SUPPLEMENTARY FIGURES Figure S-I. Fluorescence intensity and subcellular localization of transfected...
-
Upload
destiny-skinner -
Category
Documents
-
view
218 -
download
2
Transcript of SUPPLEMENTARY FIGURES Figure S-I. Fluorescence intensity and subcellular localization of transfected...

SUPPLEMENTARY FIGURES
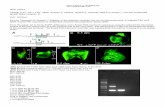
Figure S-I. Fluorescence intensity and subcellular localization of transfected EGFP fusion proteinsHeLa cells were transferred into 96-well plates (Costar) at 5 x 103/well each, incubated for 24 h and then transfected with EGFP, EGFP-importin 2 full-length and EGFP-importin 2 C-mutant constructs using Effectene Transfection Regent (Qiagen). To quantitate cytoplasmic and nuclear fluorescence, images were captured on an ArrayScan VTI instrument (Thermo Fisher) and analysed using Cellomics software and the Compartmental Analysis Bioapplication. The analysis algorithm uses DAPI fluorescence to define a mask used to measure nuclear fluorescence. The region for whole cell measurements is generated by dilating the mask to cover as much of the cytoplasm as possible without going outside the cell boundary. The cytoplasm region is generated by removal of the nuclear from whole cell regions to create a ring. Average pixel intensities of EGFP within these regions were calculated. Non-transfected cells (with autofluorescent pixel intensity similar to non-transfected controls) were gated out of the analysis. Three independent experiments were done, and 300 cells were measured for each transfection. One of the independent results was presented here.
Figure S-II. Validation of down-regulated genes by quantitative PCR HeLa cells were transfected with EGFP as a control, EGFP-importin 2 full-length or C-mutant constructs, and mRNA expression levels of indicated genes were analyzed by qPCR. The results were from three independent experiments and were presented in comparison to values in the EGFP-expressing cells as the mean ± SEM (n=3 each). **p<0.01; Student’s t-test. Primers are shown in Supplemental Table S-III.
Figure S-III. Validation of ChIP analysis (A) HeLa cells were exposed to 200 μM H2O2 for 30 min and subjected to immunoprecipitation using an
anti-importin 3 antibody in accordance with ChIP assay. The bound proteins were analyzed by Western blotting using anti-importin 3 antibody.(B) HeLa cells were exposed to 200 μM H2O2 for 30 min and subjected to immunoprecipitation using an
anti-p53 antibody. After DNA purification from the precipitated the chromatin sample, PCR was performed to amplify sequences in the indicated promoter regions.
Figure S-IV. Establishment of STK35 expressing stable cell lines and knock-down experiments by RNAi oligonucleotides (A) 293F cells were transfected with wtag, wtag-STK35S or wtag-STK35L1 and established stable cell lines were selected by single cell clones. Equal amounts of cellular proteins contained in total cell extracts were subjected to SDS-PAGE and detected by Western blotting using anti-GST antibody. The band provided with asterisk is suspected to be a degradation product of STK35L1. (B) 293F stably expressing wtag as a control (Cont.), wtag-STK35S or wtag-STK35L1 were pretreated for 30 min in the absence or presence of 50 μM zVAD-fmk and exposed to 2 mM H2O2 for 2 h or 0.5 μM STS
for 24 h, respectively. Equal amounts of cellular proteins contained in total cell extracts were subjected to SDS-PAGE and analyzed by Western blotting for PARP, and GAPDH. (C) 293F cells stably expressing wtag as a control (Cont.), wtag-STK35S or wtag-STK35L1 were exposed to 0.5 μM STS for 4 h. Cells were stained with PI and then sorted by FACS. Values are means ± SEM (n=3 each) of PI-positive cells.(D) HeLa cells were transfected with Luciferase siRNA (siLUC.) as a control or STK35 siRNA oligonucleotides (siSTK35-1 or siSTK35-2) and incubated for 48 h. The mRNA expression levels of STK35 were analyzed by qPCR. The results are from three independent experiments and are presented in comparison to the values in control siRNA-expressing cells as the mean ±SEM (n=3 each). **p<0.01; Student’s t-test.

EGFP-importin 2 Full
EGFP-importin 2 C-mut
4Total intensity
(log10)
0
10
20
5 64
N/C
rat
io
Total intensity (log10)
0
10
20
5 6 4Total intensity
(log10)
0
10
20
5 6
EGFP
Figure S-I
N/C
rat
io
N/C
rat
io

Figure S-II
TNFRSF12A
ATF3
GADD45a
CTH
HRK
TRIB3
p8a
EGR1
GADD153
p21
**
**
**
**
**
**
****
**
**
**
**
**
** ******
**
0
1
1.2
0.2
0.4
0.6
0.8
Re
lativ
e f
old
ch
an
ge
EGFP
EGFP-importin 2 Full
EGFP-importin 2 C-mut

Inp
ut
No
rma
l Ig
G (
Ra
bb
it)
An
ti-p
53
p21
STK35
B
Importin 3
IgG heavy chain
kDa
30
40
5060
80
Inpu
t
Nor
mal
IgG
(G
oat)
Ant
i-im
port
in
3A
Figure S-III

Cleaved PARP
GAPDHC
ont.
ST
K35
SS
TK
35L1
Un
tre
ate
d
H2O
2
H2O
2
+ z
-VA
D-f
mk
ST
S
ST
S+
z-V
AD
-fm
k
Con
t.S
TK
35S
ST
K35
L1C
ont.
ST
K35
SS
TK
35L1
Con
t.S
TK
35S
ST
K35
L1C
ont.
ST
K35
SS
TK
35L1
B
Figure S-IV
% P
I p
osi
tive
ce
lls
Cont.
STK35S
STK35L1
STSUntreated0
10
20
30
40
50
60
70
80
C
wta
g
wta
g-S
TK
35
S
wta
g-S
TK
35
L1
kDa
20
30
40
50
60
80
*
A
Re
lativ
e S
TK
35
mR
NA
leve
l
0
0.2
0.4
0.6
0.8
1.0
1.2
**
siS
TK
35
-2
**
siS
TK
35
-1
siL
UC
.
D

a) Fold change (≤ or ≥ 2-fold) indicates the ratio of microarray signals in the EGFP-importin 2 full-length gene expressing cells compared to those in the control EGFP transfected cells.b) Fold change (≤ or ≥ 2-fold) indicates the ratio of microarray signals of the EGFP-importin 2 C-mutant gene expressing cells to those of the control EGFP transfected cells.c) Partial molecular function/Characterization is provided according to the PANTHER classification system (http://www.pantherdb.org/) d) Abbreviations; N: Nucleus, C: Cytoplasm, M: Membrane, NM: Nuclear Membrane, PM: Plasma Membrane, Ce: Centrosom, Mit: Mitochondria, S: Secreted, G: Golgi, ER: Endoplasmic Reticulum, AJ: Adherents Junctions, CG: Cytoplasmic granular
Table S-I. Summary of up- or down-regulated genes in microarray analysis Accession Gene name Symbols a)Full b)C-mut c)Molecular Function d)Cellular
No. Characterization LocalizationNM_080836 Serine/threonine kinase 35 STK35/CLIK1 +20.14 +5.82 Protein phosphorylation N/C
Cell structureAK001520 Serine PI Kazal type 5-like 3 SPINK5L3 +2.26 +2.11 Proteolysis Unknown
NM_022107 G-protein signalling modulator 3 GPSM3/G18/AGS4 -9.12 -8.92 G-protein mediated signaling N/C/PM
NM_003707 49 kDa TATA box-binding protein- TIP49/RUVBL1 -3.25 -4.65 Transcription regulation N/C/Minteracting protein Ponti52/NMP238 Chromatin remodeling
NM_025241 UBX domain containing 1 UBXD1 -2.97 -4.12 AAA ATPase N/C/Ce
NM_005532 Interferon, alpha-inducible IFI27/IGS12/p27 -2.21 -3.94 Immune response NMprotein 27
NM_006709 Euchromatic histone-lysine EHMT2/G9A -2.72 -3.32 Chromatin remodeling NN-methyltransferase 2
NM_004024 Activating transcription factor 3 ATF3 -2.49 -3.22 Transcription regulation N
NM_005169 Paired-like homeobox 2a PHOX2a -3.05 -3.19 Transcription regulation NNeurogenesis
NM_199424 WW domain containing E3 WWP2 -2.92 -3.06 Proteolysis N/Cubiquitin protein ligase 2
NM_001924 Growth arrest and DNA-damage- GADD45a -2.28 -2.85 Stress response Ninducible, alpha Cell/cycle
NM_001902 Cystathionase CTH/CGL/CSE -3.48 -2.80 Amino acid metabolism C(cystathionine gamma-lyase)
NM_139320 CHRNA7-FAM7A fusion isoform 1 CHRFAM7A -2.53 -2.78 Cation transport Unknown
NM_016639 Tumor necrosis factor receptor TNFRSF12A/FN14 -2.83 -2.69 Apoptosis/angiogenesis M/Gsuperfamily, member 12A TWEAKR
NM_001673 Asparagine synthetase ASNS -3.22 -2.65 Amino acid biosynthesis C
NM_012323 Transcription factor MAFF MAFF -2.36 -2.48 Transcription regulation N
NM_032621 Brain expressed X-linked 2 BEX2 -2.99 -2.40 Transcription regulation N
NM_004563 Phosphoenolpyruvate PCK2 -2.32 -2.33 Gluconeogenesis Mitcarboxykinase 2
NM_022168 Interferon induced with helicase IFIH1/MDA5 -2.01 -2.32 Nucleic acid metabolism CC domain 1 Virus recognition
AK056421 Trinucleotide repeat containing TNRC6C -2.11 -2.31 Unknown N/A6C
NM_003806 Harakiri, BCL2 interacting HRK/DP5 -2.39 -2.29 Apoptosis CG/Mitprotein
NM_001670 Armadillo repeat gene deletes ARVCF -3.01 -2.26 Cell adhesion AJ/Nin velocardiofacial syndrom
NM_021158 Tribbles 3 TRIB3 -2.53 -2.21 Transcription regulation NApoptosis
NM_000476 Adenylate kinase 1 AK1 -2.15 -2.20 Nucleic acid metabolism C
NM_012385 p8 protein isoform a NUPR1/p8a/COM1 -2.34 -2.19 Cell cycle/Apoptosis N
NM_002985 Chemokine (C-C motif) CCL5/RANTES -2.05 -2.18 Immune response Sligand 5
NM_001751 Cysteinyl-tRNA synthetase CARS/CRS -2.30 -2.17 Protein metabolism C
NM_004904 cAMP responsive element CREB5/CRE-BPa -2.07 -2.17 Transcription regulation Nbinding protein 5
NM_003714 Stanniocalcin 2 STC2 -2.31 -2.17 Hormone/Cell survival S/ER//G
NM_013376 SERTA domain containing 1 SERTAD1/SEI-1 -2.48 -2.14 Transcription regulation NTRIP-Br1 Cell cycle
NM_001964 Early growth response 1 EGR1/Zif268 -2.28 -2.10 Transcription regulation N
NM_004907 Immediate early response 2 IER2/pip92 -2.01 -2.07 Apoptosis N/CNeuronal differenciation
NM_004083 DNA-damage-inducible CHOP/GADD153 -2.45 -2.05 Transcription regulation Ntranscript 3 Stress response

Table S-II. Summary of replication-dependent histone genes H1 H2A H2B H3 H4
Chromosome F C F C F C F C F CHIST1 H1A 1.00 1.00 H2AA -1.06 -1.13 H2BA 1.00 1.00 H3A -1.07 +1.03 H4A +1.13 1.00
6p21-22 H1B 1.00 1.00 H2AB -2.08 -1.72 H2BB -1.64 -1.95 H3B -1.85 -1.16 H4B +1.03 -1.63 H1C -3.06 -2.73 H2AC -1.54 -1.77 H2BC -1.71 -1.88 H3B -1.28 -1.94 H4C +1.04 -1.62 H1D -2.66 -2.42 H2AC +1.02 +1.15 H2BD -1.95 -1.87 H3C -1.18 -1.08 H4D +1.04 -1.58 H1E -2.50 -2.22 H2AD -4.22 -3.33 H2BE -1.73 -2.09 H3D -1.41 -2.21 H4E -1.14 -1.16 H1T -2.22 +1.17 H2AE -4.09 -3.31 H2BF -1.81 -2.16 H3D -2.17 -1.30 H4F -2.49 -2.09
H2AG -2.77 -2.30 H2BG -1.50 -1.87 H3E -1.10 +1.02 H4G -3.82 -3.29 H2AG -2.66 -1.64 H2BH -1.73 -2.09 H3F -1.28 -1.14 H4H -2.51 -2.16 H2AK -2.59 -1.94 H2BI -1.61 -1.94 H3G +1.10 -1.25 H4I -2.21 -2.02 H2AK -3.55 -1.83 H2BJ +1.13 -1.83 H3G -1.32 1.02 H4J -1.63 -1.63 H2AH -3.42 -2.94 H2BJ -1.91 +1.10 H3H -1.07 -1.08 H4K -1.61 -1.69 H2AJ -3.28 -2.51 H2BK -1.20 -1.76 H3H -1.37 -1.34 H4L +1.06 -1.55 H2AJ -3.51 -2.86 H2BL -1.79 -2.09 H3H -1.43 -1.21 H2AL -3.22 -3.03 H2BM -1.59 -1.89 H3I -1.17 -1.05 H2AM -3.25 -1.04 H2BN -1.66 +1.24 H3J -1.04 1.05 H2AM -1.87 -2.79 H2BN -1.46 -1.88
H2B0 -1.73 -2.05
HIST2 H2AA -5.41 -4.00 H2BE -1.82 -1.54 H3C -1.22 -2.77 H4 -2.32 -2.021q21 H2AA -3.21 -2.53 H2BE -1.48 -1.79 H3C -3.45 -1.12
H2AB -3.78 -3.08 H2BE -2.21 -1.94 H3C -1.23 -1.12 H2AB -1.95 -1.98 H3C -1.16 -1.06 H2AC -3.52 -2.96 H2AC +1.08 -1.16
HIST3 H2A -3.71 -3.02 H2BB -1.53 -1.83 H3 -1.09 -1.04 H4 -1.56 -1.341q42.13 H2BB +1.32 -1.15 H4 -1.20 -1.01
Others H2AB -1.19 -1.30 H2BS -1.48 -1.76 H2AJ -1.43 -1.14 H2AJ -2.37 -2.05 H2AJ -2.31 -1.93 H2AV +1.06 -1.12 H2AV +1.23 +1.34 H2AX -1.58 -1.32 H2AY 1.00 -1.12 H2AY2 +1.28 +1.23 H2AZ -1.13 +1.08 H2AZ -1.13 +1.09 H2AZ -1.12 +1.07

ATF3 5’-ATGTCCTCTGCGCTGGAATC-3’ 5’-TCTTGTTTCGGCACTTTGCAG-3’
GADD45a 5’-GCTCAGCAAAGCCCTGAGTC-3’ 5’-ATCTGCAGAGCCACATCTCTGTC-3’
CTH 5’-GCCTTTGCTTCAGGTTTAGCAG-3’ 5’-CCAAATTCAGATGCCACTTGC-3’
TNFRSF12A 5’-TGCTTTGGCCCATCCTTG-3’ 5’-TCCGCCGGTCTCCTCTATG-3’
HRK 5’-GCAGGCGGAACTTGTAGGAAC-3’ 5’-GAACCAGCTTTCTCCAAGGACAC-3’
TRIB3 5’-GCCCTGCACTGCCCTACAG-3’ 5’-GGTACCAGCCAGGACCTCAGT-3’
P8a 5’-AAGCTGCTGCCAACACCAAC-3’ 5’-TGGGTGTAGTGTCCATGGTCTG-3’
EGR1 5’-CAGCCCTACGAGCACCTGAC-3’ 5’-AAAGCGGCCAGTATAGGTGATG-3’
GADD153 5’-GCAAGAGGTCCTGTCTTCAGATG-3’ 5’-AGCAGGGTCAAGAGTGGTGAAG-3’
ACTB 5'-GCACTCTTCCAGCCTTCCTTC-3' 5'-TGTGTTGGCGTACAGGTCTTTG-3'
p21 5'-CCTCATCCCGTGTTCTCCTTT-3' 5'-GTACCAACCCAGCGGACAAGT-3'
H1 5’-CCTGTAAAGAAGAAGGCGGCCAAA-3’ 5’-CAGAGAAACTCCGCTACGCTCTTT-3’
H2A 5’-AGCTCAACAAGCTTCTGGGCAA-3’ 5’-TTGTGGTGGCTCTCGGTCTTCTT-3’
H2B 5’-TGCGCCCAAGAAGGGTTCTAAA-3’ 5’-ACGAAGGAGTTCATGATGCCCA-3’
H3 5’-TGCTCATCCGCAAACTGCCATT-3’ 5’-AGTGACACGCTTGGCGTGAATA-3’
H4 5’-ACCGTAAAGTACTGCGCGACAA-3’ 5’-TTCTCCAGGAACACCTTCAGCA-3’
STK35 5’-GTCCTGCAGCGCAATGG-3’ 5’-GGTAAAGCTGCGAGCTCTTGTT-3’
GAPDH 5’-TATTGCCATCAATGACCCCTT-3’ 5’-CTCCACGACGTACTCAGCG-3’
Table S-III. Primers for qPCR

Broad Institute, Cambridge, MA, USA (http://www.broadinstitute.org/gsea/index.jsp)a) number of genes in a particular gene setb) enrichment scorec) normalized enrichment scored) nominal pvaluee) false discovery rate* these gene sets were below a NOM p-val of 0.05 and a FDR q-val of 0.25
Full Up-regulated *
Down-regulated
C-mut Up-regulated
Down-regulated
CHR21Q22CHR1P36CHR14Q24CHR8Q24
CHR6P21CHR19P13CHR1Q42CHR16P13CHR20Q11CHR20Q13CHR10Q26CHR14Q32CHR9Q34
CHR14Q24CHR21Q22CHR10Q26CHR8Q24
CHR20Q11CHR19P13CHR6P21CHR9Q34CHR16P13CHR20Q13CHR1Q42CHR1P36CHR14Q32
Gene Set Name
38262015
261815181727193025
20381915
171826251827152630
a)Size
0.5150.3270.2920.229
-0.535-0.504-0.449-0.415-0.405-0.326-0.339-0.309-0.273
0.4590.3940.2900.302
-0.523-0.492-0.434-0.401-0.378-0.341-0.385-0.259-0.221
b)ES
1.5880.9240.7820.567
-1.433-1.211-1.038-1.013-0.962-0.871-0.847-0.840-0.722
1.2881.2790.8100.783
-1.332-1.255-1.204-1.101-0.959-0.947-0.942-0.721-0.635
c)NES
0.0230.5770.7350.954
0.0550.2100.4380.4530.5380.6570.6840.7100.882
0.1420.1230.7460.753
0.1300.1910.2020.3210.5400.5580.5510.8670.969
d)NOM p-val
0.1031.0001.0000.967
0.4740.9641.0001.0000.9500.9940.9000.7980.860
0.5900.3080.9870.781
1.0000.7780.6600.7490.9490.8200.7130.9710.938
e)FDR q-val
Table S-IV. GSEA tests for enrichment of MSigDB C1 gene setsAnalysis of the expression data was performed using the GSEA software (Broad Institute, Cambridge, MA, USA). GSEA examines ranked lists of genes for enrichment of biological pathways contained within datasets for different databases of MSigDB: C1 (positional gene sets), C2 (curated gene sets), C3 (motif gene sets), C4 (computational gene sets), and C5 (GO gene sets).

***
Full Up-regulated HSC_HSCANDPROGENITORS_ADULTHSC_HSCANDPROGENITORS_FETALHSC_HSCANDPROGENITORS_SHAREDBRCA_ER_NEGVHL_NORMAL_UPSTEMCELL_HEMATOPOIETIC_UPALZHEIMERS_DISEASE_UPSTEMCELL_NEURAL_UP
Down-regulated RUTELLA_HEMATOGFSNDCS_DIFFHSC_EARLYPROGENITORS_ADULTCMV_HCMV_TIMECOURSE_ALL_DNHSC_EARLYPROGENITORS_FETALHSC_EARLYPROGENITORS_SHAREDPOD1_KO_DNLEE_TCELLS2_UPHSC_LATEPROGENITORS_FETALHSC_LATEPROGENITORS_ADULTHSC_LATEPROGENITORS_SHAREDBRCA_ER_POSRCC_NL_UPSTEMCELL_EMBRYONIC_UPAGEING_KIDNEY_UPALZHEIMERS_DISEASE_DN
C-mut Up-regulated HSC_HSCANDPROGENITORS_ADULTHSC_HSCANDPROGENITORS_FETALHSC_HSCANDPROGENITORS_SHAREDBRCA_ER_NEGSTEMCELL_NEURAL_UPSTEMCELL_EMBRYONIC_UPVHL_NORMAL_UPSTEMCELL_HEMATOPOIETIC_UPRCC_NL_UPALZHEIMERS_DISEASE_DN
Down-regulated RUTELLA_HEMATOGFSNDCS_DIFFCMV_HCMV_TIMECOURSE_ALL_DNBRCA_ER_POSHSC_LATEPROGENITORS_FETALHSC_EARLYPROGENITORS_ADULTHSC_LATEPROGENITORS_ADULTHSC_LATEPROGENITORS_SHAREDALZHEIMERS_DISEASE_UPPOD1_KO_DNLEE_TCELLS2_UPHSC_EARLYPROGENITORS_SHAREDHSC_EARLYPROGENITORS_FETALAGEING_KIDNEY_UP
Table S-IV. GSEA tests for enrichment of MSigDB C2 gene sets
Gene Set Name
2323231820382947
201715161616361716162020312021
23232318473120382021
20152017171616291636161620
a)Size
0.3990.3990.3990.3820.3200.2390.2510.139
-0.568-0.528-0.525-0.495-0.495-0.469-0.393-0.431-0.422-0.422-0.368-0.350-0.302-0.238-0.209
0.5500.5500.5500.5300.2950.2980.3220.2760.2820.272
-0.541-0.530-0.438-0.466-0.455-0.444-0.444-0.341-0.371-0.292-0.338-0.338-0.195
b)ES
1.0881.0841.0830.9690.8500.7270.7150.455
-1.447-1.271-1.238-1.198-1.155-1.122-1.108-1.050-1.006-1.000-0.937-0.862-0.830-0.588-0.526
1.6421.5981.5931.4700.9950.9250.9220.8950.7870.757
-1.416-1.313-1.156-1.153-1.146-1.102-1.084-0.977-0.925-0.882-0.840-0.832-0.505
c)NES
0.3200.3490.3360.4820.6830.9010.8741.000
0.0480.1660.1890.2440.2880.3150.3160.4060.4760.4880.5670.7070.7670.9500.986
0.0220.0230.0170.0600.4540.5980.5840.6240.7510.805
0.0650.1490.2740.2980.2980.3520.3730.5030.5940.6750.6870.6970.990
d)NOM p-val
1.0001.0000.9200.9951.0001.0000.9770.996
0.7141.0000.9260.8430.8170.7780.7060.7650.7840.7170.7750.8470.8371.0000.984
0.2040.1310.0900.1440.9190.9400.8120.7640.8620.816
0.8630.8181.0000.8980.7400.7200.6570.8080.8310.8310.8300.7740.990
e)FDR q-val

Full Up-regulated YTATTTTNR_V$MEF2_02TGCTGAY_UNKNOWNCACGTG_V$MYC_Q2AACTTT_UNKNOWNGATTGGY_V$NFY_Q6_01
Down-regulated TGANTCA_V$AP1_CTGACATY_UNKNOWNGTGACGY_V$E4F1_Q6TGCGCANK_UNKNOWNTAATTA_V$CHX10_01CAGGTG_V$E12_Q6TTGTTT_V$FOXO4_01MGGAAGTG_V$GABP_BCTTTAAR_UNKNOWNCAGGTA_V$AREB6_01CTGCAGY_UNKNOWNGGGAGGRR_V$MAZ_Q6GGGCGGR_V$SP1_Q6GGGTGGRR_V$PAX4_03TGTTTGY_V$HNF3_Q6TGACCTY_V$ERR1_Q2TATAAA_V$TATA_01SCGGAAGY_V$ELK1_02WTGAAAT_UNKNOWNCTTTGT_V$LEF1_Q2TGGAAA_V$NFAT_Q4_01RTAAACA_V$FREAC2_01RYTTCCTG_V$ETS2_BCTTTGA_V$LEF1_Q2CAGCTG_V$AP4_Q5TGCCAAR_V$NF1_Q6RCGCANGCGY_V$NRF1_Q6TGAYRTCA_V$ATF3_Q6TTANTCA_UNKNOWNGCANCTGNY_V$MYOD_Q6TGACAGNY_V$MEIS1_01GGGYGTGNY_UNKNOWNRNGTGGGC_UNKNOWN
C-mut Up-regulated None
Down-regulated CTGCAGY_UNKNOWNTAATTA_V$CHX10_01TTGTTT_V$FOXO4_01CTTTAAR_UNKNOWNGGGAGGRR_V$MAZ_Q6TATAAA_V$TATA_01RTAAACA_V$FREAC2_01TGACATY_UNKNOWNGTGACGY_V$E4F1_Q6TGANTCA_V$AP1_CCTTTGA_V$LEF1_Q2CTTTGT_V$LEF1_Q2CAGGTA_V$AREB6_01GGGTGGRR_V$PAX4_03GCANCTGNY_V$MYOD_Q6GATTGGY_V$NFY_Q6_01TGCGCANK_UNKNOWNTGACAGNY_V$MEIS1_01TGGAAA_V$NFAT_Q4_01CAGGTG_V$E12_Q6YTATTTTNR_V$MEF2_02TGTTTGY_V$HNF3_Q6WTGAAAT_UNKNOWNMGGAAGTG_V$GABP_BTGCTGAY_UNKNOWNTTANTCA_UNKNOWNCACGTG_V$MYC_Q2TGCCAAR_V$NF1_Q6TGAYRTCA_V$ATF3_Q6GGGYGTGNY_UNKNOWNRCGCANGCGY_V$NRF1_Q6AACTTT_UNKNOWNCAGCTG_V$AP4_Q5RNGTGGGC_UNKNOWNGGGCGGR_V$SP1_Q6TGACCTY_V$ERR1_Q2SCGGAAGY_V$ELK1_02RYTTCCTG_V$ETS2_B
Table S-IV. GSEA tests for enrichment of MSigDB C3 gene sets
1815345126
242125191951622922181947872219342348215745202735351550162618231719
1919622247232021252435571822182619234551181921291526341516175051351987344827
0.4000.3900.2980.2780.230
-0.604-0.511-0.473-0.502-0.487-0.396-0.388-0.426-0.436-0.430-0.400-0.341-0.283-0.352-0.359-0.315-0.333-0.284-0.331-0.262-0.270-0.311-0.289-0.270-0.261-0.306-0.239-0.294-0.265-0.271-0.246-0.233-0.195
-0.610-0.537-0.416-0.508-0.420-0.486-0.500-0.490-0.452-0.455-0.398-0.350-0.442-0.420-0.429-0.390-0.425-0.402-0.340-0.333-0.397-0.389-0.373-0.341-0.359-0.313-0.288-0.337-0.328-0.315-0.235-0.232-0.237-0.272-0.198-0.216-0.196-0.220
1.0400.9600.9060.9020.641
-1.579-1.297-1.257-1.254-1.205-1.204-1.201-1.125-1.112-1.043-1.002-0.996-0.922-0.911-0.901-0.894-0.857-0.842-0.838-0.815-0.794-0.792-0.770-0.755-0.729-0.727-0.717-0.701-0.687-0.667-0.640-0.564-0.483
-1.578-1.381-1.368-1.357-1.327-1.308-1.307-1.288-1.251-1.241-1.185-1.143-1.138-1.135-1.115-1.098-1.091-1.074-1.071-1.059-1.018-0.993-0.988-0.958-0.878-0.868-0.842-0.821-0.808-0.796-0.736-0.734-0.706-0.692-0.674-0.632-0.624-0.612
0.4060.5100.6180.6570.954
0.0120.1420.1780.1810.2370.1720.1680.3130.3080.4190.4720.4620.6170.5810.5940.6170.6840.7360.6970.8020.8080.7700.8220.8730.8620.8270.9000.8650.9090.9040.9290.9680.991
0.0050.0690.0540.0980.0890.1130.1250.1390.1760.1650.2090.2510.2950.2850.3290.3370.3590.3880.3500.3650.4430.4800.4680.5330.6390.6540.7110.7160.7570.7600.9150.8990.8950.8810.9640.9550.9780.974
1.0001.0000.9960.7550.940
0.4241.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0001.0000.995
0.5511.0001.0000.8390.8130.7610.6560.6370.6910.6570.7910.8670.8170.7690.7850.7850.7590.7640.7340.7280.7970.8230.7990.8390.9980.9841.0001.0000.9980.9881.0001.0001.0001.0001.0001.0000.9830.963
Gene Set Name a)Size b)ES c)NES d)NOM p-val e)FDR q-val

Gene Set Name a)Size b)ES c)NES d)NOM p-val e)FDR q-val
Full Up-regulated None
21 -0.195 -0.497 0.995 0.995Down-regulated GCM_NF2
C-mut 21 0.267 0.753 0.812 0.812Up-regulated GCM_NF2
Down-regulated None
Table S-IV. GSEA tests for enrichment of MSigDB C4 gene sets
Gene Set Name a)Size b)ES c)NES d)NOM p-val e)FDR q-val
Full Up-regulated None
Down-regulated None
C-mut Up-regulated None
Down-regulated None
Table S-IV. GSEA tests for enrichment of MSigDB C5 gene sets