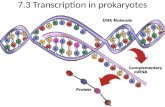
Transcription in Prokaryotes
description
Transcript of Transcription in Prokaryotes

Transcription in Prokaryotes

Transcription: production of mRNA copy of the DNA gene.
Transcription
Eukaryote model

Not all RNA is translated into protein:
• Some RNA is structural - e.g. ribosomal RNA (rRNA)• Some RNA is functional - e.g. transfer RNA (tRNA)• Some RNA is chromosomal (some viruses)
The production of protein-encoding RNA in bacteria is the subject of this lecture.
RNA
Transcription

From which DNA strand is RNA synthesized?
Transcription usually takes place on only ONE of the DNA strands (though not necessarily the same strand throughout the entire chromosome).
Transcription

5'-GTCACCCATGGAGG-3' Nontemplate strand
3'-CAGTGGGTACCTCC-5' Template strand
5'-GUCACCCAUGGAGG-3' mRNA
RNA growth always in the 5' 3' direction
5'3'
3'5'
5' 5' 5'
5'3'
3' 3' 3'
mRNA
mRNA mRNA mRNA
DNADNA
Transcription

RNA Polymerase
The synthesis of RNA from a DNA template is carried out by enzymes known formally as DNA-dependent RNA polymerases, now simply referred to as RNA polymerases
Transcription

ACCCATGG C A5'-GT GG-3' Nontemplate “CODING”strand
3'-CA CC-5‘ Template strand G T TGGGTACC
CAUG C C A5'-GUC
RNA polymerase
Transcription

RNA polymerases have the following properties:
The enzymes are template dependent, requiring double-stranded DNA
The enzymes require the four nucleoside triphosphates (ATP, GTP, CTP, and UTP)
The enzymes copy (read) the template DNA strand in the 3' to 5' direction
The enzymes synthesize the RNA in the 5' to 3' direction
RNA Polymerase

Order of events in Transcription1) Binding of polymerases to the initiation site, the promoter.
Prokaryotic polymerases can recognise the promoter and bind to it directly.
2) Unwinding (melting) of the DNA double helix by a helicase. In prokaryotes the polymerase has the helicase activity.
3) Synthesis of RNA based on the sequence of the DNA template strand, using nucleoside triphosphates (NTPs) to construct RNA.
4) Termination of synthesis. NOTE: the “STOP” codon in the genetic code for the end of peptide synthesis is NOT the end of termination.
Transcription

Prokaryotic RNA Polymerase: Core Enzyme
Chain initiation and interaction with regulatory proteins
Catalytic center: chain initiation and elongation
DNA binding
RNA Polymerase

RNA Polymerase
The core enzyme has the ability to synthesize RNA, however, the initiation point of RNA synthesis is non-specific.
An additional subunit, the sigma factor, is required to initiate RNA synthesis at specific locations in the DNA, termed
the promoter.

RNA Polymerase
Prokaryotic RNA Polymerase: Holoenzyme
Promoter recognition

• For any given gene, RNA synthesis always starts at the same point on the DNA, the promoter. What is a promoter?
• Hypothesis: Because one RNA polymerase copies every gene and binds to the promoter in each gene to do so, the promoters in different genes must have similarities. Similarities in DNA must lie in the sequence of nucleotides so the promoters of every gene must have the same sequence of nucleotides.
Promoters

David Pribnow tested this by comparing the sequences in the promoter regions of five genes from E. coli. He found a conserved sequence of nucleotides in each. This was called the Pribnow box.
Pribnow
Promoters

• The Pribnow box lies 10 nucleotides from the transcription start point (TSP). A second was later found 80 nucleotides away.
-80 -10 TSPTTGACA TATAAT
DNA
Pribnow box
Promoters
5'3'
3'5'
RNA

• The sequences found in promoters are to some extent imaginary. Very few genes actually contain these sequences but they all contain a sequence that is only a few nucleotides different. The consensus sequence is a “best average”.
Consensus sequences
Promoters

GCGTTGTCATGC gene1 AATGTGACAGCT gene2 TGCTAGACACAG gene3 GAATTGAGAAAA gene4 CTTTTCACATTC gene5 AGCTAGACAGGG gene6 TCGTTGGCACCA gene7 CCAATGACCATT gene8 ATGTTGACTTGC gene9 TTGACA consensus not actually
in any of the genes
Promoters

• Just because consensus sequences have been found, this doesn’t mean that they are functional. What is the evidence that they actually work?
Promoters

• Although sequences can vary from the consensus, some mutations stop the promoter from working. In these cases, it demonstrates that the consensus sequence is a functional promoter.
• Genes that are transcribed strongly have sequences more like the ideal consensus than genes that are transcribed weakly.
Promoters

Transcription
Promoter region in DNARNA polymerase
RNA polymerase scans DNA double helix, searching for a promoter site.

(1) Sigma binds to promoter region.
Sigma residues Y425, Y430 and W434 directly involved in the unwinding (melting) of the double helix.
Initiation

(1) Sigma binds to promoter region, recognizing both the -35 and -10 regions. The resulting structure is termed a closed promoter complex.
The promoter is rich in A and T. The AT pair involves two hydrogen
bonds whereas the CG pair involves three hydrogen bonds. Therefore, AT pairs
are easier to separate.
Initiation

Initiation
(2) After the DNA strands have been separated at the promoter region by the helicase activity of the sigma subunit, forming an open promoter complex. The core subunit () can then start to synthesize RNA.
(3) Following initiation, the sigma subunit is released after approx. 10 ribonucleotides have been polymerized,

Elongation
Synthesis of the RNA strand continues until the core polymerase reaches the termination site.

Termination
In prokaryotes, the transcription is terminated by two major mechanisms:
Rho-independent (intrinsic) and Rho-dependent.

Termination
The Rho-independent termination signal is a stretch of 30-40 bp sequence, consisting of many GC residues followed by a series of T ("U" in the transcribed RNA).
The resulting RNA transcript will form a stem-loop structure to terminate transcription
Rho-independent

The terminator has the following structure:
GC rich GC rich PolyA
GC rich GC rich PolyU
DNA
RNA
Complementary
Termination

U CU G G C A U C G C G C G G C C G C G
C
UAAUCCCACAG CAUUUU
GC rich regions
Poly U
RNA
stem-loop structure
Termination

TAG GC rich 1 GC rich 2 ATrich
RNA
RNA polymerase
As transcription proceeds, the two GC rich regions base pair. This leaves a short poly U rich region, which cannot pair strongly enough to hold the RNA onto the DNA. The polymerase comes off with it.
Termination
3 5
35

Rho-dependent.
In vitro, E. coli RNA polymerase holoenzyme transcribes DNA into a very long RNA.
The ability of the in vitro reaction to make natural length RNA is restored by the addition of a protein factor, called rho ().
RNA transcript length:
By holo polymerase in vitro
In vivo
By holo polymerase + rho
Termination

Termination
Analysis of termination sites dependent on rho revel a stem loop structure near the 3‘ end of the RNA, with NO U-rich tale.
Rho binds to RNA and can, if provided with ATP, move along the RNA.
Rho also has ATP-dependent helicase activity.

It has been established that six Rho proteins form a hexamer to terminate transcription, but the precise mechanism is not clear.
(1) The Rho hexamer first binds to the RNA transcript at an upstream site which is 70-80 nucleotides long and rich in C residues .
Model for rho termination
Termination

(2) Upon binding, the Rho hexamer moves along the RNA in the 5-3 direction, trying to catch up with the RNA polymerase.
(3) When the polymerase pauses, which happens when secondary structures form near the 3 end of the RNA, rho catches up and melts the
RNA-DNA duplex in the replication bubble, causing termination.
Termination

Protein
Ribosomes
DNA
RNA
Transcription and translation
Termination

DNA
UAG
Ribosome dissociates from the RNA when they encounter a stop codon.
Termination

DNA
UAG Rho factor binds to specific sites on naked RNA.
(i.e. RNA without ribosomes)
Termination

DNA
UAG
Termination
RNA polymerase pauses at stem loop, while rho moves along RNA, 5-3.

Termination
DNA
UAG
Rho catches up with polymerase, melting RNA-DNA duplex, causing polymerase to dissociate.

Termination of transcription can serve a role in regulating gene expression in prokaryotes
This is the subject of the final lecture in this series.
Termination

Suggested reading
Transcription (2000) In: An Introduction to Genetic Analysis. pp 300-306. Griffiths, A. J. F,. Miller, J. H., Suzuki, D. T., Lewontin, R. C. and Gelbart, W. M. (Eds). Freeman and Company, New York.
http://www.nottingham.ac.uk/bennett-lab/lee.html