Electronic Supplementary Material Supplementary Figures S1 ...
Supplementary Information Spatial segregation of polarity factors … · 2013-05-14 ·...
Transcript of Supplementary Information Spatial segregation of polarity factors … · 2013-05-14 ·...

Supplementary Information
Spatial segregation of polarity factors into distinct cortical clusters is required
for cell polarity control
James Dodgson 1*
, Anatole Chessel 1*
, Miki Yamamoto 2, Federico Vaggi
3, Susan Cox
4, Edward
Rosten 5, David Albrecht
6, Marco Geymonat
1, Attila Csikasz-Nagy
3, Masamitsu Sato
2, Rafael
E. Carazo-Salas 1, 6
1The Gurdon Institute, University of Cambridge, Cambridge, CB2 1QN, United Kingdom.
2Department of Biophysics and Biochemistry, University of Tokyo, Tokyo, 113-0032, Japan.
3The
Microsoft Research-University of Trento Centre for Computational Systems Biology, Piazza
Manci, 17, Trento, 38123, Italy. 4Randall Division of Cell and Molecular Biophysics, New Hunt's
House, King's College London, London, SE1 1UL. 5Department of Engineering, University of
Cambridge, Cambridge CB2 1PZ, UK. 6Institute of Biochemistry, ETH Zurich, Schafmattstrasse
18, HPM G16.2, Zurich, CH-8093, Switzerland.
* These authors contributed equally to this work
Correspondence: Email: [email protected] (R.E.C.S.), phone: +44 1223 334130, fax:
+441223334089.
Supplementary Figures S1 to S8
Supplementary Table S1
Supplementary Methods
Supplementary References

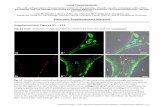
Supplementary Figure S1.
a. Tea1, Tea3 and Tea4 tagged with a monomeric GFP variant form nodes that are
indistinguishable from those tagged with the conventional S65T variant GFP. Images are
maximum intensity projections of 3D-deconvolved, wide-field z-stacks Scale bar: 2m. b.
Distribution of cluster sizes measured with dSTORM (FWHM, Full Width at Half Maximum; n =
83 clusters).

Supplementary Figure S2. Polarity factor cluster formation both in silico and in vivo is not
an inevitable consequence of polarity factor oligomerization capacity.
a. In silico T: a simulated oligomerizing polarity factor delivered by microtubule-mimicking
events onto a 2D hemispherical cell cortex. M: a cortically constrained, 2D diffusing protein that
anchors T. 48 different simulations were computed by varying the dissociation rate of T (Z axis),
the interaction radius between pairs of T particles (X axis) and the interaction radius between T
and M particle pairs (Y axis). All axes are in a –log scale. Spot size reflects the mean size of
clusters in pixels. Spot colour denotes the mean number of clusters formed in the simulated cell
end. As can be seen from the XYZ plot, a range of cluster sizes can be obtained - from many small
clusters (equivalent to a non-oligomerizing protein) to one big cluster encompassing most of the
simulated hemispherical cell cortex -, dependent on the specific parameter values chosen. b. In
vivo. Formation of artificial nodes by oligomerization between 3GFP and 3GBP-mCh-CaaX which
is constitutively localized to the plasma-membrane. Images are a maximum intensity projection of
a 3D-deconvolved, wide-field z-stack. Scale bar: 2 μm.

Supplementary Figure S3. Schematic representation of the co-localization analysis
framework.
a. An experimental cumulative nearest neighbour curve (‘G‘) is computed by putting together data
from ‘n‘ available images of cell ends displaying two different populations of nodes (circles,
triangles). b. A corresponding simulated G curve is computed using ‘n’ simulated uncorrelated cell
end point patterns having a similar distribution to that from the experimental data. c. ‘m’ distinct
simulated G curves are thus produced, to provide a mean (right panel, red dotted line) and a range
of variation around it (bottom and right panels, grey area). By construction, if an experimental
curve (top and right panels, smooth black line) falls outside the grey area, the two node
populations used to compute the experimental curve are considered to be significantly correlated
in space, with a significance 1/(m+1). Bootstrap resampling is then used to estimate a 95%
confidence band (red band around the black line). d. Simulated G curves of point patterns with
different degrees of correlation (coloured lines). Pairs of partly correlated point patterns were
computed by generating a random point distribution and a second point distribution identical to the
first, but modified by randomly displacing each of its points by one step of free diffusion of
dissusion rate σ. The different radii evaluated gave rise the different coloured lines displayed. As
expected, the bigger that radius of displacement the lower the correlation of the data, as measured
by the G curve.

Supplementary Figure S4. Dimerization of Tea1-GBP-mCh and Tea3-GFP in cells causes an
aberrant cell polarity phenotype.
Images are maximum intensity projections of wide-field z-stacks. Scale bar: 2 μm.

Supplementary Figure S5. A genetic fusion of Tea1 to Tea3 at the Tea3 locus results in a
lateral cortical mislocalization of both polarity factors, as well as in gross polarity defects.
Only the GFP channel image is shown. The image is a maximum intensity projection of a 3D-
deconvolved, wide-field z-stack. Scale bar: 2 μm.

Supplementary Figure S6. There is no detectable, significant correlation between the growth
status of a cell end and the co-localization of Tea1 and Tea3.
Left: example maximum intensity projection of one of the three-colour images used; Arp2-CFP is
used as a growth marker. Middle: distribution of the number of Arp2-CFP nodes counted; the 25%
of cell ends with the highest abundance of actin patches were defined as ‘growing’ and the 25% of
cell ends with the lowest actin patch abundance as ‘non-growing’ (n=27 cells and n=29 cells,
respectively). Right: spatial correlation plot comparing the growing and the non-growing cell end
sub-populations; both curves are well within the confidence band of the other and they were found
to be not significantly different (p>0.05).

Supplementary Figure S7. Co-localization of Tea1-3mCh and Tea3-GFP nodes at cell ends is
stronger in mitosis than in interphase.
a. Cell cycle stage can be monitored by co-expressing in cells the B-cyclin Cdc13-GFP, which
accumulates in the cell nucleus during interphase and is rapidly degraded in mitosis b. Spatial
correlation plot comparing interphase and mitosis sub-populations of cell ends; the curves are
outside the confidence band of one another and the difference was found to be statistically
significant (p<0.05; n=8 cells for both conditions). c. Cell cycle stage in assigned using an SPB
marker, a single SPB indicates the cell is in interphase; two separated SPBs within a 4m radius of
each other indicates the cell is in mitosis. Images are from 3D-deconvolved, wide-field z-stacks.
Scale bar: 2 μm.

Supplementary Figure S8. Tea1 cell end enrichment persists even during prolonged mitosis,
but only in the presence of Tea3.
Cold-sensitive Tea3-GFP Tea1-3mCh nda3-KM311 and Tea1-GFP nda3-KM311cells were
arrested in early mitosis by incubation at the restrictive temperature. DAPI staining was used to
identify mitotically arrested cells with condensed chromosomes. Images are from 2-D
deconvolved 6m OAI sweeps. Scale bar: 2 μm.

10
Supplementary Table S1. List of cell strains used.
S. pombe strains:
Strain Genotype Source or
Reference
MH120 Tea1-GFP-L-nat Z2-mCh-atb2-hph h- his7-366
leu1-32 ura4-D18 ade6-M216
This Study
MH121 Tea3-L-GFP-nat Z2-mCh-atb2-hph h- his7-366 leu1-
32 ura4-D18 ade6-M216
This Study
MH0075 Tea4-L-GFP-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
KS706 kanMX6:nmt81GFP-mod5 h- ade6-M210 leu1-32
ura4-D18
11
MH0097 Pom1-L-GFP-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
FC1798 bud6-3GFP:kanMX6 ade6-M216 leu1-32 ura4-D18
h+
58
MH225 Tea4-3GFP-LK-nat h- his7 leu1 ura4 ade6-M216 This Study
RCS492 Tea1-LL-GBP-mCh-Hph Tea4-GFP-Nat
This Study
MH230 Tea1-mGFP-LK-kan h- his7 leu1 ura4 ade6-M216
This Study
MH232 Tea3-mGFP-LK-kan h- his7 leu1 ura4 ade6-M216 This Study

11
MH234 Tea4-mGFP-LK-kan h- his7 leu1 ura4 ade6-M216
This Study
MH596 co2::Ptea1-3GFP-Tadh1-bsd lys1::Padh1-3GBP-
mCherry-CaaX-Tadh1-hph leu1 ura4 ade6-M216 h+
This Study
MH222 Tea1-3GFP-LK-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
MH220 Tea3-3GFP-LK-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
RCS460 h- ade-210 leu1-32 ura4-D18 pREP-GFP-CaaX This Study
972 h- Own
Collection
MH107 Tea1-L-GFP-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
MH095 Tea3-L-GFP-nat h- his7-366 leu1-32 ura4-D18
ade6-M216
This Study
RCS263 kanMX6:nmt81GFP-mod5 Tea1-3mCh-hph leu1-32
ura4-D18
This Study
KS5980 h+ Tea1∆(726-762aa)-GFP-kanMX ade6-M210
leu1-32 ura4-D18
8
MH573 h+ Tea1∆(726-762aa)-GFP-Kan co2::Ptea1-3GBP-
mCh-bsd leu1-32 ura4-D18 ade6-M210
This Study
RCS586 Tea1-L-GFP-nat co2::Ptea1-3GBP-mCh-bsd This Study
RCS260 Tea3-GFP-L-nat Tea1-3mCh-hph leu1-32 ura4-D18 This Study

12
ade6-M216
RCS261 Tea4-GFP-L-nat Tea1-3mCh-hph leu1-32 ura4-D18
ade6-M216
This Study
RCS262 Pom1-L-GFP-nat Tea1-3mCh-hph leu1-32 ura4-
D18 ade6-M216
This Study
RCS355 Arp3-3GFP-L-nat Tea1-3mCh-hph leu1-32 ura4-
D18 ade6-M216
This Study
RCS462 Tea1-L-GFP-nat Tea3-3mCh leu1-32 ura4-D18
ade6-M216
This Study
RCS461 kanMX6:nmt81GFP-mod5 Tea3-3mCh-hph leu1-32
ura4-D18
This Study
RCS463 Pom1-L-GFP-nat Tea3-3mCh-hph leu1-32 ura4-
D18 ade6-M216
This Study
RCS486 Tea4-L-GFP-nat Tea3-3mCh-hph leu1-32 ura4-D18
ade6-M216
This Study
RCS487 Arp3-L-3GFP-nat Tea3-3mCh-hph leu1-32 ura4-
D18 ade6-M216
This Study
RCS485 Tea1-LL-GBP-mCh-Hph Leu1::pShk1-GFP This Study
RCS491 Tea1-LL-GBP-mCh-Hph Tea3-GFP-L-nat This Study
MH196 mCh-L-Tea1-Tea3-GFP-L-Kan Tea1::nat leu1 ura4
ade6-M216
This Study
RCS496 Arp2-CFP-kan Tea1-YFP-kan Tea3-3mCh-hph This Study
RCS537 Tea3-L-GFP-nat Tea1-3mCh-Hph Leu1::Cdc13- This Study

13
GFP-Leu1
RCS434 Alp6-L-GFP-kan Tea3-L-GFP-nat Tea1-3mCh-Hph This Study
RCS554 Tea1-L-GFP-nat Tea3::kan nat Z2-mCh-atb2-hph This Study
RCS593 Tea3-GFP-nat Tea1-3mCh-hph nda3-KM311 This Study
RCS601 Tea1-GFP-nat Tea3D-kan nda3-KM311 This Study
S. cerevisiae strains:
Strain Genotype Source and
Reference
SY159 MATa, KEL1::KEL1-GFP(KANR)
59
MGY242 MATa, SPA2::SPA2-GFP(URA3)
Derived from
15D 60

14
SUPPLEMENTARY METHODS
S. pombe Filipin Staining
Cells of strain MH222 in exponential growth in YES were incubated with 5mg/ml Filipin (from a
5mg/ml stock n DMSO; Sigma, F9765) for 10 minutes on lectin pre-coated glass-bottomed dishes.
Following incubation cells were washed and suspended in EMM as described above. Imaging was
performed with a DeltaVision (Applied Precision), comprising an Olympus 1X71 widefield
microscope, an Olympus UPlanSapo x100 oil immersion lens (NA1.4) and an Photometrics
CoolSNAP HQ2
camera. Filipin and GFP were imaged with DAPI and FITC filter sets
respectively. Stacks were taken of steps 0.2µm apart for 21 planes. Images were captured and
analyzed using SoftWoRx (Applied Precision).
S. pombe Strain and Plasmid Construction
For L-GFP-nat tagging, we constructed and used pMS-GFP-nat, a pFA6a-based plasmid
containing a linker sequence (encoding GAGAGAGAGAFSVPITT). For tagging of tandem copies
of fluorescent proteins, plasmids pMS-3GFP-nat and pMS-3mCherry-hph, which contain the same
linker sequence 45
, were used. In the Z2-mCherry-Atb2 strains the mCherry-atb2 gene exists as an
extra copy of endogenous atb2+ and is expressed under the control of the atb2
+ promoter (Patb2)
and terminator (Tatb2). The Z2-mCherry-Atb2 strain was constructed by amplification of a Patb2-
mCherry-Atb2-Tatb2-hph DNA fragment and integrated into the gene-free region adjacent to zfs1+
on chromosome II of the atb2+ strain.
For construction of the GFP-CaaX plasmid, the DNA encoding EGFP was amplified with a pair of
oligonucleotides. The 5’ oligo contains the NdeI restriction site, whereas the 3’ one harbours the
sequence encoding the membrane-targeting CaaX motif of K-Ras

15
(KMSKDGKKKKKKSTKTKCVIM) followed by the termination codon and the BamHI
restriction site. The amplified fragment was then cloned into pREP1 using the same restriction
sites. The plasmid was then transformed into a wild-type S. pombe strain based on a standard
protocol 46
.
Cdc13-GFP was constructed by amplification of cdc13 and 650bps of its upstream region,
followed by insertion into the integrative plasmid pJK148-GFP at the SalI and NotI sites. The
plasmid pJK148-Cdc13-GFP was then digested with XbaI and integrated at the endogenous cdc13
locus.
To construct the GFP binding protein (GBP) strains, a standard plasmid was adapted. A plasmid
hargouring the GBP 12
coding sequence was a gift from J. Pines (Gurdon Institute). To make the
plasmid for tagging of GBP-mCherry, pMS-mCherry-hph was linearised with NheI and the
amplified GFP sequence flanked with NheI and XbaI was cloned there. The standard methods
above were used to tag the genes. The cytoplasmic (untagged) ‘GFP alone’ strain was generated
by insertion of a 524bp section of the shk1 promoter upstream of the GFP cassette in plasmid
pJK148. The pShk1-GFP construct was then inserted into the leu1-32 locus. Live-cell imaging of
the GFP- and GBP–tagged strains was performed on lectin pre-coated glass-bottomed dishes on
the DeltaVision microscope as decribed above. For scoring of mono-polar versus bipolar cells and
of bent versus unbent cells, strains were first fixed and washed as described above. Following
fixation cells were mounted on lectin coated glass-bottomed dishes as described above. Following
washing of cells once in PBS to remove un-adhered cells, 1ml of working Calcofluor solution was
pipetted over the cells (from a 10x stock of CalcofluorWhite Solution; Sigma: 18909). The stained
cells were imaged using the DAPI channel on the DeltaVision and scored into monopolar versus
bipolar as described previously 47
.

16
To generate a fusion of Tea1 and Tea3, the ura4+ cassette gene was inserted in between the
promoter and the coding region of the tea3+ gene of the tea3-GFP-kanR strain with the ura4
background. The integrant is denoted Ptea3-ura4+-tea3ORF-GFP-kanR. Next we prepared a
plasmid containing the fusion gene of mCherry and tea1+ coding regions (mCherry-tea1ORF).
Using this plasmid as a template, a PCR reaction was performed to flank the fusion gene with a
part of the promoter and the coding region of the tea3+ gene (Ptea3-mCherry-tea1ORF-tea3ORF).
The amplified fragment was introduced to the Ptea3-ura4+-tea3ORF-GFP-kanR strain so that the
ura4+ gene is replaced by the mCherry-tea1ORF construct. The ura4 auxotrophic cells were
selected by 1 mg/ml 5-fluoroorotic acid (5-FOA) and correct integration was confirmed by PCR.
The resultant strain expresses the fusion protein mCherry-Tea1-Tea3-GFP. Live and fixed cell
imaging was performed as above.
Lateral (‘Side On’) Imaging of S. pombe cells
Cells were mounted onto glass-bottomed MatTek dishes pre-coated with lectin as described above.
Imaging was performed with a DeltaVision microscope (Applied Precision) and images were
captured and analyzed using SoftWoRx (Applied Precision). Standard Z-stacks were imaged at
0.4m intervals over 16 focal planes using the appropriate filters. Cell length and curvature
phenotype data was extracted from analysis of septated cells. Live-cells were stained with 1/100
dilution of Calcofluor White Stain (Sigma: 18909) in YES. Cell curvature was visually scored.
Time-lapse imaging of apical node drift was performed with the Optical Axis Integration (OAI)
function of the DeltaVision microscope with a 6m continuous sweep of the whole cell taken at 6
minute intervals.
Quantitation of cell end fluorescence enrichment from laterally (‘Side-on’) imaged cells was done

17
using custom-written MATLAB (Mathworks) code. Firstly the outlines of cells were automatically
segmented in 2D from the transmitted light image. Then, cell end fluorescence enrichment in the
GFP channel was automatically computed from an average projection of the original 3D z-stack by
calculating the ratio between the mean fluorescence at the cell end divided by the mean
fluorescence elsewhere in the cell. For each condition ‘dark’ (non-GFP containing) cells were used
to estimate auto-fluorescence. Cell cycle stage was manually assigned to each cell by visual
inspection of the microtubule marker mCh-Atb2.
For imaging of cold-sensitive nda3-km311 strains, cells were switched from incubation at the
permissive temperature of 320C to the restrictive temperature of 20
oC for two hours prior to
imaging. To visualize the nucleus by DAPI staining, cells were washed and kept in ice-cold
1xPBS, then 1l of cell suspension was mixed with 1l of 50g/ml DAPI on a glass-slide.
Cortical Cluster (‘Node’) Detection and Quantitation
For node counting, full 3D z-stacks acquired from cell ends of interest (as described in the
Experimental Procedures in the main text) were used. ‘Head on’ cell ends that were in focus and
properly aligned were manually cropped from the deconvolved images and were centred in a 4.5 x
4.5 µm2 image region. Fluorescent (GFP- or mCherry-labelled) node detection was then performed
by using the ‘spot detector’ module of the open source software ICY from Institut Pasteur
(http://www.bioimageanalysis.org/icy/index.php; 48
), which uses wavelet thresholding 49
. Only the
first planes were used with a threshold of 40 (arbitrary units). All spots detected within 2 µm of
the center were kept and taken to belong to the cell end analyzed. Quantitation of node number and
intensity in Fig. 2d was performed from the sum projections of deconvolved wide-field z-stacks,
using custom-written MATLAB (Mathworks) code. Nodes were detected based on local maxima,

18
and a 2D Gaussian was iteratively fitted to each node and subtracted from the image, starting from
the dimmest nodes.
Statistical Assessment of Spatial Correlation Between Nodes
Assessing the co-localisation of two fluorescently-labelled proteins of interest is a long-standing
problem and, while many methods have been proposed, solutions still are very problem dependent.
In our case, we needed not only to assess the spatial co-localisation between nodes of different
proteins of interest (i.e. how many of the nodes of one protein are together with those of another),
but more generally their spatial correlation (are the different node populations suspiciously close
to one another and how much), in a way that could reveal subtle but significant changes. To that
aim, we used the spatial statistics framework initially developed for ecology, which looks at the
point patterns formed by detected objects and assumes they are typical examples of an underlying
point process. The point process seeks to capture the generality behind the collection of point
patterns 13,50,51
. Given a set of n ‘head on’ cell ends decorated by two types of protein nodes
(‘green’ nodes of protein X-GFP and ‘red’ nodes of protein Y-mCherry), we first detected the
green and red node positions in them in 3D, using the software ICY as described above. The 3D
x,y,z coordinates where converted to 2D spherical coordinates, latitude and longitude . A half-
sphere was fitted to the detected nodes and we took, for each point, the coordinates of its
projection on the sphere. To correct for a bias in surface element area in spherical coordinates, the
transformation (q,j)® (cos(q),j) was applied. The rest of the computations were done on those
2D (cos(q),j) coordinates, using geodesic distances between them (see 13 for details).
We then computed the cumulative distribution
Gobs(r) of distances between each Y-mCherry point
found and its nearest detected X-GFP point, using the edge-corrected estimator provided by the

19
SpatStat package (http://www.spatstat.org/; 52
) in the R statistical environment (http://www.r-
project.org/). It is noteworthy that we always examined the distribution of the nearest green point
with respect to each red point and not the reverse, given that the fluorescent signal captured for
mCherry-tagged proteins was consistently less bright and underwent quicker and more severe
photobleaching during imaging.
Confidence band by bootstrap. We used the bootstrap resampling method to compute a confidence
interval to the computed cumulative distribution 53
. Let T = T1,...,Tn{ } be the n cell ends under
study, m bootstrap resample datasets were then computed by random resampling with replacement
from T. A cumulative distribution
Gobs(r) was computed for each of those and the 95% confidence
band was defined as the band that encompassed 95% of the computed
Gobs(r) curves of the
resamples.
Test of spatial independence. We assessed whether a given pair of (X-GFP and Y-mCherry) node
populations were spatially distributed independently of one another or not by simulation-based
Monte-Carlo hypothesis testing 54
against the null hypothesis H0: ’the spatial distribution of the
two collections of protein nodes are independent’. To simulate H0, each channel was modeled as
an independent inhomogeneous spatial Poisson process, i.e. a random distribution of points with a
spatially varying density. The density was estimated from the experimental data, using the R built-
in kernel density estimator. A number m of simulations was done, each with n cell ends, and the
summary statistics Gi for i=1…m was computed for all of them along with its mean, Gavg. We then
computed
gmax = maxi
maxrGavg (r) -Gi(r)| |, and defined the upper and lower envelopes as Gavg + gmax
and Gavg – gmax, respectively. Thus, H0 was rejected with a significance
a =1
M+1 if the summary
statistics of the observed data Gobs laid outside of the region flanked by the upper and lower

20
envelopes (see Supplementary Fig. S3 for a visual summary of the process).
Comparing two pairs of proteins. To assess whether a pair of proteins was significantly more
spatially correlated (i.e. ‘co-localized more’) than another pair, we used the bootstrap method
again. If T a = T1
a,...,Tnaa{ } and T b = T1
b,...,Tnbb{ } were the two collections of cell ends to compare,
we did a hypothesis test against H0:’ Ta and T b are samples from the same population’. (Ba,Bb )
were two datasets of respectively na and nb cell ends obtained by random resampling with
replacement from T a ÈT b . The maximum of the difference between the respective
Gobs(r) curves
of the two resample was then compared to the maximum of the difference between the measured
Gobs(r) of the two original samples. If one was above the other k times out of m resamples, the p-
value of the test was p =k
m. This tested how improbable it would be to find datasets as separated
as the observed ones if their observations were merged together.
As a way to evaluate the procedure, we used correlated simulated point patterns. A red channel
was generated by a random point distribution and a variably correlated green channel was created
by randomly displacing each red point by one step of a random walk on the sphere with diffusion
constant σ. When σ was varied from 0.2 to 0.8, we could see the computed summary statistics
evolving as expected (Supplementary Fig. S3d). On the simulated point patterns, the curves
compared two by two σ = 0.2 and σ = 0.4, σ = 0.4 and σ = 0.6, σ = 0.6 and σ = 0.8 were all seen as
significantly different (p < 0.01). When compared to uncorrelated point patterns, randomly
displaced point patterns were seen as significantly different at a 5% significance for σ ≤ 0.8.
While the proposed framework does not give any insight on the biological origin of a spatial
correlation found, it allows a quantitative comparison of the spatial proximity of node population
pairs and a ‘ranking’ of the proximities for many different pairs of node populations, ranging from

21
full spatial co-alignment to spatial independence. It does so on a data-based basis, without making
unwieldy hypothesis on the processes under study.
In Silico Computer Simulation of Cortical Node Formation
To simulate the formation of polarity factor complexes at the ends of fission yeast cells, we used a
particle-based framework largely based on the theoretical framework developed by Andrews and
Bray in Smoldyn (http://www.smoldyn.org/index.html; 55,56
). Two species were simulated: a) a
Tea1-like polarity factor with potentially two properties, the ablility to oligomerise and the ability
to be delivered either uniformly to the cell end or at localized points of the cell end in a
microtubule-like manner (hereafter called ‘T’); and b) a Mod5-like cortically bound anchor protein
(hereafter called ‘M’) able to interact with T.
Geometry:
We limited our simulations to the end of a fission yeast cell, which we simplified as a half-sphere.
Since we used spherical coordinates, we normalized all parameters by the spherical radius of the
cell end, and characterized the position of every particle using (θ,φ) with 0 < θ < 2π and 0 < φ <
π/2.
M:
In our model, M was assumed to be a point-like particle undergoing Brownian motion at the end of
a fission yeast cell 57
. The amount of free M was assumed to be constant across the cell (due to
very fast cortical diffusion of non-oligomerized M). The enrichment of M at the cell end was due
to the accumulation of M into aggregates like those discussed in the main text.
T:
Once delivered to the cell end, either uniformly or at localized points of the cell end in a

22
microtubule-like manner, T quickly unbound from the simulated cell cortex unless it interacted
with M. The T:M complex was more stable and could grow in size by incorporating more T and M
molecules. As T was the only protein able to oligomerize and T had a single binding site for M,
aggregates could only incorporate M if the total amount of T was greater than the amount of M.
Microtubules:
T could be delivered by microtubule-mimicking events. Microtubules could arrive anywhere on
the cell end (the landing density was empirically calculated from microscopy data). Once
microtubules made contact with cell ends, they delivered a large amount of T in a burst. It is
important to note that the average amount of T delivered per unit time was constant and
corresponded to the experimentally measured value for Tea1.
Reactions:
As simulations occurred in discrete timesteps, zeroth order reactions were pre-calculated as
Poisson events until the cumulative distribution reached 99.95%. First order reactions were
assumed to be exponentially decaying events, and second order reactions (such as protein-protein
interactions) were calculated using the Andrews & Bray framework for discrete Smoluchowski
interactions.
Protein - Protein interactions:
Protein-protein interactions only occurred when two particles were within a distance smaller than
their radius of interaction. Monomeric T could interact with monomeric M to form aggregates, and
aggregates could interact with monomeric T, monomeric M and other aggregates. In the (rare) case
where multiple aggregates were within a radius of interaction of each other, fusion between
aggregates could occur.
Rendering:

23
The history of every particle was saved during the simulations. Later, we rendered the simulations
into videos by calculating a 2D histogram of the number of particles at every time step for both T
and M, using the microscope resolution as the size of the histogram. We applied a Gaussian blur
(to simulate the point-spread function of the microscope) to the histogram data, and assigned the
resulting data to different colour channels (T in red, M in green).
Parameter Sweep:
We investigated the effect of three parameters in node formation: the unbinding rate of T, the
interaction radius of T (used for T-aggregate and for aggregate-aggregate interaction) and the
interaction radius of M (used for T-M and for M-aggregate interactions). Each parameter was
varied across a range of values, and for each point the average of ten simulation runs was
computed (Supplementary Fig. S2).

24
SUPPLEMENTARY REFERENCES
45 Sato, M., Toya, M. & Toda, T. Visualization of fluorescence-tagged proteins in fission
yeast: the analysis of mitotic spindle dynamics using GFP-tubulin under the native
promoter. Methods Mol Biol 545, 185-203, doi:10.1007/978-1-60327-993-2_11 (2009).
46 Kanter-Smoler, G., Dahlkvist, A. & Sunnerhagen, P. Improved method for rapid
transformation of intact Schizosaccharomyces pombe cells. Biotechniques 16, 798-800
(1994).
47 Martin, S. G. & Chang, F. New end take off: regulating cell polarity during the fission
yeast cell cycle. Cell Cycle 4, 1046-1049, doi:1853 [pii] (2005).
48 de Chaumont, F., Dallongeville, S. & Olivo-Marin, J. C. ICY: a new open-source
community image processing software. . IEEE International Symposium on Biomedical
Imaging (ISBI). (2011).
49 Olivo-Marin, J. C. Extraction of spots in biological images using multiscale products. .
Pattern Recognition. 35, 1989-1996 (2002).
50 Baddeley, A. Spatial Point Processes and their Applications. Vol. 1892 1-7 (2007).
51 Helmuth, J. A., Paul, G. & Sbalzarini, I. F. Beyond co-localization: inferring spatial
interactions between sub-cellular structures from microscopy images. BMC Bioinformatics
11, 372, doi:1471-2105-11-372 [pii] 10.1186/1471-2105-11-372 (2010).
52 Baddeley, A. & Turner, R. spatstat: An R Package for Analyzing Spatial Point Patterns.
Journal of statistical software 12 (2005).
53 Efron, B. & Tibshirani, R. An introduction to the bootstrap. (Chapman & Hall, 1993).
54 Mattfeldt, T. A brief introduction to computer-intensive methods, with a view towards

25
applications in spatial statistics and stereology. J Microsc 242, 1-9, doi:10.1111/j.1365-
2818.2010.03452.x (2011).
55 Andrews, S. S., Addy, N. J., Brent, R. & Arkin, A. P. Detailed simulations of cell biology
with Smoldyn 2.1. PLoS Comput Biol 6, e1000705, doi:10.1371/journal.pcbi.1000705
(2010).
56 Andrews, S. S. & Bray, D. Stochastic simulation of chemical reactions with spatial
resolution and single molecule detail. Phys Biol 1, 137-151, doi:S1478-3967(04)83249-2
[pii] 10.1088/1478-3967/1/3/001 (2004).
57 Brillinger, R. A Particle Migrating Randomly on a Sphere Journal of Theoretical
Probability 10, 449-443 (1997).
58 Minc, N., Bratman, S. V., Basu, R. & Chang, F. Establishing new sites of polarization by
microtubules. Current biology : CB 19, 83-94, doi:10.1016/j.cub.2008.12.008 (2009).
59 Geymonat, M., Spanos, A., Jensen, S. & Sedgwick, S. G. Phosphorylation of Lte1 by Cdk
prevents polarized growth during mitotic arrest in S. cerevisiae. J Cell Biol 191, 1097-
1112, doi:jcb.201005070 [pii] 10.1083/jcb.201005070 (2010).
60 Richardson, H. E., Wittenberg, C., Cross, F. & Reed, S. I. An essential G1 function for
cyclin-like proteins in yeast. Cell 59, 1127-1133, doi:0092-8674(89)90768-X [pii] (1989).