymerase binding specificity?
28
The predominant factor in E. coli is 70.
Promoter recognized by 70 contains two conserved sequences (-35 and –10 regions/elements) separated by a non-specific stretch of 17-19 nt.
Position +1 is the transcription start site.
Promoters recognized by E. coli factor
29
bacterial promoter
The distance is conserved
1. 70 promoters contain recognizable –35 and –10 regions, but the sequences are not identical.
2. Comparison of many different promoters derives the consensus sequences reflecting preferred –10 and –35 regions
31
3.Promoters with sequences closer to the consensus are generally “stronger” than those match less well. (What does “stronger” mean?)
4.The strength of the promoter describes how many transcripts it initiates in a given time.
32
bacterial promoter
Confers additional specificity
UP-element is an additional DNA elements that increases polymerase binding by providing the additional interaction site for RNA polymerase
33
bacterial promoter
Another class of 70 promoter lacks a –35 region and has an “extended –10 element” compensating for the absence of –35 region
34
The The factor mediates factor mediates binding of polymerase to binding of polymerase to the promoterthe promoter
Th
e tra
nscrip
tion
cycle
in b
acte
ria
The 70 factor comprises four regions called region 1 to region 4.
35
regions of
Region 4 recognizes -35 element Region 2 recognizes -10 element
Region 3 recognizes the extended –10 element
36
Binding of –35 Two helices within region 4 form a common DNA-binding motif, called a helix-turn-helix motif
Helix-turn-helix DNA-binding motif
One helix inserts into the DNA major groove interacting with the bases at the –35 region. The other helix lies across the top of the groove, contacting the DNA backbone
37
Interaction with –10 is more elaborate ( 精细 ) and less understood
The -10 region is within DNA melting region
The helix recognizing –10 can interacts with bases on the non-template strand to stabilize the melted DNA.
38
UP-element is recognized by a carboxyl terminal domain of the -subunit (CTD), but not by factor
and subunits recruit RNA pol core enzyme to the promoter
39
Transition to the open Transition to the open complex involves structural complex involves structural changes in RNA polymerase changes in RNA polymerase and in the promoter DNAand in the promoter DNA
This transition is called Isomerization ( 异构化 )
Th
e tra
nscrip
tion
cycle
in b
acte
ria
40
For 70 –containing RNA polymerase, isomerization is a spontaneous conformational change in the DNA-enzyme complex to a more energetically favorable form. (No extra energy requirement)
41
the opening of the DNA double helix, called “melting”, at positions -11 and +3.
Change of the promoter DNA
42
The striking structural change in the polymerase
1. the and ’ pincers down tightly on the downstream DNA
2. A major shift occurs in the N-terminal region of (region 1.1) shifts. In the closed complex, region 1.1 is in the active center; in the open complex, the region 1.1 shift to the outside of the center, allowing DNA access to the cleft
44
Transcription is initiated by Transcription is initiated by RNA polymerase RNA polymerase withoutwithout the need for the need for a primera primer
Initiation requires: The initiating NTP (usually an A)
is placed in the active site The initiating ATP is held tightly
in the correct orientation by extensive interactions with the holoenzyme
Th
e tra
nscrip
tion
cycle
in b
acte
ria
45
RNA polymerase RNA polymerase synthesizes several short synthesizes several short RNAs before entering the RNAs before entering the elongation phaseelongation phase
Abortive initiation: the enzyme synthesizes and releases short RNA molecules less than 10 nt.
Th
e tra
nscrip
tion
cycle
in b
acte
ria
46
Structural barrier for the abortive initiation
The 3.2 region of factor lies in the middle of the RNA exit channel in the open complex.
Ejection of this region from the channel (1) is necessary for further RNA elongation, (2) takes the enzyme several attempts
48
The elongating polymerase The elongating polymerase is a processive machine is a processive machine that that synthesizessynthesizes and and proofreadsproofreads RNA RNA
Th
e tra
nscrip
tion
cycle
in b
acte
ria
49
1. DNA enters the polymerase between the pincers
2. Strand separation in the catalytic cleft
3. NTP addition4. RNA product spooling out (Only 8-
9 nts of the growing RNA remain base-paired with the DNA template at any given time)
5. DNA strand annealing in behind
Synthesizing by RNA polymerase
50
Pyrohosphorolytic (焦磷酸键解) editing: the enzyme catalyzes the removal of an incorrectly inserted ribonucleotide by reincorporation of PPi.
Hydrolytic (水解) editing: the enzyme backtracks by one or more nucleotides and removes the error-containing sequence. This is stimulated by Gre factor, a elongation stimulation factor.
Proofreading by RNA polymerase
51
Transcription is terminated Transcription is terminated by signals within the RNA by signals within the RNA sequencesequence
Terminators: the sequences that trigger the elongation polymerase to dissociate from the DNA Rho-dependent (requires Rho
protein) Rho-independent, also called
intrinsic ( 内在 ) terminator
Th
e tra
nscrip
tion
cycle
in b
acte
ria
52
Rho-independent terminator contains a short inverted repeat (~20 bp) and a stretch of ~8 A:T base pairs.
54
Rho () -dependent terminators
Have less well-characterized RNA elements, and requires Rho protein for termination
Rho is a ring-shaped single-stranded RNA binding protein, like SSB
Rho binding can wrest ( 夺取 ) the RNA from the polymerase-template complex using the energy from ATP hydrolysis
Rho binds to rut ( utilization) RNA sites Rho does not bind the translating RNA
57
Comparison of eukaryotic and prokaryotic RNA polymerases
Eukaryotes: Three polymerase transcribes different class of genes: Pol I-large rRNA genes; Pol II-mRNA genes; Pol III- tRNA, 5S rRNA and small nuclear RNA genes (U6)
Prokaryotes: one polymerase transcribes all genes
58
Comparison of eukaryotic and prokaryotic promoter recognition
Eukaryotes: general transcription factors (GTFs). TFI factors for RNAP I, TFII factors for RNAP II and TFIII factors for RNAP III
Prokaryotes: factors
59
In addition to the RNAP and GTFs, in vivo transcription also requires Mediator complex DNA-binding regulatory
proteins chromatin-modifying
enzymesWhy??
60
RNA polymerase II RNA polymerase II core core promoterspromoters are made up of are made up of combinations of combinations of 44 different different sequence elementssequence elements
Eukaryotic core promoter (~40 nt): the minimal set of sequence elements required for accurate transcription initiation by the Pol II machinery in vitro
Th
e tra
nscrip
tion
in e
ukary
ote
s
61
TFIIB recognition element (BRE) The TATA element/box Initiator (Inr) The downstream promoter element
(DPE)
Pol II core promoter
62
The sequence elements other than the core promoter that are required to regulate the transcription efficiency
Those increasing transcription: Promoter proximal elements Upstream activator sequences
(UASs) EnhancersThose repressing elements:
silencers, boundary elements, insulators ( 绝缘体 )
Regulatory sequences
63
RNA Pol II forms a pre-RNA Pol II forms a pre-initiation complex with initiation complex with GTFs at the promoterGTFs at the promoter
The involved GTFIIs (general transcription factor for Pol II) TFIID=TBP (TATA box
binding protein) + TAFs (TBP association factors)
TFIIA, B, F, E, H
Th
e tra
nscrip
tion
in e
ukary
ote
s
64
1. TBP in TFIID binds to the TATA box
2. TFIIA and TFIIB are recruited with TFIIB binding to the BRE
3. RNA Pol II-TFIIF complex is then recruited
4. TFIIE and TFIIH then bind upstream of Pol II to form the pre-initiation complex
5. Promoter melting using energy from ATP hydrolysis by TFIIH )
6. Promoter escapes after the phosphorylation of the CTD tail
65
Promoter escape Stimulated by phosphorylation of
the CTD (C-terminal domain) tail of the RNAP II CTD contains the heptapeptide
repeat Tyr-Ser-Pro-Thr-Ser-Pro-Ser Phosphorylation of the CTD “tail” is
conducted by a number of specific kinases including a subunit of TFIIH
66
TBP binds to and distorts TBP binds to and distorts DNA using aDNA using a sheet sheet inserted into inserted into the minor the minor groovegroove
Unusual (P367 for the detailed mechanism)
The need for that protein to distort the local DNA structure
Th
e tra
nscrip
tion
in e
ukary
ote
s
67
A:T base pairs (TATA box) are favored because they are more readily distorted to allow initial opening of the minor groove
68
The other GTFs also The other GTFs also have specific roles in have specific roles in initiationinitiation
~ 10 TAFs: (1) two of them bind DNA elements at the promoter (Inr and DPE); (2) several are histone-like TAFs and might bind to DNA similar to that histone does; (3) one regulates the binding of TBP to DNA
Th
e tra
nscrip
tion
in e
ukary
ote
s
69
TFIIB: (1) a single polypeptide chain, (2) asymmetric binding to TBP and the promoter DNA (BRE), (3)bridging TBP and the polymerase, (4) the N-terminal inserting in the RNA exit channel resembles the ..
TFIIB-TBP-promoter complex
70
TFIIF: (1) a two subunit factor, (2) binding of Pol II-TFIIF stabilizes the DNA-TBP-TFIIB complex, which is required for the followed factor binding
TFIIE: recruits and regulates TFIIH TFIIH: (1) controls the ATP-dependent
transition of the pre-initiation complex to the open complex, (2) contains 9 subunits and is the largest GTF; two functions as ATPase and one is protein kinase. (3) important for promoter melting and escape. (4) ATPase functions in nucleotide mismatch repair, called transcription-coupled repair.
71
in vivoin vivo, transcription , transcription initiation requires initiation requires additional proteinsadditional proteins The mediator complex Transcriptional regulatory
proteins Nucleosome-modifying enzymesTo counter the real situation that
the DNA template in vivo is packed into nucleosome and chromatin
Th
e tra
nscrip
tion
in e
ukary
ote
s
72
assembly of the pre-initiation complex in presence of mediator, nucleosome modifiers and remodelers, and transcriptional activators
73
MediatorMediator consists of consists of many subunits, some many subunits, some conserved from yeast to conserved from yeast to humanhuman
More than 20 subunits 7 subunits show significant sequence
homology between yeast and human Only subunit Srb4 is essential for
transcription of essentially all Pol II genes in vivo
Organized in modules ( 模块 )
Th
e tra
nscrip
tion
in e
ukary
ote
s
75
Eukaryotic RNA Pol II holoenzyme is a putative preformed complex:
Pol II + mediator + some of GTFs
Prokaryotic RNA Polymerase holoenzyme:
core polymerase + factor
76
A new set of factors A new set of factors stimulate Pol II stimulate Pol II elongation and RNA elongation and RNA proofreadingproofreading
Th
e tra
nscrip
tion
in e
ukary
ote
s
77
Transition from the initiation to elongation involves the Pol II enzyme shedding ( 摆脱 ) most of its initiation factors (GTF and mediators) and recruiting other factors:
(1) Elongation factors: factors that stimulate elongation, such as TFIIS and hSPT5.
(2) RNA processing (RNA 加工 ) factors Recruited to the C-terminal tail of the
CTD of RNAP II to phosphorylate the tail for elongation stimulation, proofreading, and RNA processing like splicing and polyadenylation.
79
Some elongation factors
P-TEFb: phosphorylates CTD Activates hSPT5 Activates TAT-SF1
TFIIS: Stimulates the overall rate of
elongation by resolving the polymerase pausing
Proofreading
80
Elongation polymerase is Elongation polymerase is associated with a new associated with a new set of protein factors set of protein factors required for various required for various types of RNA processingtypes of RNA processing
RNA processing: Capping of the 5’ end of the RNA Splicing of the introns (most
complicated) Poly adenylation ( 多聚腺苷化 ) of the
3’ end
Th
e tra
nscrip
tion
in e
ukary
ote
s
81
Evidence: this is an overlap in proteins involving in those events The elongation factor hSPT5 also
recruits and stimulates the 5’ capping enzyme
The elongation factor TAT-SF1 recruits components for splicing
Elongation, termination of transcription, and RNA processing are interconnected/ coupled ( 偶联的 ) to ensure the coordination ( 协同性 ) of these events
82
Function of poly(A) tail
Increased mRNA stability Increased translational
efficiency Splicing of last intron
83
Function of 5´cap
Protection from degradation Increased translational
efficiency Transport to cytoplasm Splicing of first intron
84
RNA processing 15’ end cappingRNA processing 15’ end capping
The “cap”: a methylated guanine joined to the RNA transcript by a 5’-5’ linkage
The linkage contains 3 phosphates
3 sequential enzymatic reactions
Occurs early
85
Splicing: joining the protein coding sequences Dephosphorylation of Ser5 within
the CTD tail leads to dissociation of capping machinery
Further phosphorylation of Ser2 recruits the splicing machinery
86
3’ end polyadenylation Linked with the termination of
transcription The CTD tail is involved in
recruiting the polyadenylation enzymes
The transcribed poly-A signal triggers the reactions
1. Cleavage of the message2. Addition of poly-A3. Termination of transcription
87
1. CPSF (cleavage and polyadenylation specificity factor) & CstF (cleavage stimulation factor) bind to the poly-A signal, leading to the RNA cleavage 2. Poly-A polymerase (PAP) adds ~ 200 As at the 3’ end of the RNA, using ATP as a substrate
polyadenylation and termination
89
Models to explain the link between polyadenylation and termination (see the animation on your CD) Model 1: The transfer of the 3’
processing enzyme to RNAP II induces conformational change—RNAP II processivity reduces—spontaneous termination
Model 2: absence of a 5’cap on the second RNA molecule—recognized by the RNAP II as improper—terminate transcription
90
RNA Pol I & III recognize RNA Pol I & III recognize distinct promoters , using distinct promoters , using distinct sets of distinct sets of transcription factors, but transcription factors, but still require TBPstill require TBP Pol I: transcribes rRNA precursor
encoding gene (multi-copy gene) Pol III: transcribes tRNA genes,
snRNA genes and 5S rRNA genes
Th
e tra
nscrip
tion
in e
ukary
ote
s
91
Pol I promoter recognition
Pol I promoter region
Upstream control element
UBF binds to the upstream of UCE, bring SL1 to the downstream part of UCE. SL1 in turn recruits RNAP I to the core promoter for transcription
92Pol III core promoter
TFIIIC binds to the promoter, recruiting TFIIIB, which in turn recruits RNAP III
Pol III promoter recognition1. Different forms, 2. locates downstream of the transcription site
93
1. RNA polymerases (RNAP, 真核和原核的异同 ) and transcription cycle (Initiation is more complicate, details in bacteria)
2. Transcription cycle in bacteria: (1) promoters (elements), factor (4
domains), CTD, abortive initiation (why?)
(2) Structures accounting for formation of the closed complex, transitions to open complex and then stable ternary complex.
(3) Elongation and editing by polymerase (10-4)
(4) Termination: Rho-independent and Rho-dependent mechanism
Key points of the chapter
94
3. Transcription cycle in eukaryotes: (1)Promoters (elements), general
transcription factors (GTF), (2)RNAP II transcription---the roles of GTFs and the CTD tail of
RNAP II in promoter recognition, formation of the pre-initiation complex, promoter melting, promoter escape
---in vivo requires mediator complex, nucleosome modifying enzymes and transcription regulatory proteins.
---elongation and proofreading involve a new set of GTFs (What)
---coupled with RNA processing (How)
95
(3)RNAP I and III transcription---GTFs and promoter recognition,
formation of the initiation complex
98
Most of the eukaryotic genes are mosaic ( 嵌合体 ), consisting of intervening sequences separating the coding sequence
Exons ( 外显子 ): the coding sequences Introns ( 内含子 ) : the intervening
sequences RNA splicing: the process by which introns
are removed from the pre-mRNA. Alternative splicing ( 可变剪接 ): some pre-
mRNAs can be spliced in more than one way , generating alternative mRNAs. 60% of the human genes are spliced in this manner.
100
Sequences within the RNA Determine Where Splicing Occurs
Th
e c
hem
istry
of R
NA
sp
licin
g
The borders between introns and exons are marked by specific nucleotide sequences within the pre-mRNAs.
102
5’splice site (5’ 剪接位点 ): the exon-intron boundary at the 5’ end of the intron
3’ splice site (3’ 剪接位点 ): the exon-intron boundary at the 3’ end of the intron
Branch point site ( 分枝位点 ): an A close to the 3’ end of the intron, which is followed by a polypyrimidine tract (Py tract).
103
The intron is removed in a Form Called a Lariat ( 套马索 ) as the Flanking Exons are joined
Two successive transesterification:Step 1: The OH of the conserved A at
the branch site attacks the phosphoryl group of the conserved G in the 5’ splice site. As a result, the 5’ exon is released and the 5’-end of the intron forms a three-way junction structure.
Th
e c
hem
istry
of R
NA
sp
licin
g
106
Step 2: The OH of the 5’ exon attacks the phosphoryl group at the 3’ splice site. As a consequence, the 5’ and 3’ exons are joined and the intron is liberated in the shape of a lariat.
108
Exons from different RNA molecules can be fused by Trans-splicing
Trans-splicing: the process in which two exons carried on different RNA molecules can be spliced together.
Th
e c
hem
istry
of R
NA
sp
licin
g
111
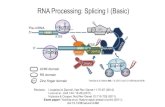
RNA splicing is carried out by a large complex called spliceosome
The above described splicing of introns from pre-mRNA are mediated by the spliceosome.
The spliceosome comprises about 150 proteins and 5 snRNAs.
Many functions of the spliceosome are carried out by its RNA components.
Th
e s
plic
eosom
e m
ach
inery
112
The five RNAs (U1, U2, U4, U5, and U6, 100-300 nt) are called small nuclear RNAs (snRNAs).
The complexes of snRNA and proteins are called small nuclear ribonuclear proteins (snRNP, pronounces “snurps”).
The spliceosome is the largest snRNP, and the exact makeup differs at different stages of the splicing reaction
113
Three roles of snRNPs in splicing1. Recognizing the 5’ splice site and
the branch site.2. Bringing those sites together.3. Catalyzing (or helping to catalyze)
the RNA cleavage.RNA-RNA, RNA-protein and protein-
protein interactions are all important during splicing.
116
Assembly, rearrangement, and catalysis within the spliceosome: the splicing pathway (Fig. 13-8) Assembly step 11. U1 recognize 5’ splice site. 2. One subunit of U2AF binds to Py
tract and the other to the 3’ splice site. The former subunits interacts with BBP and helps it bind to the branch point.
3. Early (E) complex is formed
Sp
licin
g p
ath
ways
117
Assembly step 21. U2 binds to the branch site, and
then A complex is formed.2. The base-pairing between the U2
and the branch site is such that the branch site A is extruded(Figure 13-6). This A residue is available to react with the 5’ splice site.
119
Assembly step 31. U4, U5 and U6 form the tri-snRNP
Particle. 2. With the entry of the tri-snRNP, the A
complex is converted into the B complex.
121
Assembly step 4U1 leaves the complex, and U6
replaces it at the 5’ splice site.U4 is released from the complex,
allowing U6 to interact with U2 (Figure 13-6c).This arrangement called the C complex.
123
Catalysis Step 1:
Formation of the C complexC complex produces the active site, with U2 and U6 RNAs being brought together
Formation of the active siteactive site juxtaposes ( 并置 ) the 5’ splice site of the pre-mRNA and the branch site, allowing the branched A residue to attack the 5’ splice site to accomplish the first transesterfication ( 转酯 ) reaction.
124
Catalysis Step 2:
U5 snRNPU5 snRNP helps to bring the two exons together, and aids the second transesterification reaction, in which the 3’-OH of the 5’ exon attacks the 3’ splice site.
Final Step: Release of the mRNA product
and the snRNPs
127
How does spliceosome find the splice sites reliably
Sp
licin
g p
ath
ways
Two kinds of splice-site recognition errors
Splice sites can be skipped. “Pseudo” splice sites could be
mistakenly recognized, particularly the 3’ splice site.
129
Reasons for the recognition errors
(1) The average exon is 150 nt, and the average intron is about 3,000 nt long (some introns are near 800,000 nt)
It is quite challenging for the spliceosome to identify the exons within a vast ocean of the intronic sequences.
130
(2) The splice site consensus sequence are rather loose. For example, only AGG tri-nucleotides is required for the 3’ splice site, and this consensus sequence occurs every 64 nt theoretically.
131
1. Because the C-terminal tail of the RNA polymerase II carries various splicing proteins, co-transcriptional loading of these proteins to the newly synthesized RNA ensures all the splice sites emerging from RNAP II are readily recognized, thus preventing exon skipping.
Two ways to enhance the accuracy of the splice-site selection
132
2. There is a mechanism to ensure that the splice sites close to exons are recognized preferentially. SR proteins bind to the ESEs (exonic splicing enhancers) present in the exons and promote the use of the nearby splice sites by recruiting the splicing machinery to those sites
133
SR proteins, bound to exonic splicing enhancers (ESEs),
interact with components of splicing machinery, recruiting
them to the nearby splice sites.
134
1. Ensure the accuracy and efficacy of constitutive splicing
2. Regulate alternative splicing3. There are many varieties of SR
proteins. Some are expressed preferentially in certain cell types and control splicing in cell-type specific patterns
SR proteins are essential for splicing
136
Many genes in higher eukaryotes encode RNAs that can be spliced in alternative ways to generate two or more different mRNAs and, thus, different protein products.
Single genes can produce multiple products by alternative splicing
Alte
rnativ
e s
plic
ing
139
Alternative splicing can be either constitutive or regulated
Constitutive alternative splicing: more than one product is always made from a pre-mRNA
Regulative alternative splicing: different forms of mRNA are produced at different time, under different conditions, or in different cell or tissue types
141
Alternative splicing is regulated by activators and repressors
The regulating sequences : exonic (or intronic) splicing enhancers (ESE or ISE) or silencers (ESS and ISS). The former enhance and the latter repress splicing.
Proteins that regulate splicing bind to these specific sites for their action
Alte
rnativ
e s
plic
ing
142
SR proteins binding to enhancers act as activators.
(1) One domain is the RNA-recognition motif (RRM)
(2) The other domain is RS domain rich in arginine and serine. This domain mediates interactions between the SR proteins and proteins within the splicing machinery.
143
hnRNPs binds RNA and act as repressors
1. Most silencers are recognized by hnRNP ( heterogeneous nuclear ribonucleoprotein) family.
2. These proteins bind RNA, but lack the RS domains. Therefore, (1) They cannot recruit the splicing machinery. (2) they block the use of the specific splice sites that they bind.
145
Binds at each end of the exon and conceals ( 隐藏 ) it
Coats the RNA and makes the exons invisible to the splicing machinery
An example of repressors: inhibition of splicing by hnRNPI
146
The outcome of alternative splicing:
1. Producing multiple protein products, called isoforms.
2. Switching on and off the expression of a given gene. In this case, one functional protein is produced by a splicing pattern, and the non-functional proteins are resulted from other splicing patterns.
147
A small group of intron are spliced by minor spliceosome
This spliceosome works on a minority of exons, and those have distinct splice-site sequence.
The chemical pathway is the same as the major spliceosome.
Alte
rnativ
e s
plic
ing
150
Self-splicing introns reveal that RNA can catalyze RNA splicing
Self-splicing introns: the intron itself folds into a specific conformation within the precursor RNA and catalyzes the chemistry of its own release and the exon ligation
Sp
licin
g p
ath
ways
152
Practical definition for self-splicing introns: the introns that can remove themselves from pre-RNAs in the test tube in the absence of any proteins or other RNAs.
There are two classes of self-splicing introns, group I and group II self-splicing introns.
153
Three class of RNA SplicingClass Abundance Mechanism Catalytic
Machinery
Nuclear pre-
mRNA
Very common; used for most eukaryotic
genes
Two transesterification reactions; branch
site A
Major spliceosome
Group II introns
Rare; some eu-Karyotic genes from
organelles and prokaryotes
Same as pre-mRNA
RNA enzyme encoded by
intron (ribozyme)
Group I introns
Rare; nuclear rRNA in some eukaryotics,
organlle genes, and a few prokaryotic
genes
Two transesterific-ation reactions; exogenous G
Same as group II introns
154
The chemistry of group II intron splicing and RNA intermediates produced are the same as that of the nuclear pre-mRNA.
156
Group I introns release a linear intron rather than a lariat
Instead of using a branch point A, group I introns use a free G to attack the 5’ splice site.
This G is attached to the 5’ end of the intron.The 3’-OH group of the 5’ exon attacks the 5’ splice site.
The two-step transesterification reactions are the same as that of splicing of the group II intron and pre-mRNA introns.
Sp
licin
g p
ath
ways
158
1. Smaller than group II introns2. Share a conserved secondary
structure, which includes an “internal guide sequence” base-pairing with the 5’ splice site sequence in the upstream exon.
3. The tertiary structure contains a binding pocket that will accommodate the guanine nucleotide or nucleoside cofactor
Group I introns
159
The similarity of the structures of group II introns and U2-U6
snRNA complex formed to process first transesterification
162
RNA editing is another way of changing the sequence of an mRNA
I. Site specific deamination :1. A specifically targeted C residue
within mRNA is converted into U by the deaminase.
2. The process occurs only in certain tissues or cell types and in a regulated manner.
RN
A e
ditin
g
165
3. Adenosine deamination also occurs in cells. The enzyme ADAR (adenosine deaminase acting on RNA) convert A into Inosine. Insone can base-pair with C, and this change can alter the sequence of the protein.
4. An ion channel expressed in mammalian brains is the target of Adenosine deamination.
166
II Guide RNA-directed uridine insertion or deletion.
1. This form of RNA editing is found in the mitochondria of trypanosomes.
2. Multiple Us are inserted into specific region of mRNAs after transcription (or US may be deleted).
167
3. The addition of Us to the message changes codons and reading frames, completely altering the “meaning” of the message.
4. Us are inserted into the message by guide RNAs (gRNAs) .
168
Having three regions: anchor– directing the gRNAs to the
region of mRNAs it will edit. editing region – determining where
the Us will be inserted poly-U stretch
gRNAs
171
Once processed, mRNA is packaged and exported from the nucleus into the cytoplasm for translation
mR
NA
tran
sp
ort
All the fully processed mRNAs are transported to the cytoplasm for translation into proteins
172
Movement from the nucleus to the cytoplasm is an active and carefully regulated process.
The damaged, misprocessed and liberated introns are retained in the nucleus and degraded.
1.A typical mature mRNA carries a collection of proteins that identifies it as being ready for transport.
2.Export takes place through the nuclear pore complex.
173
3.Once in the cytoplasm, some proteins are discarded and are then imported back to the nucleus for another cycle of mRNA transport. Some proteins stay on the mRNA to facilitate translation.
175
1. Why RNA splicing is important? 2. Chemical reaction: determination of the
splice sites, the products, trans-splicing3. Spliceosome: splicing pathway and
finding the splice sites4. Self-splicing introns and mechanisms5. Alternative splicing and regulation,
alternative spliceosome6. Two different mechanisms of RNA
editing7. mRNA transport-a link to translation
Key points