Trans-splicing of organelle introns – a detour to ... · dicted from DNA sequences derived from a...
Transcript of Trans-splicing of organelle introns – a detour to ... · dicted from DNA sequences derived from a...

DOI 10.1002/bies.200900036 Review article
Trans-splicing of organelle introns –a detour to continuous RNAsStephanie Glanz and Ulrich Kuck*
¨
Lehrstuhl fur Allgemeine und Molekulare Botanik, Ruhr-Universitat Bochum, Bochum, GermanyIn eukaryotes, RNA trans-splicing is an important RNA-processing form for the end-to-end ligation of primarytranscripts that are derived from separately transcribedexons. So far, three different categories of RNA trans-splicing have been found in organisms as diverse asalgae to man. Here, we review one of these categories:the trans-splicing of discontinuous group II introns,which occurs in chloroplasts and mitochondria of lowereukaryotes and plants. Trans-spliced exons can be pre-dicted from DNA sequences derived from a large numberof sequenced organelle genomes. Further moleculargenetic analysis of mutants has unravelled proteins,some of which being part of high-molecular-weight com-plexes that promote the splicing process. Based on dataderived from the alga Chlamydomonas reinhardtii, amodel is provided which defines the composition of anorganelle spliceosome. This will have a general rele-vance for understanding the function of RNA-processingmachineries in eukaryotic organelles.
Keywords: chloroplasts and mitochondria; group II introns;
organelle spliceosome; trans-splicing
Introduction
Introns were first discovered in 1977 and were subsequently
identified in organisms from all three kingdoms, namely
prokaryotes, archaea and eukaryotes.(1–3) On the basis of
their splicing mechanisms and conserved RNA-folding
patterns, introns are classified into the following categories:
group I and group II introns, nuclear tRNA introns, archaeal
introns and spliceosomal mRNA introns. Group I introns are
widely distributed in genomes of prokaryotic and eukaryotic
organisms but not in archaea,(4) while the tRNA and/or
archaeal introns are found in eukaryotic nuclear tRNAs as
well as in archaeal mRNAs, rRNAs and tRNAs.(5) Spliceo-
somal mRNA introns were exclusively discovered in nuclear
genomes of eukaryotes (see Glossary),(6) whereas group II
introns are restricted to chloroplasts and mitochondria of
lower eukaryotes, plants and some prokaryotes. These
prokaryotes belong to cyanobacterial and proteobacterial
*Correspondence to: U. Kuck, Lehrstuhl fur Allgemeine und Molekulare
Botanik, Fakultat fur Biologie und Biotechnologie, Ruhr-Universitat Bochum,
44780 Bochum, Germany.
E-mail: [email protected]
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
lineages and are believed to be potential ancestors of
chloroplasts and mitochondria. More recently, group II introns
have been discovered outside these eukaryote organelles in
the genome of the archaean Methanosarcina sp.(7) and in the
nuclear genome of the bilaterian Nephtys sp.(8)
During splicing, introns are removed from a precursor
RNA, and the concomitant ligation of exons results in the
formation of a mature transcript. This process of intramole-
cular ligation involves only a single RNA molecule and is
called cis-splicing. In cases, however, when more than one
primary transcript is involved in an intermolecular ligation, the
RNA is processed by trans-splicing. So far, three different
categories of RNA trans-splicing have been found in genomes
as diverse as archaeans to man: the spliced-leader (SL) trans-
splicing, the alternative trans-splicing and the trans-splicing of
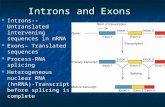
discontinuous group II introns (Fig. 1).
The term SL trans-splicing describes the spliceosomal
transfer of a short RNA sequence (the SL, 15–50 nt) from the 50-
end of a particular non-coding RNA donor molecule (the SL
RNA, 45–140 nt) to unpaired splice acceptor sites on pre-
mRNA molecules. As a result, diverse mRNAs, ranging from a
small proportion to 100% of the mRNA population in different
organisms, acquire a common 50-sequence.(9) This whole
process converts a polycistronic transcript into translatable
monocistronic mRNAs (Fig. 1A). The phenomenon of SL trans-
splicing was first discovered in pre-mRNAs from nuclear genes
of trypanosomes. In this organism, the capped 50-terminal
sequence of SL RNAs is a mini-exon containing an AUG start
codon, which is trans-spliced onto the 50-end of each mRNA.(10)
Since then, SL trans-splicing has been found in diverse groups
of eukaryotes including ascidians, cnidarians, dinoflagellates,
euglenozoans, flatworms, nematodes and rotifers;(11–13) how-
ever, it has an as yet unknown evolutionary origin.
Alternative trans-splicing has recently been discovered in
Drosophila and mammals. In this case, exons located on
separate primary transcripts are selectively joined to produce
mature mRNAs encoding proteins with distinct structures and
functions. Alternative trans-splicing can essentially be
differentiated into intragenic and intergenic trans-splicing
processes. Intragenic trans-splicing is known to occur in rat
and involves exon repetitions, whereas intergenic trans-
splicing was found in man and mouse and involves trans-
splicing of two RNA molecules originating from two different
genes (Fig. 1B).(14–16)
921

Figure 1. Three categories of trans-splicing. A: Spliced leader (SL) trans-splicing. SL trans-splicing accurately joins sequences derived from
separately transcribed small non-coding RNAs and independently transcribed pre-mRNAs. The SL sequence is donated from the SL RNA to pre-
mRNAs to form the 50-terminal mini-exon of the mature mRNA. Outron indicates the 50-segment of a trans-spliced pre-mRNA upstream of the
trans-splice acceptor site. B: Alternative trans-splicing. Intragenic trans-splicing generates mRNAs containing tandem duplications of specific
exons (dark blue) and intergenic trans-splicing generates chimeric mRNAs (grey and light blue) between pre-mRNAs originating from two
different genes (A or B). C: Group II intron trans-splicing. Primary transcripts derived from distantly located exonic sequences are joined end to
end and ligated after assembly and splicing of the flanking group II intron sequences. Exons are shown as boxes and waved lines represent non-
translated sequences.
Review article S. Glanz and U. Kuck
Finally, trans-splicing occurs between transcripts derived
from scrambled gene fragments flanked by discontinuous
group II introns (Fig. 1C). Group II introns are characterised by
a conserved secondary structure configuration. This structure
consists of six major stem-loops, corresponding to domains
D1–D6, radiating from a central core of single-stranded RNA
segments that brings the 50- and 30-splice junctions into close
proximity. For correct folding and catalytic function, the
formation of tertiary interactions is essential. Within group II
introns, two main subclasses of secondary structures, IIA and
IIB, each consisting of two forms (IIA1, IIA2 and IIB1, IIB2),
have been found and more recently, two further classes, IIC
and IID, have been discovered in bacteria.(6,17) A model for the
secondary structure of a typical group IIB intron is shown in
Fig. 2 and excellent reviews concerning the structure and
folding of these introns have been published.(18,19)
Discontinuous group II introns are found in chloroplasts of
algae as well as in chloroplasts and mitochondria of higher
plants and will be the focus of this review. The genes involved
consist of exons that are distributed throughout the genome
and flanked by 50- and 30-regions of group II intron consensus
sequences. Due to the assembly of these regions, a functional
group II intron secondary structure is restored in trans. In
many cases, the correct assembly and splicing reaction
seems to depend on trans-acting factors, which could be RNA
922
molecules and/or proteins. Discontinuous group II introns
share a common splicing mechanism with spliceosomal
introns and are therefore considered as an evolutionary link
between cis-splicing group II introns and nuclear spliceoso-
mal introns. These nuclear spliceosomal introns depend
functionally on the trans-acting spliceosome machinery.(20,21)
Previously, it was assumed that DNA rearrangements within
group II introns result in discontinuous mosaic genes with
exons scattered over the genome.(22)
Chloroplast trans-splicing
The plastid DNA (plastome) of plants and algae codes for
100–140 genes. Most of these genes are organised in
operons and the corresponding polycistronic precursor
transcripts undergo complex posttranscriptional processes.
These processes include the stabilisation of the RNA as well
as cis- and trans-splicing processes, endonucleolytic clea-
vage, RNA editing, and terminal nucleotide additions and/or
deletions. We have analysed a total of 120 sequenced
chloroplast genomes with regard to trans-splicing processes
(Table S1). Our analysis revealed that no trans-splicing events
take place in the three members of the Apicomplexa and in a
single cercozoan species sequenced so far, but that 101
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

Figure 2. Secondary structure model of a typical group IIB intron.
Intron and exon sequences are given as thin and thick lines, respec-
tively. Arabic numerals denote the six conserved domains of group II
introns (D1–D6). The dashed domain D4 is the most variable intron
region and sometimes contains an open reading frame. A branch
point involved in group II intron splicing is circled. Arrows indicate 50 to
30 strand polarity. Potential fragmentation sites of trans-spliced introns
are mapped with arrowheads in domain D1, D2, D3 and D4. Typical
tertiary interactions between exon-binding sites (EBS) and intron-
binding sites (IBS) are indicated. For a complete set of tertiary
interactions in group II introns see Pyle et al.(95) and Fedorova et al.(18)
S. Glanz and U. Kuck Review article
trans-splicing events take place in the other 116 analysed
eukaryotic genomes. Table 1 summarises all characterised
chloroplast trans-splicing introns known to date, and Fig. 3A
shows examples for the organisation of exon sequences in
chloroplast genomes from selected organisms. The first
examples of chloroplast trans-splicing were discovered for the
group II introns from the Marchantia polymorpha and
Nicotiana tabacum rps12 gene, encoding the 30S ribosomal
protein S12, and from the C. reinhardtii psaA gene, encoding
the P700 chloropyll a-apoprotein of photosystem I reaction
centre.(23–26)
Maturation of the rps12 mRNA represents a complex
trans-splicing process, and the corresponding gene shows a
highly similar organisation in chloroplasts of charyophytes
and embryophytes. The mosaic rps12 genes, like other
discontinuous chloroplast genes, contain introns encoding
cis- and/or trans-spliced primary transcripts that are flanked
by sequences showing features of group II introns (Table S1).
In Fig. 3A, two out of four possible organisations of the rps12
genes are depicted. In all cases, splicing of exon 1 and exon 2
occurs in trans, with an intronic fragmentation site in domain
D3, while exon 2 can harbour further two exonic sequences
that can be processed on the RNA level by cis-splicing.
Alternatively, the continuous exon 2 sequence or the
discontinuous exon 2-exon 3 sequences can be located
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
either in a large single copy region or in the two inverted
repeats of chloroplasts genomes. Both the 50- and 30-rps12
gene fragments are organised in operons and are expressed
as polycistronic transcripts.(27,28) Maturation of the rps12
mRNA comprises both endonucleolytic cleavage of the
polycistronic transcripts and trans-splicing of exon 1 with
exon 2 as well as cis-splicing of exons 2 and 3.
Numerous sequencing projects have enabled the in silico
analysis of further genes with putative trans-spliced introns
that has revealed other genes than those mentioned above.
For example, these genes include psaA of the green alga
Scenedesmus obliquus, pbsA (heme oxygenase) of the red
alga Rhodella violacea, petD (subunit IV of cytochrome-b6/f-
complex) and psaC (subunit VII of photosystem I) of the green
algaeStigeoclonium helveticum and Oedogonium cardiacum,
rbcL (large subunit of ribulose 1,5-bisphosphate carboxylase/
oxygenase) of the green algae S. helveticum and Floydiella
terrestris, and ndhH (subunit of the NA(P)DH dehydrogenase
complex) of Triticum aestivum.(29–33)
The highest number of trans-spliced introns within a plastid
genome was predicted from complete sequencing of the
chloroplast DNA from the green alga S. helveticum.(29) Four
discontinuous group II introns were identified in the pre-
mRNAs, one each in the petD and psaC genes and two in the
rbcL gene. Recently, further chloroplast DNAs from the green
algae F. terrestris and O. cardiacum were reported to encode
petD, psaC or rbcL pre-mRNAs that are spliced in trans
(Table 1, Table S1 and Fig. 3B).
To date, the best-analysed trans-splicing process is the
one of the psaA mRNA in the unicellular green alga
C. reinhardtii. This alga can be regarded as the model
organism for the analysis of plastid gene expression during
photosynthesis and the communication between the nucleus
and the chloroplast.(34) Mutant strains with a defective
photosystem I finally served as base for the identification
of trans-splicing processes.(35) As early as 1987, it was
already known that the three exons of the psaA gene are
distributed on the plastome and transcribed separately from
each other(25). Two trans-splicing steps are necessary to form
the mature mRNA. For intron 1, three independently trans-
cribed RNA molecules assemble into a functional group II
intron structure by base pairings and tertiary interactions. This
tripartite group IIB intron is interrupted in domains D1 and D4;
thereby exon 1 is flanked by a portion of domain D1 and exon 2
by the entire domains D4 and D5 as well as a portion of D6.
The rest of domains D1 and D4 as well as the entire domains
D2 and D3 are delivered from a third RNA molecule, the
plastid-encoded tscA RNA, which is 450 nt in length.(36)
The tscA RNA was also detected in C. gelatinosa (376 nt)
and in C. zebra (466 nt) and exhibits sequence identity of
approximately 55% to the tscARNA ofC. reinhardtii for both of
the species. Analysis of the secondary structure of the three
tscA RNAs showed also a high degree of similarity with the
923

Table 1. Distribution of discontinuous chloroplast group II introns from selected algae, higher plants and mosses.
Gene IntronaSplicing
type
Intron typeb,
fragmented
domain Organismc
pbsA pbsA-i1 trans n.d. Rhodella violacea(33)
pbsA-i2 cis? n.d.
petD petD-i1 trans IIB, bi, D1 Oedogonium cardiacum, Stigeoclonium helveticum(29,30)
psaA psaA-i1 trans IIB, tri, D1þD4 Chlamydomonas reinhardtii(25)
psaA-i2d trans IIB, bi, D4
psaA-i1d trans IIB, bi, D4 Scenedesmus obliquus(31)
psaC psaC-i1 trans IIB, bi, D1 Oedogonium cardiacum, Stigeoclonium helveticum(29,30)
rbcL rbcL-i1 trans IIB, bi, D1 Floydiella terrestris, Stigeoclonium helveticum(29,30)
rbcL-i2 trans IIA, bi, D2 Floydiella terrestris, Stigeoclonium helveticum(29,30)
rps12 rps12-i1 trans IIB, bi, D3 Epifagus virginiana, Hordeum vulgare L.,
Marchantia polymorpha, Nicotiana tabacum(23,24,26,98,99)rps12-i2 cis IIA, bi
rps12-i1 trans IIA, bi, D3 Cuscuta europaea, Staurastrum punctulatum,
Zygnema circumcarinatum(100,101)
This list contains examples that have thoroughly been analysed by cDNA and/or Northern or sequence analyses. A complete list of chloroplast
introns shows that in Chlorophyta, 6 out of 27 genomes encode nine trans-spliced RNAs. Similarly, in charyophytes such as Chara vulgaris, 4 out
of 6 genomes encode rps12 RNAs that are predicted to be trans-spliced. The same is true for the 83 embryophytes whose plastomes are
completely sequenced. An exception seem to be the ndhA and ndhH transcripts that are most probably trans-spliced in wheat (Table S1).(97)
Abbreviations and gene products: bi, bipartite intron; D1-4, domains D1-D4 of a typical group II intron; n.d., not determined; pbsA, heme
oxygenase; petD, subunit IV of cytochrome-b6/f-complex; psaA, P700 chloropyll a-apoprotein of photosystem I reaction centre; psaC, subunit VII
of photosystem I; rbcL, large subunit of RubisCO; rps12, 30S ribosomal protein S12; tri, tripartite intronaThe intron nomenclature is based on the flowering plant mitochondrial literature used by Bonen.(43)
bPrediction of the secondary structure and the classification into subclasses IIA and IIB rely on sequence analyses, based on models of Michel(6)
and Michel and Ferat.(42)
Review article S. Glanz and U. Kuck
formation of domains D2 and D3 and partial domains D1 and
D4, all of which are indicative for group II introns.(37) Plastome
sequencing of the green alga S. obliquus revealed that the
psaA gene is split into two exons, which are likewise ligated
by a trans-splicing process (Fig. 3B). This discontinuous
group II intron is located and interrupted within domain D4 at
the same position as the second trans-spliced group II intron
in the psaA gene of C. reinhardtii.(31)
The large number of so far characterised trans-spliced
RNAs allows a comparison of the sites of fragmentation within
the conserved group II intron structure. With the exception of
domains D5 and D6 (Fig. 2), of which domain D5 shows the
most conserved sequence similarity within all group II introns,
all other domains can be fragmented as listed in Table 1. As
described in the next section, this list of multipartite
chloroplast genes can be extended by a number of
mitochondrial genes showing similar fragmented group II
intron structures (Table 2).
Mitochondrial trans-splicing
Mitochondrial genomes (chondriomes) of eukaryotes show a
great variation in size, ranging from about 15 kb in Metazoans
and a few algae to about 2 000 kb in species of the
Cucurbitaceae.(38) Chondriomes that are larger than 200 kb
924
are almost exclusively found in vascular plants with the
exception of the protist species ichthyosporean Amoebidium
parasiticum with a chondriome size of >200 kb.(39) The size
difference of plant chondriomes compared to other eukaryotic
chondriomes is mostly due to the presence of an additional set
of genes, promiscuous DNAs of nuclear or plastid origin,
repetitive DNAs, and numerous group I or group II
introns.(40,41)
Our analysis of 59 sequenced algal and plant chondriomes
identified 19 genomes with genes whose pre-mRNA is
predicted to be trans-spliced (Table S2). Soon after the
discovery of several trans-spliced group II introns in
chloroplasts, mainly DNA sequencing work led to the
detection of split group II introns in a range of mitochondria.
Phylogenetic analyses demonstrated that group II introns of
plant chondriomes can be distinguished from those found in
the chloroplast genome.(17,42) In addition, sequence analyses
revealed that many mitochondrial group II introns of flowering
plants, as compared with bacterial and chloroplast introns,
show variations in the sequence, structure and/or length of
typical group II introns.(43)
Most group II introns in higher plant mitochondria are
processed by cis-splicing; however, a distinct set of tran-
scripts, encoding subunits of the NADH dehydrogenase
complex, are trans-spliced. PCR and phylogenetic analyses
of cis-homologue introns in early branching land plants such
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

Table 2. Distribution of discontinuous mitochondrial introns from selected organisms.
Gene Intron
Splicing
type
Intron typea,
fragmented
domain Organismb
cox1 cox1-i1 trans – Diplonema papilatum, Emiliana huxleyi(54,102)
cox3 cox3-i1 trans – Karlodinium micrum(103)
nad1 nad1-i1 trans IIB, bi, D4 Arabidopsis thaliana, Brassica napus, Oenothera berteriana,
Petunia hybrida, Triticum aestivum, Vicia faba, Zea mays(45–47,104–107)nad1-i2 cis bi
nad1-i3 trans IIB, bi, D4
nad1-i4 cis bi Arabidopsis thaliana, Brassica napus, Oenothera berteriana,
Vicia faba(47,105–107)
trans IIB, bi, D4 Petunia hybrida, Triticum aestivum, Zea mays(45,46,104)
nad2 nad2-i1 cis IIA, bi Arabidopsis thaliana, Brassica napus, Oenothera berteriana,
Triticum aestivum, Zea mays (sterile line) (48,51,108,109)nad2-i2 trans IIA, bi, D4
nad2-i3 cis IIA, bi
nad2-i4 cis IIA, bi
nad3 nad3-i1 trans IIA, bi, D4 Mesostigma viride(110)
nad3-i2 trans IIA, bi, D4
nad5 nad5-i1 cis IIA, bi Arabidopsis thaliana, Brassica napus, Oenothera berteriana,
Triticum aestivum, Vicia faba, Zea mays(49,105,111,112)nad5-i2 trans IIA, bi, D4
nad5-i3 trans IIB, bi, D4 Arabidopsis thaliana, Brassica napus, Triticum aestivum, Vicia faba,
Zea mays(49,105,111,112)
trans IIB, tri, D4 Oenothera berteriana(49)
nad5-i4 cis IIA, bi Arabidopsis thaliana, Brassica napus, Oenothera berteriana,
Triticum aestivum, Vicia faba, Zea mays(49,105,111,112)
This list contains examples that have thoroughly been analysed by cDNA and/or Northern or sequence analyses. A complete list of all organelle
introns predicted to be trans-spliced is given in the supplemental material (Table S2).
Abbreviations and gene products: bi, bipartite intron; cox1, cox3, subunits of cytochrome c oxidase; D4, domain D4 of a group II intron; nad1,
nad2, nad3, nad5, subunits of NADH dehydrogenase complex; n.d., not determined; tri, tripartite intron.aThe prediction of the secondary structure as well as the classification into the subclasses IIA or IIB are based on sequence analyses, according
to the models of Michel(6) and Michel and Ferat.(42)
bAccession numbers are given in the supplemental material (Table S2).
S. Glanz and U. Kuck Review article
as ferns, horsetails, hornworts and mosses have suggested
that trans-spliced introns might have evolved from originally
cis-arranged continuous exon–intron structures by disruption
due to DNA rearrangements.(44) These genes include
nad1,(45–47) nad2(48) and nad5(49) (see Table 2 and Fig. 3C)
and recent sequencing of the first mitochondrial genome of a
gymnosperm, the cycad Cycas taitungensis, revealed trans-
spliced group II introns within the homologous genes.(50)
Similar to their chloroplast counterparts, these mosaic genes
contain introns encoding cis- or trans-spliced primary
transcripts that are flanked by sequences showing features
of group II introns (Fig. 3C).(6,34)
The genomic organisation, e.g. the exon/intron boundaries
as well as the high degree of sequence identity, is conserved
in different organisms. For instance, the intron nad2-i2 is split
at the same position in angiosperms and shows 98%
sequence identity in exons of Arabidopsis, Brassica,
Oenothera and Triticum.(51)
Another conserved example is the third intron of the nad5
gene, which is trans-spliced in all angiosperms investigated.
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
However, this intron can have a bipartite or a tripartite
organisation. In O. berteriana, sequence analyses of the
tripartite organisation showed that an intronic region down-
stream of exon 3 is missing, which is encoded by a distant
genomic region named tix locus (trans-splicing intron
fragment (Fig. 3C)).(52) This tripartite structure is reminiscent
of intron 1 of the chloroplast psaA RNA from C. reinhardtii that
requires the tscA RNA in order to form the correct secondary
structure.(36) Finally, despite their sequence dissimilarities,
both tix and tscA show a highly conserved secondary
structure with fragmentation sites in domains D1 and D4 at
homologous sites.(52)
An unusual trans-splicing mechanism was predicted in
both the dinoflagellate Karlodinium micrum (Alveolata)
(cox3)(53) and the diplonemid Diplonema papillatum (Eugle-
nozoa) (cox1).(54) In D. papillatum, a member of diplonemids,
which are a sister group of kinetoplastids, a fragmented cox1
gene encoded on two different chromosomes was found.
Interestingly, the flanking regions do not exhibit any
characteristics of organelle or nuclear introns nor contain
925

Review article S. Glanz and U. Kuck
conserved sequences adjacent to coding regions, and
therefore, this trans-splicing mechanism can be predicted
to be different from those processes described above for
group II introns.(55) The second remarkable example is the
bipartite cox3 gene (cytochrome c oxidase subunit 3) from the
dinoflagellate K. micrum. Similar to the example mentioned
above, no evidence of flanking group II introns was found.
Instead, numerous inverted repeats in the intergenic
sequences, which might form secondary structures, led to
the assumption that they play a role in splicing. At the splice
site, five adenine nucleotides are found that seem to be
derived from the polyA-tail of the 50-upstream fragment.
Therefore, ligation of exonic sequences seems to occur
without involvement of group II intron sequences, and the
exact mechanism of the splicing process has still to be
resolved.(56)
Trans-acting factors
Although some group II introns exhibit autocatalytic splicing
activity in vitro (see Glossary), both cis- and trans-splicing
introns require cofactors for efficient splicing in vivo.(57) In
principle, factors encoded by organelle or nuclear genomes
can be distinguished, and most of our current knowledge
stems from work with mutants having a defect in RNA
splicing.(34,58) The organelle-encoded components can be
differentiated into RNA and protein factors (Table 3). As
already mentioned above, the tscA RNA from algal chlor-
oplasts and the tix RNA from plant mitochondria are the only
Table 3. Examples of nuclear-encoded factors controlling trans-splicing
Affected
RNA Gene Organism Local. Function
nad1 OTP43 A. thaliana(78) mt trans-splicing of
psaA Raa1 C. reinhardtii(80) cp, m trans-splicing of
(class B, 30 en
RNA; group II
Raa2 C. reinhardtii(92) cp, LDM trans-splicing of
(class A; grou
Raa3 C. reinhardtii(89) cp, sþm trans-splicing of
(class C; grou
Rat1 C. reinhardtii(88) cp, m trans-splicing of
(class C, 30 en
group IIB)
Rat2 C. reinhardtii(88) n.b. trans-splicing of
(class C, 30 en
group IIB)
rps12 ppr4 Z. mays(79) cp, s trans-splicing of
biogenesis of
An extended list of trans-acting factors is given in the supplemental mat
Abbreviations: cp, chloroplast; LDM, low-density membrane; local., local
determined; OTP, organelle transcript processing defect; ppr, pentatricop
psaA tscA RNA; s, stroma.
926
organelle-encoded RNA factors so far known to support the
splicing process in trans.(36,52) As described in the next
chapter, the tscA RNA is most probably part of an organelle
spliceosome that similar to the nuclear spliceosome contains
protein as well as RNA components.(34)
Maturases are highly conserved organelle-encoded pro-
teins, and are usually encoded in domain D4 of some of the
characterised group II introns. These enzymes catalyse the
excision of non-autocatalytic introns, e.g. the excision of the
intron from its own primary transcript, and together with the
intron RNA, they form a ribonucleoprotein (RNP) com-
plex.(59,60) Moreover, maturases have reverse transcriptase
activity, mediating the integration of their ‘mobile’ introns into
new DNA sites (see Glossary).(61) Functional maturases
encoded by bacterial introns were shown to promote splicing,
e.g. of the group II intron Ll.LtrB from Lactococcus lactis.(62)
Recently, the Ll.LtrB intron was used as a model system to
study group II intron trans-splicing in bacteria. A highly
sensitive splicing/conjugation assay was developed and it was
demonstrated that assembly and trans-splicing of a frag-
mented group II intron can efficiently take place in bacterial
cells. The authors mimicked naturally occurring fragmentation
sites, e.g. the site in domain D1 of psaA, and further showed
that the Ll.LtrB intron-encoded maturase LtrA is essential for
trans-splicing.(63,64)
Nuclear-encoded trans-acting factors are the second class
of components that are able to compensate for the loss of
autocatalytic splicing activity in organelle introns (Table S3). It
is generally accepted that mitochondria and chloroplasts are
the result of an endosymbiosis of a-proteobacteria-like and
of group II introns.
Sequence homology
nad1 intron 1 PPR protein
psaA
d processing of tscA
B)
n.d. (possible PPR protein)
psaA
p IIB)
Pseudouridine synthases
psaA
p IIB)
Pyridoxamine 50-phosphate oxidases
psaA
d processing of tscA RNA;
NADþ-binding domain of poly
(ADP-ribose) polymerases
psaA
d processing of tscA RNA;
Domain of a putative RNA-binding
protein of Synechococcus spec.
WH8102
rps12 (intron 1),
ribosomes
PPR protein
erial (Table S3).
isation of gene products; m, membrane; mt, mitochondrion; n.d., not
eptide repeat; Raa, RNA maturation of psaA; Rat, RNA maturation of
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

Figure 3. Organisation of selected chloroplast and mitochondrial genes with trans-spliced RNAs. A: Two possible genomic organisations of
rps12 genes from different sources. The bottom example is representative for higher plants with duplicated 30-exons in the inverted repeat (IR)
regions IRa and IRb and a 50-exon in the large single copy region (LSC). B: Examples of chloroplast genes from diverse algae. C: Schemes of
mitochondrial loci from the five exons of nad1 and nad5. The transcripts of nad1 and nad5 are polycistronic (data not shown).(96) The genome of
O. berteriana was not yet completely sequenced. Exons are represented by black boxes with their corresponding size in base pairs. Arrows
indicate direction of transcription. Cis-spliced introns are depicted as red boxes and trans-spliced introns are marked in yellow. Black double
slashes indicate split gene fragments, which are separately transcribed. Distances in kb were determined from a clockwise orientation of the
chloroplast genomes. Abbreviations of species are as follows: C.v., Chara vulgaris (NC_008097); N.t., Nicotiana tabacum (NC_001879); O.b.,
Oenothera berteriana (X07566, X60046, X60049, X99516); S.o., Scenedesmus obliquus (NC_008101); S.h., Stigeoclonium helveticum
(NC_008372); T.a., Triticum aestivum (NC_007579). Abbreviated gene designations are explained in the legend of Tables 1 and 2.
S. Glanz and U. Kuck Review article
cyanobacteria-like prokaryotes, respectively. This process is
accompanied by the relocation of a major part of the
prokaryotic genomes into the chromosome of the host cell.
As a consequence, the nuclear-encoded organelle proteins
have to be retargeted to their ancestral compartments.(65,66)
In Fig. 4, this situation is depicted for the chloroplast of the
unicellular green alga C. reinhardtii. RNA-processing, trans-
lation, as well as assembly of membrane or membrane-
associated complexes, is dependent on both, organelle- and
nuclear-encoded proteins. The latter are translated on
cytosolic ribosomes and will be transported through the
chloroplast membranes into the inner space of the orga-
nelle.(67–69) Table 3 summarises nuclear-encoded proteins
involved in trans-splicing of group II introns together with the
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
functionally characterised gene products. When known, we
also give the homology of the proteins to other factors as well
as their subcellular localisation.
While some of these trans-acting factors seem to be
specific for only a single intron, other factors are involved
in splicing of a set of introns. Moreover, some of these
nuclear-encoded proteins have acquired, in addition to
their catalytic role, further organelle functions during
splicing. It is therefore assumed that during evolution some
of these nuclear-encoded factors were adapted to the
binding of intron structures, thus playing a role in the splicing
process.(22)
The trans-acting factors involved in splicing of a set of
introns can be divided into three groups. The first group of
927

Figure 4. Dependence of chloroplast biogenesis on nuclear-
encoded factors. During chloroplast biogenesis, coordination of gene
expression is achieved by nuclear-encoded factors that affect RNA
processing, translation, and also assembly of complexes. Chloroplast
multisubunit complexes are thus formed by both nuclear- (arrows) and
chloroplast- (dashed arrows) encoded polypeptides.
Review article S. Glanz and U. Kuck
nuclear-encoded factors comprises enzymes involved in RNA
maturation processes, and recently, new members of this
group with homologies to mitochondrial maturases were
detected. Four genes for group II intron maturases, nMat-1a,
nMat-1b, nMat-2a and nMat-2b, were identified in the nuclear
genomes of both A. thaliana and Oryza sativa. Interestingly,
these maturase-like proteins are not intron-encoded. The
predicted mature proteins show homology to mitochondrial
counterparts and contain putative mitochondrial import
sequences. It is assumed that they were transferred during
evolution from the chondriome to the nuclear genome and
may have retained their role in splicing of mitochondrial group
II introns, as was functionally demonstrated with an
A. thaliana mutant analysis.(70,71) Nuclear-encoded RNA-
editing factors also play a role in splicing. RNA editing in plant
organelles is mediated by specific nuclear-encoded factors(72)
and is essential for the formation and stabilisation of splicing-
competent primary and secondary structures of several
mitochondrial group II introns.(48,51) Almost all protein-coding
transcripts as well as some introns and tRNAs in mitochondria
928
of higher plants are edited. The editing event comprises
mostly C-to-U substitutions.(73) For example, fusion of the
trans-splicing intron nad1-i3 from O. berteriana with
sequences from an autocatalytic splicing intron from
yeast revealed that unedited intron sequences are not able
to form a functional, splicing-competent group II intron
structure.(74)
The second group of nuclear-encoded factors exhibit
repeated motifs of 34–38 amino acids, e.g. like in penta-
tricopeptide repeat (PPR) proteins. The PPR protein family is
characterised by tandem repeats of a motif consisting of a
degenerate 35 amino acid repeat. Several PPR proteins are
encoded in the genomes of animals, fungi and trypano-
somes,(75) and most of these proteins are found in genomes
of higher plants.(76) Already characterised PPR proteins show
RNA-binding features and affect the processing or the
translation of specific RNA molecules in mitochondria and
chloroplasts.(77) Recently, the nuclear OTP43 (organelle
transcript processing defect) gene was shown to be
specifically required for trans-splicing of mitochondrial
nad1-i1 in A. thaliana.(78) Another example is PPR4 of Zea
mays, which is responsible for trans-splicing of the first intron
of the chloroplast rps12 RNA by directly binding to this
intron.(79) Finally, Raa1 (RNA maturation of psaA) from
C. reinhardtii, which is involved in the trans-splicing process of
both psaA introns, also harbours tandem repeats similar to
those found in PPR proteins.(80)
The third and final group of nuclear-encoded factors
represent proteins that cannot be assigned to any classified
function. CRS1 (chloroplast RNA splicing 1) from Z. mays, for
example, contains a new RNA-binding domain, the CRM
domain (chloroplast RNA splicing and ribosome maturation),
which is also found in archaeal and bacterial proteins, involved
in the maturation of ribosomes.(81) This CRM domain is
likewise found in CRS2, another splicing factor from maize.(82)
Moreover, the trans-acting factors in Z. mays are known to be
part of a high-molecular-weight ribonucleoprotein complex
that also contains spliced intron sequences.(79) Similar
data are available from C. reinhardtii, which in recent years
was the subject of intense mutational analyses all of which
show a defect in trans-splicing of the psaA RNA.(80,83) As
detailed below, these studies led to the notion of a complex
chloroplast spliceosome being involved in the splicing
process.
Putative chloroplast spliceosome of themodel alga C. reinhardtii
Trans-splicing of the psaA RNA from C. reinhardtii requires a
plastid-encoded tscA RNA and at least 14 nuclear-encoded
chloroplast factors.(36,84) The corresponding nuclear mutants
are grouped into three classes according to their mode of
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

Figure 5. Trans-splicing of the chloroplast psaA RNA of C. reinhardtii. The psaA gene is fragmented into three independently transcribed
exons, which are flanked by consensus sequences of group II introns (green wavy lines). To generate a mature psaA mRNA, two trans-splicing
steps are necessary. For the formation of the first group II intron, a small chloroplast-encoded RNA (tscA) is required that interacts with precursor
transcripts. The tscA RNA is co-transcribed with chlN and is the subject of various 30 end processing events. Ovals represent nuclear mutant
classes, which are affected in different steps of the trans-splicing process. Colours indicate class A, B and C mutants, respectively. The first group
II intron is labelled as denoted in Fig. 2. Arrowheads indicate the sites of fragmentation. Abbreviations: EBS2, exon binding site 2; IBS2, intron
binding site 2.
S. Glanz and U. Kuck Review article
action (Fig. 5): class A mutants fail to trans-splice exon 2 and 3
primary transcripts; class B mutants neither splice exon 1 and
2 nor exon 2 and 3 primary transcripts; and class C mutants
are not able to splice exon 1 and 2 primary transcripts.(85,86)
Lack of correct splicing in class B and C mutants can take
place at two different levels of RNA processing, i.e.
either splicing of the primary psaA transcripts or 30-end
processing of the tscA RNA is affected. Of note is that
maturation of the tscA precursor is a prerequisite for correct
splicing of exon 1 and 2. To date, five trans-acting factors were
characterised in C. reinhardtii with three belonging to class C
factors (Table 3).
Based on cofractionations and sucrose density
gradient centrifugations using protein extracts of wild-type
or splicing-deficient mutants, Rochaix and co-workers
proposed at least three protein complexes, two of which
are associated with chloroplast RNAs. Thus, the latter two
can be considered as chloroplast RNP (cpRNP) complexes
that might be part of the chloroplast spliceosome. This
concept of a spliceosome-like complex was further supported
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
by reciprocal coimmunoprecipitations, showing that either
protein of the complex can be immunoprecipitated with the
other. However, these data do not yet answer the question
whether the identified splicing factors interact with each
other.(80,87)
Figure 6 provides a current model of chloroplast
spliceosomal complexes and their action in splicing and
integrates data from several experimental approaches. The
tscA RNA participating in the formation of the secondary
group II intron structure is involved in splicing of the first group
II intron (psaA-i1).(36) At least three factors are involved in the
processing of the precursor molecule containing the tscA
RNA. As mentioned above, Raa1 is related to PPR proteins
and is a factor involved in tscA RNA maturation. It contains
two distinct domains of which the C-terminal domain is
involved in processing of the tscA RNA, and the central
domain in splicing of intron 2. The function of both domains
was deciphered when different truncated versions of theRaa1
gene were used in restoration experiments analysing two
different mutants.(80)
929

Figure 6. Model of chloroplast psaA RNA trans-splicing complexes in C. reinhardtii. The scheme integrates data from several groups of
investigators as described in the text. Depicted are proteins and protein complexes required for trans-splicing of three psaA mRNA
precursors. The two secondary structures resemble the folding of 50- and 30-intronic RNAs flanking exon 1, 2 and 3 sequences modified
after Goldschmidt-Clermont et al.(84) For details of the secondary structure from the first intron see Fig. 5. Abbreviations: chlN, subunit of the
light-independent protochlorophyllide reductase; Cpn60, chaperonine 60; cNAPL, chloroplast nucleosome assembly protein-like; pL118B and
pL137H, class B factors, and pL121G, class A factor, which are defined genetically;(87) Raa1-6, RNA maturation of psaA; Rat1-3, RNA
maturation of psaA tscA RNA; tscA, trans-splicing chloroplast. Abbreviations are as described in the legend of Fig. 2, and see also the text for
further details.
Review article S. Glanz and U. Kuck
Rat1 and Rat2 (RNA maturation of psaA tscA RNA), both
of which are encoded by two adjacently located nuclear
genes, are also part of the maturation process. Interestingly,
only when both genes are simultaneously transferred into the
corresponding splicing-deficient mutant, they are able to
restore the wild-type phenotype. The deduced amino acid
sequence of Rat1, which directly interacts with tscA RNA,
shows 26% sequence homology to the conserved NADþ-
binding domain of poly(ADP-ribose) polymerases (PARP).(88)
All proteins involved in processing of the tscA precursor are
associated with the thylakoid membrane. The processed tscA
RNA is also associated with a stromal 1 700 kDa protein
complex that additionally contains the exon 1 primary
transcript with its 50-intron. A component of this protein
complex is Raa3, showing homologies to pyridoxamine 50-
phosphate oxidases. The cofractionation of these two RNAs
together with Raa3 was shown by size exclusion chromato-
graphy.(89) Recently, another factor (Raa4) that shares a small
protein domain with tRNA synthetases was shown to be
involved in splicing of the first group II intron, and it remains to
930
be determined whether it is also part of a high-molecular-
weight complex (Glanz, unpublished).
A biochemical approach including UV-crosslinking experi-
ments, yeast three-hybrid analysis and mass spectrometry
identified three further chloroplast proteins with a more
general affinity to group II introns. These include a 31 kDa
protein with a 39% sequence homology to the NADþ-binding
domain of 6-phosphogluconate dehydrogenases Cpn60, a
bacterial homologue of GroEL ATPases, and a chloroplast-
localised cNAPL protein, showing high similarity to nucleo-
some assembly proteins.(83,90,91)
For splicing of the second intron, apparently two
membrane-associated complexes are involved (Fig. 6). The
first is the 670 kDa complex containing the above-mentioned
Raa1, together with so far uncharacterised RNA molecules
and protein factors. The second is a 400–500 kDa multiprotein
Raa1/Raa2 complex, which is probably not associated with
RNA, since no direct interaction with psaA-i2 or solubilised
chloroplast extracts in vitro was detected.(87) The Raa2
polypeptide contains conserved motifs with significant
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

Glossary
Autocatalytic splicing: Self-splicing of group I, II and III
introns in vitro under non-physiological reaction conditions
in the absence of protein factors. These introns are
referred to as ribozymes (ribonucleic acid enzymes) and
can catalyse their own cleavage or the cleavage of other
RNAs. However, efficient in vivo splicing almost always
requires the assistance of a catalytic enzyme, RNA
molecules and/or other protein factors that are either
encoded by the nucleus or the intron itself (maturases).
Mobile group II introns: Mobile group II introns are
found in bacterial and organelle genomes. They are both
catalytic RNAs and retrotransposable elements with an
intron-encoded protein that has reverse transcriptase
activity. Group II introns can transpose with high
efficiencies (retrohoming) into defined sites or can invade
at ectopic sites (retrotransposition).
Nuclear spliceosome: Nuclear pre-mRNA introns are
not able to splice autocatalytically without the assistance
of trans-acting RNA or protein factors. Eukaryotic pre-
mRNA splicing takes place in the spliceosome, a
ribonucleoprotein (RNP) complex of �60S that assembles
from the five U-rich small nuclear ribonucleoproteins
(snRNPs) U1, U2, U4, U5 and U6, which are temporarily
associated with more than 70 proteins such as RNA
helicases and SR proteins. For accurate spliceosome
assembly, a range of dynamic protein-protein, RNA-
protein and RNA-RNA interactions are required.
Splicing mechanism of group II introns: Splicing
S. Glanz and U. Kuck Review article
sequence similarity to two domains of pseudouridine
synthases; however, this enzyme activity is not a prerequisite
for trans-splicing. Therefore, Raa2 was speculated to be a
bifunctional protein acting in pseudouridination as well as
trans-splicing.(92) Moreover, it was suggested that this
complex represents a pre-spliceosome, which is assembled
and/or stabilised via three genetically defined factors. It was
discussed that this complex has an indirect role in recognition
and assembly of primary exon 2 and 3 RNAs, and thus this
complex may be involved in the storage of trans-splicing
factors. Finally, upon gene activation, this complex may
specifically be redistributed to the site of transcription.(87,92,93)
The spatial separation of the complexes into membrane
and stromal chloroplast fractions indicates that they may act in
different modes and at different steps in the psaA trans-
splicing process. The first reaction probably takes place in the
stromal phase, whereas the second reaction is associated
with the membrane. It can be further speculated that
membranous splicing of the second intron is coupled with
the translation and integration of the psaA protein into the
thylakoid membrane system.(87)
Although no homologues of C. reinhardtii factors have been
identified in higher plant chloroplasts (see Table 3), it may be
envisioned that proteins promoting trans-splicing act as RNA
chaperones and stabilise or support the correct folding of intron
structures. Alternatively, they may mediate splicing indirectly by
interaction with other protein factors. Indeed, cpRNPs in
tobacco were shown to act as stabilising factors for a number of
non-ribosome-bound stromal chloroplast mRNAs.(94) Even
though the exact functions of so far characterised factors in the
trans-splicing process have to be elucidated, the presented
high-molecular RNP complexes and splicing factors provide a
basis for the isolation and characterisation of further trans-
splicing factors and for the analyses of their general functional
role in an organelle spliceosome.
occurs via two sequential transesterification reactions.
First, the 20OH of a specific branch point nucleotide within
the intron performs a nucleophilic attack on the first
nucleotide of the intron at the 50-splice site forming the
lariat intermediate. Second, the 30OH of the released
50-exon performs a nucleophilic attack at the last
nucleotide of the intron at the 30-splice site, thereby joining
the exons and releasing the intron lariat.
Conclusions
Trans-splicing of discontinuous group II introns is a
phenomenon that occurs in a huge number of organelles
from plants and diverse lower eukaryotes. We provide a
complete survey of 187 organelle trans-spliced introns that
were predicted from the complete sequencing data of 179
organelle genomes. Furthermore, a summary of trans-
splicing factors that are supposed to promote group II intron
splicing is given. Genetic and biochemical data from splicing-
deficient mutants support the assumption that intron trans-
splicing is promoted by a set of trans-acting factors as part of
high-molecular-weight complexes. The presented model
predicts that these complexes may be involved in the storage
of trans-splicing factors and can specifically be redistributed
to the site of transcription. A spatial separation of the complex
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
in membrane and non-membrane fractions indicates further
different modes of action during the trans-splicing process.
Supporting Information available online
Acknowledgments: The experimented work of the authors is
supported by the Deutsche Forschungsgemeinschaft
(SFB480, B3).
931

Review article S. Glanz and U. Kuck
References
1. Berget, S. M., Moore, C. and Sharp, P. A., Spliced segments at the 5(terminus of adenovirus 2 late mRNA. Proc Natl Acad Sci USA 1977. 74:
3171–3175.
2. Breathnach, R., Mandel, J. L. and Chambon, P., Ovalbumin gene is
split in chicken DNA. Nature 1977. 270: 314–319.
3. Chow, L. T., Gelinas, R. E., Broker, T. R. and Roberts, R. J.,
An amazing sequence arrangement at the 5( ends of adenovirus 2
messenger RNA. Cell 1977. 12: 1–8.
4. Haugen, P., Simon, D. M. and Bhattacharya, D., The natural history of
group I introns. Trends Genet 2005. 21: 111–119.
5. Cavalier-Smith, T., 1991. Intron phylogeny: a new hypothesis. Trends
Genet 7: 145–148.
6. Michel, F., Umesono, K. and Ozeki, H., Comparative and functional
anatomy of group II catalytic introns - a review. Gene 1989. 82:
5–30.
7. Toro, N., Bacteria and archaea group II introns: additional mobile
genetic elements in the environment. EnvironMicrobiol 2003. 5: 143–151.
8. Valles, Y., Halanych, K. M. and Boore, J. L., Group II introns break new
boundaries: presence in a bilaterian’s genome. PLoS ONE 2008. 3:
e1488.
9. Hastings, K. E. M., SL trans-splicing: easy come or easy go? Trends
Genet 2005. 21: 240–247.
10. Liang, X. H., Haritan, A., Uliel, S. and Michaeli, S., trans and cis
splicing in trypanosomatids: mechanism, factors, and regulation. Eukar-
yotic Cell 2003. 2: 830–840.
11. Stover, N. A., Kaye, M. S. and Cavalcanti, A. R. O., Spliced leader
trans-splicing. Curr Biol 2006. 16: R8–R9.
12. Zhang, H., Hou, Y., Miranda, L., Campbell, D. A., Sturm, N. R., et al.
Spliced leader RNA trans-splicing in dinoflagellates. Proc Natl Acad Sci
USA 2007. 104: 4618–4623.
13. Blumenthal T., 2005. Trans-splicing and operons. WormBook, ed. The
C. elegans Research Community, WormBook, http://www.wormboo-
k.org.
14. Horiuchi, T. and Aigaki, T., Alternative trans-splicing: a novel mode of
pre-mRNA processing. Biol Cell 2006. 98: 135–140.
15. Dorn, R., Reuter, G. and Loewendorf, A., Transgene analysis proves
mRNA trans-splicing at the complex mod(mdg4) locus in Drosophila.
Proc Natl Acad Sci USA 2001. 98: 9724–9729.
16. Pirrotta, V., trans-splicing in Drosophila. BioEssays 2002. 24: 988–991.
17. Toor, N., Hausner, G. and Zimmerly, S., Coevolution of group II intron
RNA structures with their intron-encoded reverse transcriptases. RNA
2001. 7: 1142–1152.
18. Fedorova, O. and Zingler, N., Group II introns: structure, folding and
splicing mechanism. Biol Chem 2007. 388: 665–678.
19. Pyle, A. M., Fedorova, O. and Waldsich, C., Folding of group II introns:
a model system for large, multidomain RNAs? Trends Biochem Sci 2007.
32: 138–145.
20. Valadkhan, S., The spliceosome: a ribozyme at heart? Biol Chem 2007.
388: 693–697.
21. Nilsen, T. W., The spliceosome: the most complex macromolecular
machine in the cell? Bioessays 2003. 25: 1147–1149.
22. Lehmann, K. and Schmidt, U., Group II introns: Structure and catalytic
versatility of large natural ribozymes. Crit Rev Biochem Mol 2003. 38:
249–303.
23. Fromm, H., Edelman, M., Koller, B., Goloubinoff, P. and Galun, E.,
The enigma of the gene coding for ribosomal protein S12 in the chlor-
oplasts of Nicotiana. Nucleic Acids Res 1986. 14: 883–898.
24. Fukuzawa, H., Kohchi, T., Shirai, H., Ohyama, K., Umesono, K., et al.
Coding sequences for chloroplast ribosomal protein S12 from the liver-
wort, Marchantia polymorpha, are separated far apart on the different
DNA strands. FEBS Lett 1986. 198: 11–15.
25. Kuck, U., Choquet, Y., Schneider, M., Dron, M. and Bennoun, P.,
Structural and transcriptional analysis of two homologous genes for
the P700 chlorophyll a-apoproteins in Chlamydomonas reinhardtii:
evidence for in vivo trans-splicing. EMBO J 1987. 6: 2185–2195.
26. Torazawa, K., Hayashida, N., Obokata, J., Shinozaki, K. and Sugiura,
M., The 5( part of the gene for ribosomal protein S12 is located 30 kb
932
downstream from its 3( part in tobacco chloroplast genome. Nucleic
Acids Res 1986. 14: 3143–3143.
27. Hildebrand, M., Hallick, R. B., Passavant, C. W. and Bourque, D. P.,
Trans-splicing in chloroplasts: the rps12 loci of Nicotiana tabacum. Proc
Natl Acad Sci USA 1988. 85: 372–376.
28. Koller, B., Fromm, H., Galun, E. and Edelman, M., Evidence for in vivo
trans-splicing of pre-mRNAs in tobacco chloroplasts. Cell 1987. 48: 111–
119.
29. Belanger, A. S., Brouard, J. S., Charlebois, P., Otis, C., Lemieux, C.
and Turmel, M., Distinctive architecture of the chloroplast genome in the
chlorophycean green alga Stigeoclonium helveticum. Mol Genet Geno-
mics 2006. 276: 464–477.
30. Brouard, J. S., Otis, C., Lemieux, C. and Turmel, M., Chloroplast DNA
sequence of the green alga Oedogonium cardiacum (Chlorophyceae):
unique genome architecture, derived characters shared with the Chae-
tophorales and novel genes acquired through horizontal transfer. BMC
Genomics 2008. 9: 290.
31. de Cambiaire, J. C., Otis, C., Lemieux, C. and Turmel, M., The
complete chloroplast genome sequence of the chlorophycean green
alga Scenedesmus obliquus reveals a compact gene organization and a
biased distribution of genes on the two DNA strands. BMC Evol Biol
2006. 6: 37.
32. Ogihara, Y., Isono, K., Kojima, T., Endo, A., Hanaoka, M., et al.
Structural features of a wheat plastome as revealed by complete
sequencing of chloroplast DNA. Mol Genet Genomics 2002. 266:
740–746.
33. Richaud, C. and Zabulon, G., The heme oxygenase gene (pbsA) in the
red alga Rhodella violacea is discontinuous and transcriptionally acti-
vated during iron limitation. Proc Natl Acad Sci USA 1997. 94: 11736–
11741.
34. Barkan, A. and Goldschmidt-Clermont, M., Participation of nuclear
genes in chloroplast gene expression. Biochimie 2000. 82: 559–
572.
35. Nickelsen, J. and Kuck, U., The unicellular green alga Chlamydomonas
reinhardtii as an experimental system to study chloroplast RNA meta-
bolism. Naturwissenschaften 2000. 87: 97–107.
36. Goldschmidt-Clermont, M., Choquet, Y., Girard-Bascou, J., Michel,
F., Schirmer-Rahire, M. and Rochaix, J. D., A small chloroplast RNA
may be required for trans-splicing in Chlamydomonas reinhardtii. Cell
1991. 65: 135–143.
37. Turmel, M., Choquet, Y., Goldschmidt-Clermont, M., Rochaix, J. D.,
Otis, C. and Lemieux, C., The trans-spliced intron 1 in the psaA gene of
the Chlamydomonas chloroplast: a comparative analysis. Curr Genet
1995. 27: 270–279.
38. Ward, B. L., Anderson, R. S. and Bendich, A. J., The mitochondrial
genome is large and variable in a family of plants (Cucurbitaceae). Cell
1981. 25: 793–803.
39. Burger, G., Forget, L., Zhu, Y., Gray, M. W. and Lang, B. F., Unique
mitochondrial genome architecture in unicellular relatives of animals.
Proc Natl Acad Sci USA 2003. 100: 892–897.
40. Knoop, V., The mitochondrial DNA of land plants: peculiarities in phy-
logenetic perspective. Curr Genet 2004. 46: 123–139.
41. Kubo, T. and Mikami, T., Organization and variation of angiosperm
mitochondrial genome. Physiol Plant 2007. 129: 6–13.
42. Michel, F. and Ferat, J. L., Structure and activities of group II introns.
Annu Rev Biochem 1995. 64: 435–461.
43. Bonen, L., Cis- and trans-splicing of group II introns in plant mitochon-
dria. Mitochondrion 2008. 8: 26–34.
44. Malek, O. and Knoop, V., Trans-splicing group II introns in plant
mitochondria: The complete set of cis-arranged homologs in ferns, fern
allies and a hornwort. RNA 1998. 4: 1599–1609.
45. Chapdelaine, Y. and Bonen, L., The wheat mitochondrial gene for
subunit I of the NADH dehydrogenase complex: a trans-splicing model
for this gene-in-pieces. Cell 1991. 65: 465–472.
46. Conklin, P. L., Wilson, R. K. and Hanson, M. R., Multiple trans-splicing
events are required to produce a mature nad1 transcript in a plant
mitochondrion. Genes Dev 1991. 5: 1407–1415.
47. Wissinger, B., Schuster, W. and Brennicke, A., Trans-splicing in
Oenothera mitochondria: nad1 mRNAs are edited in exon and trans-
splicing group II intron sequences. Cell 1991. 65: 473–482.
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.

S. Glanz and U. Kuck Review article
48. Binder, S., Marchfelder, A., Brennicke, A. and Wissinger, B., RNA
editing in trans-splicing intron sequences of nad2 mRNAs in Oenothera
mitochondria. J Biol Chem 1992. 267: 7615–7623.
49. Knoop, V., Schuster, W., Wissinger, B. and Brennicke, A., Trans-
splicing integrates an exon of 22 nucleotides into the nad5 mRNA in
higher plant mitochondria. EMBO J 1991. 10: 3483–3493.
50. Chaw, S. M., Shih, A. C. C., Wang, D., Wu, Y. W., Liu, S. M. and Chou,
T. Y., The mitochondrial genome of the gymnosperm Cycas taitungensis
contains a novel family of short interspersed elements, Bpu sequences,
and abundant RNA editing sites. Mol Biol Evol 2008. 25: 603–615.
51. Morawala-Patell, V., Gualberto, J. M., Lamattina, L., Grienenberger,
J. M. and Bonnard, G., Cis- and trans-splicing and RNA editing are
required for the expression of nad2 in wheat mitochondria. Mol Gen
Genet 1998. 258: 503–511.
52. Knoop, V., Altwasser, M. and Brennicke, A., A tripartite group II intron
in mitochondria of an angiosperm plant. Mol Gen Genet 1997. 255: 269–
276.
53. Waller, R. F. and Jackson, C. J., Dinoflagellate mitochondrial genomes:
stretching the rules of molecular biology. Bioessays 2009. 31: 237–245.
54. Marande, W., Lukes, J. and Burger, G., Unique mitochondrial genome
structure in diplonemids, the sister group of kinetoplastids. Eukaryotic
Cell 2005. 4: 1137–1146.
55. Marande, W. and Burger, G., Mitochondrial DNA as a genomic jigsaw
puzzle. Science 2007. 318: 415.
56. Nash, E. A., Nisbet, R. E., Barbrook, A. C. and Howe, C. J., Dino-
flagellates: a mitochondrial genome all at sea. Trends Genet 2008. 24:
328–335.
57. Bonen, L. and Vogel, J., The ins and outs of group II introns. Trends
Genet 2001. 17: 322–331.
58. Nickelsen, J., Chloroplast RNA-binding proteins. Curr Genet 2003. 43:
392–399.
59. Dai, L. X., Chai, D. G., Gu, S. Q., Gabel, J., Noskov, S. Y., et al. A three-
dimensional model of a group II intron RNA and its interaction with the
intron-encoded reverse transcriptase. Mol Cell 2008. 30: 472–485.
60. Rambo, R. P. and Doudna, J. A., Assembly of an active group II intron-
maturase complex by protein dimerization.Biochemistry 2004. 43: 6486–
6497.
61. Kelchner, S. A., Group II introns as phylogenetic tools: structure func-
tion, and evolutionary constraints. Am J Bot 2002. 89: 1651–1669.
62. Toro, N., Jimenez-Zurdo, J. I. and Garcıa-Rodrıguez, F. M., Bacterial
group II introns: not just splicing. FEMSMicrobiol Rev 2007. 31: 342–358.
63. Belhocine, K., Mak, A. B. and Cousineau, B., Trans-splicing of the
Ll.LtrB group II intron in Lactococcus lactis. Nucleic Acids Res 2007. 35:
2257–2268.
64. Belhocine, K., Mak, A. B. and Cousineau, B., Trans-splicing versatility
of the LI.LtrB group II intron. RNA 2008. 14: 1782–1790.
65. Bhattacharya, D., Archibald, J. M., Weber, A. P. M. and Reyes-Prieto,
A., How do endosymbionts become organelles? Understanding early
events in plastid evolution. Bioessays 2007. 29: 1239–1246.
66. Gould, S. B., Waller, R. R. and McFadden, G. I., Plastid evolution. Annu
Rev Plant Biol 2008. 59: 491–517.
67. Kleine, T., Maier, U. G. and Leister, D., DNA transfer from organelles to
the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant
Biol 2008. 60: 115–138.
68. Bock, R. and Timmis, J. N., Reconstructing evolution: gene transfer
from plastids to the nucleus. Bioessays 2008. 30: 556–566.
69. Patron, N. J. and Waller, R. F., Transit peptide diversity and divergence:
a global analysis of plastid targeting signals. Bioessays 2007. 29: 1048–
1058.
70. Mohr, G. and Lambowitz, A. M., Putative proteins related to group II
intron reverse transcriptase/maturases are encoded by nuclear genes in
higher plants. Nucleic Acids Res 2003. 31: 647–652.
71. Nakagawa, N. and Sakurai, N., A mutation in At-nMat1a, which encodes
a nuclear gene having high similarity to group II intron maturase, causes
impaired splicing of mitochondrial NAD4 transcript and altered carbon
metabolism in Arabidopsis thaliana. Plant Cell Physiol 2006. 47: 772–
783.
72. Tillich, M., Poltnigg, P., Kushnir, S. and Schmitz-Linneweber, C.,
Maintenance of plastid RNA editing activities independently of their
target sites. EMBO Rep 2006. 7: 308–313.
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.
73. Wissinger, B., Brennicke, A. and Schuster, W., Regenerating good
sense - RNA editing and trans-splicing in plant mitochondria. Trends
Genet 1992. 8: 322–328.
74. Borner, G. V., Morl, M., Wissinger, B., Brennicke, A. and Schmelzer,
C., RNA editing of a group II intron in Oenothera as a prerequisite for
splicing. Mol Gen Genet 1995. 246: 739–744.
75. Mingler, M. K., Hingst, A. M., Clement, S. L., Yu, L. E., Reifur, L. and
Koslowsky, D. J., Identification of pentatricopeptide repeat proteins in
Trypanosoma brucei. Mol Biochem Parasitol 2006. 150: 37–45.
76. Lurin, C., Andres, C., Aubourg, S., Bellaoui, M., Bitton, F., et al.
Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins
reveals their essential role in organelle biogenesis. Plant Cell 2004. 16:
2089–2103.
77. Delannoy, E., Stanley, W. A., Bond, C. S. and Small, I. D., Pentatri-
copeptide repeat (PPR) proteins as sequence-specificity factors in post-
transcriptional processes in organelles. Biochem Soc Trans 2007. 35:
1643–1647.
78. de Longevialle, A. F., Meyer, E. H., Andres, C., Taylor, N. L., Lurin, C.,
et al. The pentatricopeptide repeat gene OTP43 is required for trans-
splicing of the mitochondrial nad1 intron 1 in Arabidopsis thaliana. Plant
Cell 2007. 19: 3256–3265.
79. Schmitz-Linneweber, C., Williams-Carrier, R. E., Williams-Voelker,
P. M., Kroeger, T. S., Vichas, A. and Barkan, A., A pentatricopeptide
repeat protein facilitates the trans-splicing of the maize chloroplast rps12
pre-mRNA. Plant Cell 2006. 18: 2650–2663.
80. Merendino, L., Perron, K., Rahire, M., Howald, I., Rochaix, J. D. and
Goldschmidt-Clermont, M., A novel multifunctional factor involved in
trans-splicing of chloroplast introns in Chlamydomonas. Nucleic Acids
Res 2006. 34: 262–274.
81. Barkan, A., Klipcan, L., Ostersetzer, O., Kawamura, T., Asakura, Y.
and Watkins, K. P., The CRM domain: An RNA binding module derived
from an ancient ribosome-associated protein. RNA 2007. 13: 55–64.
82. Ostheimer, G. J., Rojas, M., Hadjivassiliou, H. and Barkan, A., For-
mation of the CRS2-CAF2 group II intron splicing complex is mediated by
a 22-amino acid motif in the COOH-terminal region of CAF2. J Biol Chem
2006. 281: 4732–4738.
83. Bunse, A. A., Nickelsen, J. and Kuck, U., Intron-specific RNA binding
proteins in the chloroplast of the green alga Chlamydomonas reinhardtii.
Biochim Biophys Acta 2001. 1519: 46–54.
84. Goldschmidt-Clermont, M., Girard-Bascou, J., Choquet, Y. and
Rochaix, J. D., Trans-splicing mutants of Chlamydomonas reinhardtii.
Mol Gen Genet 1990. 223: 417–425.
85. Choquet, Y., Goldschmidt-Clermont, M., Girard-Bascou, J., Kuck, U.,
Bennoun, P. and Rochaix, J. D., Mutant phenotypes support a trans-
splicing mechanism for the expression of the tripartite psaA gene in the
C. reinhardtii chloroplast. Cell 1988. 52: 903–913.
86. Hahn, D., Nickelsen, J., Hackert, A. and Kuck, U., A single nuclear
locus is involved in both chloroplast RNA trans-splicing and 3( end
processing. Plant J 1998. 15: 575–581.
87. Perron, K., Goldschmidt-Clermont, M. and Rochaix, J. D.,
A multiprotein complex involved in chloroplast group II intron splicing.
RNA 2004. 10: 704–711.
88. Balczun, C., Bunse, A., Hahn, D., Bennoun, P., Nickelsen, J. and
Kuck, U., Two adjacent nuclear genes are required for functional
complementation of a chloroplast trans-splicing mutant from Chlamydo-
monas reinhardtii. Plant J 2005. 43: 636–648.
89. Rivier, C., Goldschmidt-Clermont, M. and Rochaix, J. D., Identifica-
tion of an RNA-protein complex involved in chloroplast group II intron
trans-splicing in Chlamydomonas reinhardtii. EMBO J 2001. 20: 1765–
1773.
90. Balczun, C., Bunse, A., Schwarz, C., Piotrowski, M. and Kuck, U.,
Chloroplast heat shock protein Cpn60 from Chlamydomonas reinhardtii
exhibits a novel function as a group II intron-specific RNA-binding
protein. FEBS Lett 2006. 580: 4527–4532.
91. Glanz, S., Bunse, A., Wimbert, A., Balczun, C. and Kuck, U.,
A nucleosome assembly protein-like polypeptide binds to chloroplast
group II intron RNA in Chlamydomonas reinhardtii. Nucleic Acids Res
2006. 34: 5337–5351.
92. Perron, K., Goldschmidt-Clermont, M. and Rochaix, J. D., A factor
related to pseudouridine synthases is required for chloroplast group II
933

Review article S. Glanz and U. Kuck
intron trans-splicing in Chlamydomonas reinhardtii. EMBO J 1999. 18:
6481–6490.
93. Zerges, W. and Rochaix, J. D., Low density membranes are associated
with RNA-binding proteins and thylakoids in the chloroplast of Chlamy-
domonas reinhardtii. J Cell Biol 1998. 140: 101–110.
94. Nakamura, T., Ohta, M., Sugiura, M. and Sugita, M., Chloroplast
ribonucleoproteins function as a stabilizing factor of ribosome-free
mRNAs in the stroma. J Biol Chem 2001. 276: 147–152.
95. Pyle, A. M. and Lambowitz, A. M., Group II introns: ribozymes that
splice RNA and invade DNA. In: Gestel, R. F., Cech, T. R. and Atkins, J. F.
editors. The RNA World, 3rd ed. Cold Spring Harbor, NY, Cold Spring
Harbor Laboratory Press, 2006. p 469–506.
96. Farre, J. C. and Araya, A., The mat-r open reading frame is transcribed
from a non-canonical promoter and contains an internal promoter to co-
transcribe exons nad1e and nad5III in wheat mitochondria. Plant Mol Biol
1999. 40: 959–967.
97. Ogihara, Y., Isono, K., Kojima, T., Endo, A., Hanaoka, M., et al.
Chinese spring wheat (Triticum aestivum L.) chloroplast genome: com-
plete sequence and contig clones. Plant Mol Biol Rep 2000. 18: 243–253.
98. Hubschmann, T., Hess, W. R. and Borner, T., Impaired splicing of the
rps12 transcript in ribosome-deficient plastids. Plant Mol Biol 1996. 30:
109–123.
99. Ems, S. C., Morden, C. W., Dixon, C. K., Wolfe, K. H., dePamphilis,
C. W. and Palmer, J. D., Transcription, splicing and editing of plastid
RNAs in the nonphotosynthetic plant Epifagus virginiana. Plant Mol Biol
1995. 29: 721–733.
100. Turmel, M., Otis, C. and Lemieux, C., The complete chloroplast DNA
sequences of the charophycean green algae Staurastrum and Zygnema
reveal that the chloroplast genome underwent extensive changes during
the evolution of the Zygnematales. BMC Biol 2005. 3: 22.
101. Freyer, R., Neckermann, K., Maier, R. M. and Kossel, H., Structural
and functional analysis of plastid genomes from parasitic plants: loss of
an intron within the genus Cuscuta. Curr Genet 1995. 27: 580–586.
102. Sanchez Puerta, M. V., Bachvaroff, T. R. and Delwiche, C. F., The
complete mitochondrial genome sequence of the haptophyte Emiliania
huxleyi and its relation to heterokonts. DNA Res 2004. 11: 1–10.
934
103. Jackson, C. J., Norman, J. E., Schnare, M. N., Gray, M. W., Keeling,
P. J. and Waller, R. F., Broad genomic and transcriptional analysis
reveals a highly derived genome in dinoflagellate mitochondria. BMC
Biol 2007. 5: 41.
104. Clifton, S. W., Minx, P., Fauron, C. M. R., Gibson, M., Allen, J. O., et al.
Sequence and comparative analysis of the maize NB mitochondrial
genome. Plant Physiol 2004. 136: 3486–3503.
105. Handa, H., The complete nucleotide sequence and RNA editing content
of the mitochondrial genome of rapeseed (Brassica napus L.): compara-
tive analysis of the mitochondrial genomes of rapeseed and Arabidopsis
thaliana. Nucleic Acids Res 2003. 31: 5907–5916.
106. Unseld, M., Marienfeld, J. R., Brandt, P. and Brennicke, A., The
mitochondrial genome of Arabidopsis thaliana contains 57 genes in
366,924 nucleotides. Nat Genet 1997. 15: 57–61.
107. Wahleithner, J. A., Macfarlane, J. L. and Wolstenholme, D. R.,
A sequence encoding a maturase-related protein in a group II intron
of a plant mitochondrial nad1 gene. Proc Natl Acad Sci USA 1990. 87:
548–552.
108. Handa, H., Mizobuchi-Fukuoka, R. and Pinyarat, W., The rapeseed
mitochondrial gene for subunit 2 of the NADH dehydrogenase
complex: a trans-spliced structure is conserved in one of the smallest
plant mitochondrial genomes. Curr Genet 1997. 31: 336–342.
109. Lippok, B., Brennicke, A. and Unseld, M., The rps4 gene is encoded
upstream of the nad2 gene in Arabidopsis mitochondria. Biol Chem
Hoppe Seyler 1996. 377: 251–257.
110. Turmel, M., Otis, C. and Lemieux, C., The complete mitochondrial DNA
sequence of Mesostigma viride identifies this green alga as the earliest
green plant divergence and predicts a highly compact mitochondrial
genome in the ancestor of all green plants. Mol Biol Evol 2002. 19: 24–38.
111. deSouza, A. P., Jubier, M. F., Delcher, E., Lancelin, D. and Lejeune,
B., A trans-splicing model for the expression of the tripartite nad5 gene in
wheat and maize mitochondria. Plant Cell 1991. 3: 1363–1378.
112. Scheepers, D., Luo, H. and Boutry, M., Variant mitochondrial tran-
scripts of a broad bean line are associated with two point mutations
located upstream of the nad5 exon c. Plant Sci 1997. 129: 203–
212.
BioEssays 31:921–934, � 2009 Wiley Periodicals, Inc.