The Blueprint of Life, From DNA to Protein Chapter 7.
-
Upload
lawrence-farrin -
Category
Documents
-
view
218 -
download
0
Transcript of The Blueprint of Life, From DNA to Protein Chapter 7.

The Blueprint of Life,From DNA to Protein
Chapter 7

The Blueprint of Life
Characteristics of each cell dictated by information contained on DNA DNA holds master blueprint
All cell structures and processes directed by DNA

Overview
Complete set of genetic information referred to as genome Genome of all cells is composed of DNA
Some viruses have RNA genome Functional unit of genome is the gene Gene codes for gene product
Gene product is most commonly protein Study of transfer of genes is genetics Study of sequence of DNA is genomic

Overview
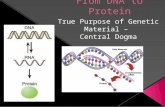
Living cells must accomplish two general tasks to multiply DNA replication DNA expression (gene expression)
Expression involves two process Transcription
Copies information in DNA to RNA Translation
Interpret RNA to synthesize protein
Flow of information from DNA to RNA to protein Central dogma of molecular biology

Overview
Characteristics of DNA Made up of deoxy-
ribonucleotides Nucleotides include:
Phosphate group 5 carbon sugar
Deoxyribose Nucleotides bond covalently
between the 5’PO4 of one nucleotide and the 3’OH of another
Joining of nucleotides creates an alternating sugar-phosphate backbone

Overview
Characteristics of DNA Each sugar (deoxyribose) molecule is
connected to a nitrogenous base Nitrogenous bases
Adenine (A) - purine Thymine (T) - pyrimidine Guanine (G) - purine Cytosine (C) – pyrimidine

Overview
Characteristics of DNA Chemical structure and joining of
nucleotide subunits causes strands to differ at the ends
One strand has a phosphate attached at the number 5 carbon of the sugar.
Termed the five prime (5’) end The other strand has a hydroxyl
group attached to the number 3 carbon of the sugar.
Termed the three prime (3’) end

Overview Characteristics of DNA
DNA occurs as double-stranded molecule
Strands are complementary to each other
Due to the specific base pairing of bases
A:T C:G
Strands are held together with hydrogen bonds
Specific hydrogen bonding between bases
A is bound to T by two hydrogen bonds
G is bound to C by three hydrogen bond

Overview
Characteristics of DNA DNA molecule is antiparallel
Strands are oriented in opposite directions
Strands differ at the ends One strand oriented in
the 5’ to 3’ direction. The other strand is
oriented in the 3’ to 5’ direction.

Overview
Characteristics of RNA RNA is made up of nucleotides
Ribonucleotides RNA contains nitrogenous bases
Adenine Guanine Cytosine Uracil
Uracil replaces thymine in RNA
RNA usually exists as single stranded molecule

Overview
Characteristics of RNA Portion of DNA acts of template for RNA
synthesis RNA molecule called transcript
Numerous transcripts can be produced from one chromosome
Either strand of DNA can act as template Three different functional groups of RNA
Messenger RNA (mRNA) Ribosomal RNA (rRNA) Transfer RNA (tRNA)

Overview
Regulating the expression of genes Nucleotide sequence codes for regulation
mechanism for gene expression Mechanisms determine duration of synthesis
of gene products Products are only made when required
Key mechanism is regulation of mRNA synthesis from DNA
Regulation of transcription

DNA Replication
DNA is replicated to create second copy of molecule Molecule is identical to
original Replication is
bidirectional Replication begins at
specific starting point Proceeds in opposite
directions Allows replication to
proceed more quickly Bi-directional repli
cation

DNA Replication
DNA replication The two strands are unwound and separated Free, unbound nucleotides match up to the
newly separated nitrogenous bases of the parent strand
The parent strand is also called the template strand

DNA Replication
DNA replication Base pairing is specific in DNA replication
Where adenine is present only thymine binds in the new strand and vice versa
Where guanine is present only cytosine binds in the new strand and vice versa
Bases that are improperly inserted are removed and replaced with the correct base
Newly added bases are added by the enzyme DNA polymerase

DNA Replication
Specifics of DNA replication As the strands of DNA
unwind, it creates an area of replication called the replication fork
As nucleotides are added, the replication fork moves down the parental strand

DNA Replication
Specifics of DNA replication DNA polymerase adds new nucleotides as
they become available. DNA polymerase can only add nucleotides to the
free hydroxyl at the 3’ end DNA polymerase replicates in 5’ to 3’ direction Enzymes READS DNA template in 3’ to 5’ direction
Because of the antiparallel nature of the strands of DNA, the two new strands will grow in opposite directions
One strand is the leading strand One strand is the lagging strand

DNA Replication
Specifics of DNA replication Leading strand
Is synthesized CONTINUOUSLY as the DNA polymerase moves towards the replication fork
Lagging strand Is synthesized DISCONTINUOUSLY in pieces as
DNA polymerase moves away from the replication fork

DNA Replication
Specifics of DNA replication DNA polymerase must bind to an RNA primer
to begin synthesis A second DNA polymerase removes any RNA
primers An RNA primer is required at each newly
synthesized section of the lagging strand DNA ligase joins the fragments of the lagging
strand

DNA Replication
Specifics of DNA replication Replication is completed when the replication
fork reaches the end of the parent strands The original parent strand and the newly
synthesized daughter strand rewind Each new strand of DNA consists of one parent
strand and one daughter strand DNA replication is referred to as semiconservative
DNA Replication

Gene Expression
Involves two separate but interrelated process Transcription
Process of synthesizing RNA from DNA template Translation
RNA is deciphered to synthesize protein

Gene Expression
Transcription Transcription is the synthesis of a strand of
mRNA from a DNA template mRNA carries the coded information from DNA to
the ribosome, which is the site of protein synthesis
mRNA also plays an important role in translation

Gene Expression
Transcription During transcription the
enzyme, RNA polymerase, synthesizes a complementary strand of mRNA from a portion of unwound DNA

Gene Expression
Specifics of Transcription RNA polymerase binds to a region of the DNA
called the promoter Only one strand of DNA acts as a template
This is called the sense strand The strand not transcribed is the nonsense strand

Genet Expression
Specifics of transcription Nucleotides in RNA are the same as those in
DNA with one exception Thymine is replaced with uracil
Binding in RNA is A:U or U:A C:G or G:C

Gene Expression
Specifics of transcription RNA polymerase continues down strand of
DNA until it reaches a site on DNA called the terminator
At the terminator RNA polymerase and the new strand of mRNA are released from strand of DNA
Transcription

Gene Expression
Translation Translation is the decoding of information held
in the mRNA to synthesize proteins Two more RNA molecules become involved in
translation Ribosomal RNA (rRNA) Transfer RNA (tRNA)

Gene Expression
rRNA forms part of the ribosomal machinery used in protein synthesis rRNA builds the ribosomes
tRNA recognizes specific sequences of mRNA and transports the required amino acids to form a polypeptide chain

Gene Expression
Translation The language of mRNA is in the form of
codons Codons are groups of three nucleotides situated
next to each other on DNA Codons are written in terms of their base
sequence in mRNA The sequence of codons determines the
sequence of amino acids in the protein

Gene Expression
Translation There are 64 codons that make up the
“alphabet” of proteins Of the 64 codons, 61 are sense codons
Each coding a specific amino acid The remaining 3 are nonsense codons
These code for termination of the message
Codons contained in mRNA are read into proteins through translation
The site of translation is the ribosome

Gene Expression
In response to each codon, tRNA brings the appropriate amino acid to the site of translation
Each codon has an anticodon The anticodon is
complementary sequence to the codon

Gene Expression
Translation Ribosomes
The 30s and the 50s ribosomal subunits join together around the mRNA
The ribosomes direct the binding of tRNA to the correct codon on the mRNA
tRNA binds to the P site and the A site of the 50s ribosomal subunit
The ribosomes bind to the mRNA to be translated

Gene Expression
Specifics of Translation The first tRNA binds to a start
codon in the P site of the ribosome
AUG is the start codon for EVERY protein
AUG codes for the amino acid methionine
When the second tRNA binds to the A site, the amino acid of the first tRNA forms a peptide bond with the amino acid of the second tRNA
P site A site

Gene Expression
Specifics of translation After the peptide bond is formed between the
two amino acids, the tRNA P site leaves the ribosome
The ribosome moves distance of one codon Amino acid in the A site moves to the P site A new tRNA fills the now empty A site

Gene Expression
Specifics of translation The ribosome continues down the strand of
mRNA Amino acids form peptide bonds along the way
Translation is terminated when the ribosomes come to a stop or nonsense codon
At this point the ribosomes separate The new polypeptide chain is released
The ribosome and the mRNA are free to begin translation again

Gene Expression
Specifics of translation As the ribosome moves down the strand of
mRNA, the start codon is exposed Once exposed, a new ribosome will attach and
begin another polypeptide chain
Translation

Regulation of Gene Expression
Microorganisms posses mechanism to synthesize maximum amount of cell material from limited energy Controls directed at metabolic pathways
Two general mechanism Allosteric inhibition of enzymes Controlling synthesis of enzymes
Directed at making only what is required

Principles of regulation Not all genes subjected to regulation Enzymes can be classified according to characteristics of
regulation Constitutive enzymes
Constantly synthesized Enzymes of glycolysis
Inducible enzymes Not regularly produced turned on in certain conditions
Β-galactosidase Repressible enzymes
Routinely synthesized Generally involved in biosynthesis
Regulation of Gene Expression

Mechanisms controlling transcription Often controlled by regulatory region near
promoter Protein binds to region and acts as “on/off” switch
Binding protein can act as repressor or activator Repressor blocks transcription Activator facilitates transcription
Set of genes controlled by protein is called an operon
Regulation of Gene Expression

Regulation of Gene Expression
Repressors Control mechanism that inhibits gene expression and
decreases the synthesis of enzymes Repression is usually in response to the overabundance
of an end product Repression decreases the rate synthesis of enzymes
leading to the formation of the particular end product Regulatory proteins called repressors mediate
repression Repressors block the ability of RNA polymerase to bind and
initiate protein synthesis

Regulation of Gene Expression
Activators Control mechanism that turns on the
transcription of a gene or set of genes Inducers are substances that act to induce
transcription Enzymes synthesized in the presence of inducers
are called inducible enzymes

Regulation of Gene Expression
Operon model of gene expression An operon is a set of genes that includes an
operator, promoter and structural genes An operon is divided into two regions, the control
region and the structural region The control region include the operator and the
promoter This region controls transcription The operator acts as the “on-off” switch
The structural region includes the structural genes This region contains the genes being transcribed

Operon structure
Promoter – Binding site for RNA polymerase
Operator – binding site for the repressor protein for the regulation of gene expression
Structural Genes – DNA sequence for specific proteins
Operator
Gene 1 Gene 3Gene 2
Promoter

Regulation of Gene Expression
Lac operon Example of induction of gene expression
Near the operon on the DNA is a regulatory gene called the “I” gene
This codes for the repressor protein When lactose is absent, the repressor protein
binds to the operator gene Binding of the repressor gene prevents RNA
polymerase from transcribing the structural genes No mRNA is made and no enzymes are
synthesized

Regulation of Gene Expression
Lac operon When lactose is present the repressor binds to
lactose instead of the operator With the repressor bound to lactose, RNA
polymerase is able to bind to the promoter and transcribes the structural genes
Lactose acts as an inducer by keeping the repressor from binding to the operator
It induces the transcription of the structural genes

Lac Operon
Operator
Gene 1 Gene 3Gene 2
Promoter
1.
2.
Lactose
3.
RNA polymerase
Repressor
Lac Operon

Gene Expression and Environmental Fluctuations Many organisms adapt to changing
environments by altering level of gene expression
Mechanisms include Signal transduction Natural selection

Signal transduction Process that transmits information from external
environment to inside cell Allows cell to respond to changes
Two-component regulatory systems Relies on sensor and response regulator proteins
Sensors recognize change in environment Response regulators activate or repress gene
expression Quorum sensing
Organisms sense density of population Enables activation of genes beneficial to the mass
Gene Expression and Environmental Fluctuations

Natural selection Mechanisms to enhance survivability
Antigenic variation Alteration in characteristics of certain surface
proteins Example: Neisseria gonorrhoeae hides from host
immunity by changing numerous surface proteins
Phase variation Routine switching on and off of certain genes
Altering expression allows portions of population to survive and multiply
Gene Expression and Environmental Fluctuations