Tombo: detection of non-standard nucleotides using the genome … · 2018-05-21 · Tombo:...
Transcript of Tombo: detection of non-standard nucleotides using the genome … · 2018-05-21 · Tombo:...

Tombo: detection of non-standard nucleotides using the genome-resolved raw nanopore signalThe Tombo software package enables investigation, detection and visualisation of modified nucleotides. Its framework enables practical, scalable expansion to detect all modifications in both DNA and RNA
Fig. 1 Modifications a) different epigenetic modifications b) nanopore sequencing of native RNA
‘Epigenetics’ refers to heritable alterations of DNA that do not change the nucleotide sequence. One of the most widespread epigenetic modifications is 5-methyl cytosine (m5C), which most frequently occurs in mammalian cells in CpG dinucleotides, but which also occurs in additional sequence contexts. CpG methylation can alter patterns of gene expression by suppressing transcription. Base modification is also widespread in RNA, though the roles effected by these changes in biological processes are less well understood. Nanopore sequencing does not require amplification or strand synthesis, meaning that during sequencing, modified bases pass through the pore, and that the signature of these bases is present in the raw signal (Fig. 1).
Tombo provides three distinct methods for the detection of modified bases, the choice of which depends on the data available and the experimental objectives. The performance of the different models has continued to improve as we have developed Tombo’s algorithms. Figure 2a shows ROC curves for the three detection methods at the dam- and dcm-modified motifs in E. coli. The de novo method (identifying deviations from the canonical base model) shows the best performance. dam- and dcm-methylation show strong and consistent shifts in the raw nanopore signal (Fig. 2b, top panel). Across the top 1,000 CCWGG-containing regions, the fraction of modified bases identified by Tombo is highest at the known m5C location (Fig. 2c, bottom panel).
Non-standard nucleotides are biologically significant and widespread in DNA and RNA
Tombo performance on known m5C and N6-methyl A (m6A) sites in the E. coli genome
Fig. 2 Identifying m6A and m5C a) ROC curves b) AUC values c) examples of dcm-methylation
Fig. 3 a) ROC curve for detection of m6A and m5C in E. coli gDNA at different levels of coverage b) m5C detection on NA12878 chr20
We applied Tombo’s de novo modified base model to the E. coli genome for detection of m6A and m5C at different levels of coverage: 1x, 30x and 376x. As might be expected, greater coverage led to higher AUCs. At 376x, the AUC for m6A is 0.975, and for m5C is 0.992, whereas at 30x the AUCs are 0.947 and 0.983 respectively (Fig. 3a). We then used Tombo to detect m5C in human genomic data generated on a PromethION from NA12878, genomic DNA. The optimisation of Tombo for use with PromethION data is not yet complete, but we obtained strong correspondence between our Tombo analysis and publicly available bisulphite data from the same genome. Fig. 3b shows raw nanopore signals (red lines) which deviate from the expected canonical levels (grey background distributions) around sites of methylation which were identified using bisulphite sequencing. The top two panels show methylation at a CpG site, which is symmetric on positive and negative strands. The bottom two panels show methylation in examples of CHG and CHH contexts. Here the methylation was asymmetric, only being present on the positive strands.
To estimate a specific m5C model for RNA, we produced a library by in vitro transcription which contains a mixture of standard NTPs and m5CTP. The modified base caused a signal-level shift only at positions containing cytosine bases, compared to a control library (Fig. 4). From these signal distributions, we created a m5C RNA model and this is now included with Tombo for the specific detection of m5C in Direct RNA sequencing experiments.
Using Tombo to detect of m6A and m5C at different levels of coverage, and m5C in a variety of sequence contexts
Model estimation for m5C in Direct RNA data
Fig. 4 Signal shifts in non-C- and C-containing motifs
© 2018 Oxford Nanopore Technologies. All rights reserved.P17020 - Version 5.0
b)
Contact: [email protected] More information at: www.nanoporetech.com and publications.nanoporetech.com
a)
CH3
CH3CO
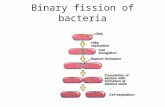
Chromosome
Chromatin
Nucleosome
Histonemodification
DNAmodification
Alteration of gene expression
Cur
rent
(pA
)
100
90
80
70
60
Position in reference210 220 230
CUGUCGAAUUAAUUCGCCCGGCGAAUGUGCC
Unmodifiedm6A
m6A
Exa
mpl
es o
fm
ethy
late
d lo
ciFr
actio
n m
odifi
ed
0.0
0.2
0.4
0.6
0.8
1.0
A C C T T T A G C T G G G C A A A A A A
C C G C T A C A C m5C
m5C
T G G A A T T C T A C
a)
b)
0
0.25
0.50
0.75
1.00
0 0.25 0.50 0.75 1.00False positive rate
True
pos
itive
rat
ec)
Modificationm5Cm6A
ModelAlternativeDe novoPCR comparison
420X
1X
Model
Mod m5C m6A
b)a)
0
0.25
0.50
0.75
1.00
0 0.25 0.50 0.75 1.00False positive rate
True
pos
itive
rat
e
Modificationm5Cm6A
Coverage1x30x376x
Amplified with 25% m5CAmplified without m5C
−2.5 0.0 2.5Signal
−2.5 0.0 2.5Signal
AAAUU
AGAAA
AGAAG
AGAAU
AGAGA
AGAGU
AGAUA
AGAUG
AGAUU
GAGAU
GAGUA
GGAAU
GGAUA
GGAUG
GGAUU
UGAAA
UGAAU
UGAUA
UGAUG
UGAUU
5mer
s
Largest shifts in non-C-containing 5mers
ACCCU
AGAAC
AGACA
AGACC
AGACG
AGACU
AGAUC
AGCCC
AGUAC
AGUCC
AGUCG
AGUCU
CCCCC
CGACA
CGACC
GGACG
GGACU
UGACA
UGACC
UGACG
Largest shifts inC-containing 5mers
0.6
0.7
0.8
0.9
AUC0.72 0.81 0.76 0.65 0.74 0.61
0.89 0.99 0.85 0.88 0.98 0.60
CHH motifchromosome 20:
position 1,333,620+ve strand
T A C T A C C C G T A A G G G C T G G Cm5C
CHG motifchromosome 20:position 404,407
+ve strandG T T T G T A T A T A G T A T T A T C Am5C
CpG motifchromosome 20:position 457,495
+ve strandG T G C T A G G A T G C A G A T G T G Am5C
CpG motifchromosome 20:position 457,496
-ve strand
C A C G A T C C T A G G T C T A C A C Tm5C
Locationof m5C
m5C-containingmotif
Bisulphite data taken from the Encode Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489 (7414) 57–74 (2012).