Sample Collection System for DNA Analysis of ... - … of Justice and prepared the following final...
-
Upload
nguyendung -
Category
Documents
-
view
219 -
download
3
Transcript of Sample Collection System for DNA Analysis of ... - … of Justice and prepared the following final...
The author(s) shown below used Federal funds provided by the U.S. Department of Justice and prepared the following final report: Document Title: Sample Collection System for DNA Analysis of
Forensic Evidence: Towards Practical, Fully‐Integrated STR Analysis
Author: Eugene Tan Document No.: 236826
Date Received: December 2011 Award Number: 2008-DN-BX-K010 This report has not been published by the U.S. Department of Justice. To provide better customer service, NCJRS has made this Federally-funded grant final report available electronically in addition to traditional paper copies.
Opinions or points of view expressed are those
of the author(s) and do not necessarily reflect the official position or policies of the U.S.
Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
1
SampleCollectionSystemforDNAAnalysisofForensicEvidence:TowardsPractical,Fully‐
IntegratedSTRAnalysisEugeneTan
NetBio,830WinterSt,Waltham,MA02451
This project was supported by Grant Number NIJ 2008-DN-BX-K010 awarded by the National
Institute of Justice, Office of Justice Programs, US Department of Justice.
I. ExecutiveSummary
The goal of this research project was to develop a sample collection system for law
enforcementagentstocollect,protect,anddocumentbiologicalevidenceandtoperforminitial
processing steps ina format compatiblewith rapid sample‐in to results‐outmicrofluidicDNA
analysis. The sample collection system consists of an evidence collection device (a collecting
swab and accompanying tube for swab insertion) and a sample processing cartridge. This
“Smart CartridgeTM” will lyse cells, solubilize and concentrate DNA, and transfer the DNA
solution to a microfluidic biochip. The microfluidic biochip will process the DNA solution to
generatepurifiedDNAcompatiblewiththerequirementsforSTRanalysis.
Theevidencecollectiondevicehasbeendesignedtoacceptblood,buccalcells, saliva,
andcellularsamples.Cotton,modifiedcellulose,foam,nylon,polyesterandrayon‐tippedswabs
wereevaluatedforreproducibilityofcellcollection,easeofcollection,tolerancetocollection
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
2
protocol, liquidabsorption,andapplication in forensicapplications.TheBodeSecurSwabwas
selectedforthesamplecollectionsystem.
To operate the system, the userwill collect a swab sample, insert the swab into the
accompanying collection tube, place the tube into the Smart Cartridge, and press a start
button—nofurtheruserinterventionisrequired.TheDNAgeneratedbythesamplecollection
systemwill thenbe transferredautomatically toNetBio’s fully integratedmicrofluidicbiochip
forSTR typing.Severalapproaches to thesampleprocessingstepswereevaluatedand those
selectedwereintegratedtodevelopanoptimizedSmartCartridge.
Thisresearchhasresultedinthedevelopmentofacrimesceneevidencecollectionand
processing system compatible with an instrument that will generate an STR profile in 45
minutes from sample introduction with minimal operating requirements. The integrated
instrument will process 16 samples in parallel and dramatically reduce the costs (including
labor, space, andvalidation)of settingupandoperatingaDNA lab. Theevidence collection
device and Smart Cartridge are designed such that the system will be easy to operate and
compatible with both forensic and microfluidic requirements. This sample collection system
representsacriticaladditiontoafullyintegratedsystemforforensicDNAanalysis.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
3
II. TableofContents
I. ExecutiveSummary ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐1
II. TableofContents ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐3
III. Introduction‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐5
A. Purpose,Goals,andObjectives‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐5
B. NetBioSample‐intoResults‐outComponents ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐8
1. SamplecollectionandDNArecovery ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐8
2. MicrofluidicDNAExtractionandPurification ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 10
3. RapidMultiplexPCRAmplificationinaMicrofluidicChip ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 13
4. MicrofluidicSeparationandDetection ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 16
C. ReviewofRelevantLiterature‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 19
D. RequirementsofaSampleCollectionSystem‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 23
IV. ResearchDesignandMethods‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 24
A. OverviewofResearchDesign ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 24
B. MaterialsandMethods ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 25
1. MockCaseworkandReferenceSamples. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 25
2. VortexExtractionofDNAfromSwabs. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 25
3. MechanicalAgitationforExtractionofDNAfromSwabs. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 26
4. MicrofluidicBiochipPurification. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 26
5. AutomatedMicrofluidicExtractionandPurificationofDNAfromSwabsinSmart
Cartridge. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 26
6. STRAmplificationReaction. ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 27
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
4
7. STRseparationanddetectioninstrumentation ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 27
V. DisseminationStrategy ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 27
VI. ImplicationsforCriminalJusticePolicyandPractice ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 28
VII.ResultsandDiscussions ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 29
A. Evidencecollectionmedia ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 29
B. ChemicalLysis ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 32
C. SmartCartridgeProcessStep2:SelectionofSilicaFiberMembrane‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 34
D. SmartCartridgeDevelopment ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 35
E. TandemEvaluationandOptimizationoftheThreeProcessSteps ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 36
F. TestingoftheEvidenceCollectionDeviceandSmartCartridge ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 37
VIII. Conclusions ‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐‐ 41
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
5
III. Introduction
A. Purpose,Goals,andObjectives
Theuseofmicrofluidicdevices limitsthesamplesforanalysisto liquids,andtheminiaturized
natureof thedevices limits thevolumeof sample that canbeanalyzed.These limitationson
microfluidic sample format preclude the direct insertion of biological samples collected by
validated forensic collection methods directly into a microfluidic biochip. To microfluidically
process a conventionally collected biological sample, extensivemanual processing would be
requiredtoextract,solubilize,andconcentratetheDNAofinterest.
Thepurposeof thisprojectwas todevelopa sample collection system thatwill allow
law enforcement agents to easily collect, protect and document biological evidence, and to
performrapidsample‐intoresults‐outmicrofluidicDNAanalysisintheforensicslaboratoryor
at the crime scene. Accordingly, this research focused on the development of an evidence
collection device and a sample processing cartridge (termed the “Smart Cartridge”) thatwill
operateintandemwithafullyintegratedforensicinstrumentandmicrofluidicbiochipthatwill
perform DNA extraction and purification, STR amplification, andmicrofluidic separation and
detection.
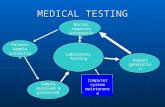
The process flow of the fully integrated STR analysis system is shown in Figure 1. A
forensicsampleiscollectedusingtheevidencecollectiondeviceandthenplacedintotheSmart
Cartridge. After pressing the start button, all required operations are completed, and STR
resultsarereported.Withinthecartridge,severalprocessingstepsareaccomplished(celllysis,
DNA solubilization, and concentration; Figure 2). On the biochip, the DNA is subjected to
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
6
purification(ModuleI),andanappropriatealiquotoftheremainingDNAistransferredtothe
PCR chamber for STR amplification (Module II). The DNA fragments are then subjected to
separationanddetection(ModuleIII),resultinginanSTRprofile.
Figure1.ProcessflowforsampleintoresultsoutSTRanalysisofcaseworksamplesinthefullyintegratedsystem.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
7
Figure2.ProcessflowontheEvidenceCollectionDeviceandSmartCartridge.
Finally, commercially available STR typing kits allow the effective generationof highly
accurateSTRprofiles,butonlywhentheinputDNAtemplatefallswithinanarrowlyoptimized
range.TheDNAadvisoryboard to theFederalBureauof Investigationhas recommendedthe
use of human‐specific DNA quantitation prior to PCR amplification of casework samples as
thereisthepotentialforsamplecontaminationfromnon‐humansourcesincludingnon‐human
mammalian,bacterial, and fungalDNA.Accordingly,a fully‐integratedmicrofluidic system for
forensic human identification should perform human‐specific DNA quantitation in order to
determinepreciselytheamountofDNAtemplatetobesubjectedtoSTRamplification.NetBio
is pursuing microfluidic quantitation under NIJ Award 2008‐DN‐BX‐K009, “RapidMicrofluidic
HumanSpecificDNAQuantitation.”
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
8
B. NetBioSampleintoResultsoutComponents
1. SamplecollectionandDNArecovery
Bode Technology has developed numerous highly advanced and innovative DNA collection
productsincludingtheCrimeScenecollectorandSliderBuccalDNACollector.TheCrimeScene
collectorwasspecificallydesignedtoprovide lawenforcementsecuremorereliableevidence
fromcrimescenes, toprotect thecollectedsamples forshippingandstorage,andtosimplify
documentation of biological evidence. Evaluation of sample collection methods show that
swabbing is more effective for collection and recovery of DNA compared to taping.
Furthermore,cottontippedswabsarethemosteffectivemediaandformatforcollectionand
recoveryofDNA(Figure3).
NetBio previously performed an initial evaluation of lysis solution formulations.
Increases in the DNA extraction efficiency of 1.2 to 2 times compared to that of the
unoptimizedlysissolutionswereachieved.Figure4showstheDNApurificationefficiencyfor10
– 300µl of humanblood in eachof five lysis solution formulations. In this earlier research,
optimizationofthecollectionmediaandlysisbuffer,intandemoverarangeofsampletypes,
was performed tomaximizeDNA recovery.Ultimately, formulations based on a guanidinium
chloride(GuHCl)protocolwereselectedforfurtheroptimization.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
9
Figure3.DNArecoveryforepithelialcellsonpolyesterblendfabriccollectedusingvarioussamplecollectionmatricesanddevices.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
10
Figure4.ComparisonofGuHClandSDSNaClbasedlysissolutionsforextractionofDNAfromwholeblood.
2. MicrofluidicDNAExtractionandPurification
Development of themicrofluidic biochip‐based extraction and purificationmodule has been
completed,andthisworkwasfundedinpartbyNIJAward2007‐DN‐BX‐K184.Preliminarywork
hasbeendemonstratedusing8‐samplebiochips incorporatinga solidphasebind/wash/elute
membrane, and a schematic of the current biochip is shown in Figure 5. For blood, lysis is
performed with a GuHCl‐based‐lysis buffer, and DNA in the lysate is bound to a silica
membrane. The operations to manipulate the fluids through the biochip are driven in an
automated fashion by the sequential actuation of pneumatic lines. Figure 6 shows that the
humangenomicDNApurifiedonthebiochiphasanaveragelengthofapproximately50kbp,a
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
11
size appropriate for STR amplification. Finally, DNA extracted using thismicrofluidic biochip
amplifies effectively and generates STR profiles (Figure 7) that are indistinguishable from
conventionaltubebasedreactions.
Figure5.Schematicofan8samplemicrofluidicchipforextractionandpurificationofDNA.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
12
Figure6.AgarosegelanalysisofDNAextractedfromwholebloodwithmicrofluidicbiochip.(1:Biochipextaction,2:Qiagencontrolextraction,λ:50kbplambdaDNA)
Figure7.STRprofilefromDNAextractedandpurifiedfromwholebloodwithmicrofluidicbiochip.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
13
3. RapidMultiplexPCRAmplificationinaMicrofluidicChip
Rapid multiplexed STR amplification was accomplished by focusing on two major areas:
rigorous optimization of all reaction mix components and cycling parameters and the
developmentof instrumentationtoallowrapidandhighly‐controlledtemperaturetransitions.
The custom thermal cycler shown in Figure 8 is designedwith a high output thermoelectric
cooler/heatermounted toahighefficiencyheat sink, together referred toas theheatpump.
This instrumentacceptsa16‐chambermicrofluidicbiochip (Figure9)which is coupled to the
heatpumpbyapplyingacompressivepressurewithaclampingmechanism.EachPCRchamber
holds 7µl of PCR reaction solution. The 16 PCR reaction solutions are placed into individual
chambers of the microfluidic biochip. The thermal cycler has the ability to heat and cool a
reactionsolutionatratesof15.8°C/sand15.4°C/srespectively,muchfasterthancommercially
available cyclers. Appropriate selection of an enzymes allows a highly multiplexed PCR
reactiontobeperformedinaslittleas17minutes(Figure10).Figure11showsarepresentative
fastSTRprofileusingtheNetBiothermalcyclerandseparatedanddetectedonGenebenchFX.
The fast PCR profiles generated using this approachmeet forensically relevant requirements
including signal strength, stutter, peak‐height ratio, complete non‐template nucleotide
addition,andinterlocusbalance(Giese2009).Figure12showstherelationshipbetweenDNA
template level and signal strength, indicating that the amplification systemhas a near single
copylimitofdetection.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
14
Figure8.NetBiofastthermalcycler.
Figure9.16samplePCRBiochipforrapidhighlymultiplexedamplificationinNetBio’sfastthermalcycler.
1
cm
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
15
Figure10.ThermalprofileforfastmultiplexedamplificationofSTRsinthefastthermalcycler.
Figure11.STRprofilefor0.5ngDNA(9947A)amplifiedinbiochipunderfastthermalcyclingconditionswithprimersfromtheAmpFlSTRProfilerPlusIDPCRamplificationkit.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
16
Figure12.SignalstrengthofvWA17and18,amplifiedinbiochipunderfastthermalcyclingconditionswiththefastthermalcyclerandseparatedanddetectedonGenebenchFX.
4. MicrofluidicSeparationandDetection
Genebench FXTM is a rapid, high resolution, and high sensitivity DNA fragment sizing and
sequencing instrument for laboratoryandfielduse. The instrumentseparatesDNAbasedon
fragment size by electrophoresis on microfluidic biochips, and excitation and detection of
labeledDNA fragments isaccomplishedby laser‐induced fluorescencedetection. Genebench
FX(Figure13)canbeoperatedinboththeforensiclaboratoryandinthefield,haslowpower
consumption,andisCEmarkedundertheLowVoltageDirective73/23/EEC.Separationofthe
DNAfragmentstakeplacewithinamicrofluidicbiochipthatisfilledwithasievingmatrix.The
biochip(Figure14)accepts16samplestoallowforsimultaneousanalysisofmultiplesamples
andrequiredcontrolreactions.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
17
DNA samples for analysis by Genebench‐FX are preparedwith conventionalmethods
andmanualpipettingintothesamplereservoirsofthebiochip.Afterloadingthesamplesand
buffers, thebiochip isplaced intothe instrumentandanalysiscontinueswithout furtheruser
manipulation, with electric fields applied to electrophoretically separate the DNA and
excitementanddetectionof the fluorophoresas theypass through thedetection zone. The
fully integrated version of the Genebench instrument, the focus of NetBio’s development
efforts,willfeatureafullyintegratedmicrofluidicbiochipthatwillacceptforensicsamplesfrom
the user and perform all DNAmanipulations, allowing for sample‐in to results‐out operation
withouttheneedforuserintervention.
Figure15showsanSTRprofileusingtheSGM+primerset(AppliedBiosystems,Foster
City,CA)generatedonGenebench‐FX.ThePCRproductresultedfromtheamplificationof1ng
of007DNAtemplateunderthemanufacturer’srecommendedcondition.Genebench‐FXhas5
color detection capability, allowing the instrument to perform 5 color multiplexed DNA
fragmentsizingassayssuchasthoserequiredbytheAmpFlSTRIdentifilerPCRAmplificationKit
(AppliedBiosystems).
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
18
Figure13.GenebenchFXseries100.Arrowindicatessiteofmicrofluidicbiochipinsertion.
Figure14.16samplemicrofluidicchipforseparationanddetectiononGenebenchFX.
Cathode Anode
SampleandWasteWells
SeparationChannel
ExcitationandDetectionRegion
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
19
Figure15.GenebenchFXSTRprofileof1ngDNA(007)withAmpFlSTRSGM+amplificationkit.
Takentogether,theseresultsshowthedevelopmentalprogressofthemodulesrequiredforafullyintegrated,sample‐intoresults‐outforensicDNAanalysissystem.
C. ReviewofRelevantLiterature
SampleCollectionandElution. Collectionofbiological evidence forDNAanalysis fromcrime
scenes is a process that effectively collects cells from a variety of surfaces, preserves the
collected cells to preventmolecular degradation, and releases thematerial for downstream
processing. Blood, semen, epithelial cells, urine, saliva, bone, and various tissues can be
associatedwiththecrimesceneandrequirecarefulandeffectivecollection(Lee1998).Theage
of the sample as well as the surface on which it is found are factors determining how the
sampleshouldbecollected.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
20
Themostcommoncollectiontechniqueisperformedusingacottonswab.Asingleswab
canbetakenfromanareaorthewet‐drydoubleswabtechniquecanbeused.Thedoubleswab
technique may be the most prevalent, and a number of different fluids including water,
buffered saline, or lysis buffers can be used tomoisten the first swab (Leemans 2006). This
technique allows for dried samples to become re‐hydrated, with the majority of material
collectedonthefirstswabandthedrysecondswabcollectingtheremainderofthesample.
Swabs can be comprised of variousmaterials including cotton or synthetic variations.
Swabbing can also be performed using gauze‐like materials, disposable brushes, or
commercially available biological sampling kits (Lauk and Schaaf 2007). Another standard
collection technique involves taking cuttings of the area of interest such as a biological fluid
fromclothing;howeverthisdestroystheintegrityoftheevidence.Adhesivetapeliftsarealso
usedonavarietyofsurfacestocollecttraceevidencethatmaycontainhumanDNA.
Ifsamplesarecollectedandarenotprocessedimmediately,theyshouldbeallowedto
dry to prevent fungal or bacterial growth. Evidentiary samples should not be immediately
sealed in plastic, which can result in microbial growth and cause degradation of the DNA.
Typically,swabsorcuttingsareplaced inbreathablecontainersmadeofpaperorcardboard.
Storingcollectedevidenceincool,dryenvironmentsalsominimizessampledeterioration(Lee
andLadd2001).
Onceasampleiscollected,thenextchallengeistoremovethebiologicalcellsfromthe
collectionmatrices.Cellscanbeextracteddirectlyfromthecollectiondevice,asisthecasein
mostcuttingsorcottonswabs,byplacingthecollectorintoanextractionbuffer.Inthecaseof
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
21
tape lifts, swabs moistened with solutions such as xylene can be utilized to dissolve the
adhesive,collectingasmuchmaterialfromthetapeaspossiblepriortosubjectingtheswabto
extraction.
In some cases,DNA from the collected cells should not be extracteddirectly and the
biological material should be removed first for examination. When using cotton swabs to
collectmaterial,therecanbeproblemsremovingbiologicalmaterialfromthecottonmatrix;as
the cotton swab dries after collection, the biological material can adhere to the swab. For
example, due to the saccharic composition of the spermatocyte membrane, spermatocytes
stick to solid supports, especially cotton (Lazzarino 2008). In order to release themaximum
amountofmaterialfromtheswabs,avarietyofbuffershavebeentestedandcomparedtothe
standarddifferentialextractionbuffer.Useofdetergentssuchas1‐2%sodiumdodecylsulfate
(SDS) has shown to increase sperm cell recovery (Norris 2007). Also, the addition of low
amounts of cellulase has shown to releasemore epithelial and sperm cells from the cotton
swabmatrixthanbufferelutionalone(Voorhees2006).
Problematic Samples. There can be many challenges to obtaining STR profiles from
biological materials including low quantity or quality of DNA. Low copy number samples
(containing less than50picogramsofDNA)aswell as lowquality,degraded samples require
highlyefficientcollection,extraction,andamplificationprocedures.Thesesamplesareseenin
avarietyofforensicevidenceincludingtouchevidenceandagedsamples.
PCR inhibitors are another challenge and must be eliminated before downstream
applications can be performed. Common inhibitors are indigo dyes from denim, heme from
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
22
blood,humicacidfoundinplantsandsoil,andcollagenfoundinvarioustissues.Themajority
of these inhibitors are effectively eliminated using silica‐based DNA extraction methods or
additionalpurificationwithchargeorsizeexclusioncolumns. Thepresenceof inhibitorscan
bedetectedbyperformingPCRwithinternalpositivecontrols.Ifpresent,someinhibitorscan
beneutralizedbyvarioustreatmentsincludingsodiumhydroxidewashesorfurtherpurification
withMilliporeMicroconYM®columns.
IntegrationofEvidenceCollectionwithPost‐CollectionProcessing.Theneedtoreconcile
the“realworld”requirementsofsamplecollectionwiththemicrofluidicrequirementsofafully
integrated microfluidic DNA processing biochip can be referred to as the “macro‐to‐micro
interface”orthe“world‐to‐chipinterface”(Fredrickson2004).Muchofthereportedresearch
onaddressingthisinterfaceisfocusedonresolvingthemismatchbetweenthemacrofluidicand
microfluidic volumetric requirements, but little of no research concerning the reconciling of
specific forensic sampling requirements and formats with microfluidic devices has been
reported.
The(non‐forensic)volumetricmismatchhasbeencommerciallyaddressedbyAgilentin
theBioanalyzer2100bytheuseofacapillarytoaspiratesamplesfromamicrotiterplatetoa
chip for enzyme assays (Lin 2003). Similarly, Gyros has developed a capillary dispenser for a
LabCDsystemwheresamplesareaspiratedfromawellplateintoadispensingnozzleandthen
directedupwardsontoarotatingdevice(Jesson2003).Thesedevices,however,donotaddress
the format incompatibility of collected forensic samples—particularly on the commonly used
collectiondevicesbasedonswabs.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
23
D. RequirementsofaSampleCollectionSystem
Sample collection and initial sample processing are critical to the development of a fully
integrated microfluidic system for STR typing. These steps are macrofluidic and must be
conducted such that theendproduct—DNA in solution—canbe transferred toamicrofluidic
biochip.TheevidencecollectiondeviceandSmartCartridgedevelopedaspartofthisresearch
addresstheformatandvolumetricmismatchthatexistsbetween“realworld”forensicsamples
that are collected and requirements for analysis by microfluidic devices, a mismatch that
precludes direct insertion of biological samples collected by validated forensic collection
methods directly into a microfluidic biochip. This research takes advantage of a series of
forensic advances in sample collection, elution, and processing of difficult samples. These
forensic advanceswere combinedwithBode’sexperiencewith sample collectionandelution
andNetBio’sexperiencewithmicrofluidicmanipulationtodevelopasamplecollectiondevice
andSmartCartridgecompatiblewiththemicrofluidicbiochiprequiredforafullyintegratedSTR
typingsystem.
Thecollectionsystemwasdesignedtopossessthefollowingproperties:
Ease of use—The evidence collection system must be capable of operation by law
enforcementagentswithminimaltraininginmolecularbiologicalorchemicalanalysisandmust
protectagainst inter‐andintra‐runsample‐to‐samplecontamination. Thesystemwillbepre‐
filledwithallreagentstoreducelaborrequiredtooperatethesystem.
• SampleTypes—Thesystemmustacceptavarietyofsampletypes.Forthisproject,we
havefocusedonliquidanddriedblood,buccalcells,epithelialcells,andsalivasamples.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
24
• Sensitivity—The systemmust be capable of accepting and processing sampleswith a
highdynamicrangeofcellcounts.
• Compatibility—Both the evidence collection device and Smart Cartridge must be
compatiblewithallrequiredreagents.
• Integration—Allevidencecollectionmatricesandsampleprocessingreagentsmustbe
compatiblewithtransferusingmicrofluidicprinciples.
• Timetoanswer—Thesampleprocessingproceduremusttakenomorethan5minutes.
IV. ResearchDesignandMethods
A. OverviewofResearchDesign
Thegoalofthisresearchwastodevelopasamplecollectionandprocessingsystemconsisting
ofanevidencecollectiondeviceandasampleprocessingdevice termeda“SmartCartridge.”
TheSmartCartridgewilllyseandsolubilizetheinputsample,purifyDNA,andconcentratethe
purifiedDNA. The concentratedDNAwill thenbe transferredautomatically toNetBio’s fully
integrated microfluidic biochip for further DNA purification, DNA quantitation, STR
amplification, and separation and detection. Figures 1 and 2 show schematics of the entire
process,fromevidencecollectiontogenerationofanSTRprofile.
Thespecificobjectivesofthisresearchwereto:
Develop an evidence collection device that provides for ease of biological sample
collection, protection, and documentation. This includes the evaluation and selection of a
collectionmatrix thatprovidesefficiencyofbiological samplecollectionoverawide rangeof
sampletypesandcompatibilitywithsubsequentsampleprocessingincludingsamplelysis,DNA
solubilization, and concentration in the Smart Cartridge, and also subsequent downstream
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
25
sampleprocessing in the integratedmicrofluidicbiochip.Theevidencecollectiondevicemust
protectthesampleforstorageandtransportandpreventsample‐to‐samplecontamination.
Develop a Smart Cartridge that accepts the evidence collection device, performs
processing steps, and transfers the resulting DNA solution to the microfluidic biochip. This
sample processing cartridge will incorporate sample lysis, DNA solubilization, and
concentration.Appropriatemechanicalandfluidicinterfacesforcouplingtoboththeevidence
collectiondeviceandthebiochipwillalsobeincorporated.
B. MaterialsandMethods
1. MockCaseworkandReferenceSamples.
Buccal cell samples were obtained bymoving the swabs up and down on the inside
cheekofahumansubject.
FreshwholebloodcontainingEDTAasanticoagulantwasobtainedonice.
Driedbloodsamplesonswabwerepreparedbyallowingthebloodtodryovernight.
Salivawascollectedbyexpectoratingintoa50mlfalcontube.
Epithelial cell sampleswere collected by rubbing the swab heads on the palm and in
betweenthefingersofahumansubject.
A set of reference and mock casework samples was also prepared by Bode. These
samplesconsistedobuccal,blood,driedblood,saliva,andepithelialcells thatwerecollected
anddriedonSecurSwabs.
2. VortexExtractionofDNAfromSwabs.
Swabswereplacedina2mLmicrocentrifugetube.Theswabheadswereseparatedfromthe
shaft by cutting them off with scissors. 500 µL of NetBio lysis solution was added to the
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
26
microcentrifugetubeandthetubewasvortexedfor5seconds.Theswabheadsweremanually
removedusingapairofcleantweezersanddiscarded.
3. MechanicalAgitationforExtractionofDNAfromSwabs.
A tube containing lysis solutionwas placed in a custombuiltmechanical agitation system.A
SecurSwabwasinsertedintotheagitationsystemandattachedtothesystembytheswabcap.
Aminiaturizedmotorwasused togeneratevigorousvibrationsand this vibrationalenergy is
coupled to the swab tip through the cap and shaft. Mechanical agitation was applied
immediatelyafterinsertionintothesystem.
4. MicrofluidicBiochipPurification.
ThemicrofluidicbiochipcontainsapurificationfilterforDNAbindingsealedwithinachamber.
Twomicrofluidicchannelsleadtothechamber,onetoallowflowofreagentstothefilterand
theother to remove reagents from the filter. The lysis solutionmixturewas loaded into the
input port and pneumatically driven through the purification filter. This was followed by
pneumaticallydrivingwashbufferthroughthefilter.BoundDNAwaselutedfromthefilterby
pneumaticallydrivingelutionbufferthroughthefilter.
5. Automated Microfluidic Extraction and Purification of DNA from Swabs in
SmartCartridge.
The Smart Cartridge is comprised of reagent chambers for holding preloaded solutions and
processchambers.OnechamberisusedtoholdthecottonswabwiththeDNAsample;fourof
the chambers are prefilled with purification reagents. The final two chambers are used for
holding solutions during the process. The SC biochip is comprised of a network of channels,
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
27
chambers, flowcontrolelements,andapurification filter.Anautomatedscriptpneumatically
manipulatessolutionswithintheSCtoextractDNAfromacottonswabthatisinserted.
6. STRAmplificationReaction.
MultiplexPCRreactionswereperformedwiththeAmpFℓSTR®Identifiler®PCRAmplificationKit
(AppliedBiosystems,FosterCity,CA).The7µLPCRreactionandcyclingprotocolwasprepared
as described in (Giese 2009). Amplification of 16‐sample in the microfluidic biochip was
completed in approximately 17minutes and the amplifiedproductsweremanually retrieved
fromtheindividual.
7. STRseparationanddetectioninstrumentation
Amplified products were separated and detected using Genebench‐FX. This instrument was
developedandoptimizedspecificallyforSTRanalysis.
V. DisseminationStrategy
Theresultsofthisresearchhavebeendisseminatedasfollows:
Presentations. Thisworkhasbeenpresentedinanoralpresentationentitled“Sample
CollectionSystemforDNAAnalysisofForensicEvidence”attheAmericanAcademyofForensics
Scientists62ndAnnualScientificMeetinginSeattle,WAonFeb22‐27,2010.
Duringandafterthecourseoftheproposedwork,wehavediscussedourprogressand
askedforsuggestionsandcommentsfromseveralforensicscientists.Wehavemaintainedand
expanded contact with the NFSTC and several state crime labs. Our hope is to continue
developing a close relationshipwith these groups to ensure that our development goals are
consistentwiththeneedsoftheforensiccommunity.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
28
VI. ImplicationsforCriminalJusticePolicyandPractice
The availability of an STR typing instrument that combines DNA extraction and purification,
amplification, and separation into a single, easy to operate instrument would represent a
substantialadvanceinforensicDNAanalysis.Significantburdensinsettingupandoperatinga
forensic DNA analysis laboratory are the costs of dedicated rooms to prevent PCR
contamination,automatingtheprocedures(eitherthroughroboticsordedicatedtechnicians),
and validating and re‐validating individual instruments and the entire series of laboratory
processes. A fully integrated instrument has the potential to be faster,more sensitive, less
susceptible to contamination, less costly, and less labor‐intensive than currently available
technologies.
Indevelopinga fully integrated system for STR typing, evidence collectionand
initial sample processing are critical steps, and, as such,must be incorporated into any fully
integratedmicrofluidicSTRtypingsystem.Thisresearchhasresulted inthedevelopmentofa
practical system for DNA analysis of forensic evidence, a system that is faster and performs
better than currently available technologies. The research represents an important step
towardsmakingfullyintegratedforensicDNAanalysisareality.
Finally, a fully integrated microfluidic STR typing instrument would offer forensic
scientistsnewcapabilitiesnotpossiblewithconventionalcapillaryinstrumentation.NetBiohas
alreadyruggedizedGenebench‐FXanddemonstratedthatitwithstandstherigorsoftransport
to and operation at the crime scene. Developing an evidence collection device and Smart
Cartridge capableof collecting sample andprocessingDNA froma varietyof biological fluids
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
29
associated with a variety of evidence will also contribute towards producing reliable DNA
profiles in a cost‐effective manner at the crime scene. The use of a field portable, fully‐
integratedDNAanalysis instrumentwill allow rapid generationof aDNAprofilewhich could
directlyaffecttheapprehensionofasuspectandtherebyreducecrimeandimprovethepublic
safetywhiledecreasingthetimeandcostsofcriminalinvestigations.
VII. ResultsandDiscussions
A. Evidencecollectionmedia
TheobjectiveofthisworkwastodeterminethebestmatrixtocollectandextractDNAfroma
varietyof cells associatedwithdifferentPCR inhibitors, varietyof surfaces, and fromvarying
cellnumbers.Aseriesofcollectionmatricesconsistingofnatural fiber (cotton)andsynthetic
matrices(modifiedcellulose,foam,Nylon,PolyesterandRayon)wereselectedforevaluation.
Fourbuccalsampleswerecollectedfromeachdonorwitheachofthecollectionmatrices.The
collection matrices were assessed based on DNA collection and extraction efficiency, lysate
retentionvolume,anduseinforensicapplications.
DNAcollectionandextractionefficiency.DNAwasextractedandpurifiedfromeachof
the 112 samples using a set of reagents thatwere optimized for purification of buccal swab
samples, compatibility with biochip DNA purification, and minimal process time. DNA yield
measuredbyUVanalysis showedthatawide rangeofDNA (300‐1900ng)wasextractedper
swabwithhigh inter‐ and intra‐ donor variation. The average recoveredDNAacross showed
thatallswabtypesperformedsimilarly(Figure16).
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
30
Figure16.DNArecoveredfromvariousswabmatrices.
Use in forensic applications. A survey of swab collection matrices used in forensic
applicationssuggestedthatcottonwasmostcommonlyusedforcrimescenesamplecollection
andforreferencesamplecollection.
Cotton was the matrix selected for incorporation into the sample collection device
becauseofitswidespreadacceptanceandusewithintheforensicscommunityandcomparable
extractionefficiencyrelativetoothertestedcollectionmatrices.
Commerciallyavailableswabswithvariousswabheadconfigurationswereassessedfor
useinthefullyintegratedsystem.Easeofuseforbuccalcellcollectionwasevaluatedbyhaving
eachdonorswabtheinsideofthecheekwiththecandidateswab.Furthermore,eachcandidate
swabwasalsousedtocollectmockcrimescenesampleswiththecandidateswabs.Subjective
feedback frombuccal cell donorsand crime scene collectorswereevaluated toestablish the
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
31
SecurSwab (Bode) as the easiest, safest, and most convenient for use; the SecurSwab was
designedforcollectionofDNAsamplesorforensicevidenceandiseasiertoopen,holdanduse.
Thereproducibilityoftheevidencecollectorwasevaluatedbycollecting4buccalswab
samplesfromeachofnineteendonors.Twoswabswerecollectedfromtheleftcheekandtwo
from the right cheek. DNA from each swab was extracted and purified. Purified DNA was
quantifiedbyUVspectrometry.Data inFigure17showsthatDNAyieldedfromthe76swabs
rangefrom200ngto1850ng,andthattheswabtoswabvariation(%CV)forthe76samples
was42%.Thecoefficientofvariation (%CV)withineachdonorranges from20to65%.These
results show that DNA collected from buccal swabbing is highly variable, however, the
minimum amount that is collected by the selected evidence collection device is more than
sufficienttogeneratethe1ngofDNArequireforSTRanalysis.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
32
Figure17.DNAyieldfrombuccalswabscollectedacross19donors.
Figure 18 shows the results of an STR analysis generated by amplifying 1 ng of DNA
templateandtheAmpFLSTRProfilerPrimerset,andseparatinganddetectingwithGenebench.
TemplateDNAwascollectedfrombuccalcellscollectedbytheevidencecollectiondeviceand
extractedandpurifiedwithNetBioreagentsfollowingvortexextraction.ThefullSTRprofilethat
isdemonstratedconfirmsthecompatibilityof theevidencecollectiondeviceand thereagent
setforSTRanalysis.
The performance of NetBio’s microfluidic purification biochip was evaluated by
generating sets of lysates from swabs, by vortex extraction, and purifying with NetBio’s
Microfluidic Biochip Purification protocol. The purified DNA was quantified and 1 ng of the
purifiedsolutionsubjected toamplification.The full STRprofile (Figure19)demonstrates the
compatibilityofthereagentsetwithmicrofluidicbiochippurifications.
B. ChemicalLysis
Biologicalmaterialcollectedwiththeevidencecollectoris lysedandDNAissolubilized
within the Smart Cartridge. The solubilized DNA is then microfluidically transported for
subsequentDNApurification.It isimportanttomaximizethequantityofDNArecoveredfrom
the sample collection device, particularly in casework sampleswhere the amount of sample
collectedmaybelimited.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
33
Figure18.STRprofileofDNApurifiedfrombuccalswabsampleswithNetBio’soptimizedreagentsetfollowingvortexextraction.
Figure19.STRprofileofDNAmicrofluidicallypurifiedinbiochipfrombuccalswabsampleswithNetBio’soptimizedreagentstefollowingvortexextraction.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
34
Initial work on cell lysis strongly suggested that chaotropic salts are effective lysis
reagents and are compatible with microfluidic biochips. Optimization of the chemical lysis
reagent components was performed to maximize DNA efficiency for sample types including
blood, buccal, touch, and saliva samples, and to minimize process time. The optimization
includes a systematic variation of the lysis reagent components, wash solution components,
andelutionbuffercomponents.Tominimizethenumberofiterations,anexperimentalmatrix
wasdeveloped to determine the effect of each componentonDNAyield andpurity, and an
optimal reagent set. Three formulations (NetBioA,B andC)wereevaluatedby collecting12
buccalswabsamplesfromthesamedonorandpurifyingtheDNAfromthebuccalswabswith
each of the formulations. The DNA yielded was quantified by spectrometry shows that
formulationCgenerates thehighestaverageDNAyieldof900ngcomparedwith850ngand
650ngforformulationsBandArespectively.
C. SmartCartridgeProcessStep2:SelectionofSilicaFiberMembrane
DNAConcentrationandpurification
Thebindingandelutionpropertiesofchaotropicsalt/silicamediasystemsallowDNAto
be concentrated from relatively dilute lysates. An added advantage of using the bind/elute
method for DNA concentration is the opportunity towash theDNAwhile it is bound to the
membrane,allowingfortheremovalofparticulatesandcrudecelldebrispriortoelution.Silica
basedbind/elutemediaareavailable in several formats including silica fibermembranesand
silicabeads.Useofthemembraneformatrequiresthemembranetobepositionedatafixed
site of the Smart Cartridge,with the attendant advantages of lowermanufacturing cost and
ease of containment of the silica media. The bead‐based format provides the flexibility of
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
35
movingthebeadsto leavebehindthedebrisassociatedwiththecollectedsamplesand lysed
cells. This format is relatively costly to implement, and the beads are somewhat difficult to
contain.Accordingly,performanceevaluationshavebeenconductedonmembranebasedsilica
fibermedia.
D. SmartCartridgeDevelopment
An initial Smart Cartridge was designed and fabricated based on the optimized
conditionsdiscussedintheprevioussections.Thisplatformformedthebasisfortheevaluation
andoptimizationofthethreeprocessstepsinseries.TheinitialSmartCartridgewasdesigned
withthefollowingfeatures:
• Flexibility – ability to rapidly implementmodifications toboth the flowconfigurations
andprotocols
• Linked processing chambers – the lysis and DNA concentration are in direct
communication.
• Optical imaging – the device functions with optical cameras to allow the research
monitoring of reactions within each chamber and fluid flow within and between
chambers.
• Fluidicdrive–pneumaticandfluidicpumpsarecoupledtotheSCtodriveprocessfluids.
• Computer‐controlled – the Smart Cartridge is computer‐controlled to allow the
automatedexecutionofaprocessbyscript
A single sample smart cartridge design and associated extraction and concentration
processflowshasbeendeveloped.ArenderingofasingleSC(Figure20)showsamacrofluidic
componentforacceptingaBodeSecurSwab,storingreagents,andprocessing.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
36
Figure20.Renderingofasinglesamplesmartcartridge.
E. TandemEvaluationandOptimizationoftheThreeProcessSteps
TheSCperformancewascharacterizedusingbuccalswabs.DNAfromtheprocesswas
quantifiedbyUVabsorbanceshowsthatbetween200–1100ngofDNAisyieldedfromthe16
swab samples (Figure 21). This yield of DNA is comparable to manually processing and
demonstratesthesuccessofautomatedsmartcartridgeprocess.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
37
Figure21.DNAextractionperformanceoftheextractioncomponentoftheSC.
F. TestingoftheEvidenceCollectionDeviceandSmartCartridge
Theabilityof thecompletedevidencecollectiondeviceandSmartCartridge tocollect
samplesandtoperformthethreeprocessstepswastested.Aseriesofmockcaseworksamples
werecollectedandprocessedasfollows:
Whole blood. Two evidence collectors were used to swab the blood directly from a
ceramicsurface.OneevidencecollectorwasprocessedbyautomatedscriptintheSCandthe
other was processed following a control Qiagen protocol. DNA from the SC process was
amplifiedandseparatedanddetectedonGenebench.Figure22showsafullSTRprofileofthe
SC processed sample. This profile is similar to that generatedwith the control protocol and
demonstratesthecompatibilityofthesamplecollectionsystemforwholebloodsamples.
Driedblood.TwoevidencecollectorswerewetwithDIwaterandusedtoswabthedried
bloodfromthesurface.Oneevidencecollectorwasprocessedbyautomatedscript intheSC,
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
38
andtheotherevidencecollectorwasprocessedfollowingtheQiagenprotocol.DNAfromthe
SCprocessandtheQiagenprocesswasamplifiedandseparatedanddetectedonGenebench.
Figure 23 shows a full STR profile of the SC processed sample. This profile is similar to that
generated with the control protocol and demonstrates the compatibility of the sample
collectionsystemfordriedbloodsamples.
Figure22.STRprofileofDNAextractedandpurifiedintheSCfromwholeblood.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
39
Figure23.STRprofileofDNAextractedandpurifiedintheSCfromdriedblood.
Salivasamples.Pooledsalivasampleswerecollectedandspottedonaceramicsurface.
Twoevidencecollectorswereusedtoswabthesalivafromthesurface.Theevidencecollectors
werethensecuredintheircollectiontubesovernight.Oneevidencecollectorwasprocessedby
automated script in the SC, and the other evidence collector was processed following the
Qiagen protocol to serve as controls. DNA from the SC process and theQiagen processwas
amplifiedandseparatedanddetectedonGenebench.Figure24showsafullSTRprofileofthe
SCprocessedsample.
Buccalsamples.Buccalsampleswerecollectedbyswabbingtheinsideofthecheekand
then secured in their collection tubes overnight. One evidence collector was processed by
automated script in the SC, and the other evidence collector was processed following the
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
40
Qiagen protocol to serve as controls. DNA from the SC process and theQiagen processwas
amplifiedandseparatedanddetectedonGenebench.Figure25showsafullSTRprofileofthe
SCprocessedsample.
Figure24.STRprofileofDNAextractedandpurifiedintheSCfromsaliva.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
41
Figure25.STRprofileofDNAextractedandpurifiedintheSCfrombuccalcells.
VIII. Conclusions
This report presents work demonstrating successful completion of all technical
milestonesofthisresearchproject.
An evidence collection device has been developed for ease of biological sample
collection,protection,anddocumentation.Theevaluationofvarietysamplecollectionmatrices
including cotton, modified cellulose, foam, nylon, polyester and rayon based on collection
efficiency and current use in forensics applications resulted in the selection of cotton.
Commerciallyavailableswabswithvariousswabheadconfigurationswereassessedforeaseof
useandresultedintheselectionoftheBodeSecurSwabforincorporationintothesystem.
A Smart Cartridge was developed to accept the evidence collection device, performs
processing steps, and transfers the resulting DNA solution to the microfluidic biochip. The
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
42
optimization of a lysis and concentration reagent set, and incorporation of mechanical lysis
resultedintherapidandefficientlysisofbuccalcellsamplestoyield200–1400ngofDNA.A
Smart Cartridge was fabricated by initially designing and fabricating the macrofluidic and
microfluidiccomponentsoftheSC,andprocessingofbuccalsamplesdemonstratedthatDNAof
between200–1100ngofDNAisyieldedbytheautomatedSCprocess.
The system (evidence collector and SC) was tested by collecting mock crime scene
samplesincludingwholeblood,diedblood,saliva,andepithelialcellsampleswiththeevidence
collectorandprocessingwiththeSCbyautomatedscript.STRprofilesofDNAgeneratedbythe
systemweresimilartothoseproducedbyconventionalextractionprotocols.
Theresultsasawholedemonstratethesuccessfuldevelopmentofanevidencecollector
and SmartCartridge capableofpurifyingDNA froma varietyof sample types and substrates
relevanttotheforensicssciencescommunity.Thesuccessfuldevelopmentofthismodulealso
represents the completion of another critical step towards the implementation of a fully
integratedinstrumentthatwillgenerateanSTRprofilein45minutesfromsampleintroduction
withminimal operating requirements. The integrated instrument will process 16 samples in
parallelanddramaticallyreducethecosts(includinglabor,space,andvalidation)ofsettingup
andoperatingaDNAlab.TheevidencecollectiondeviceandSmartCartridgearedesignedsuch
that the systemwill be easy to operate and compatible with both forensic andmicrofluidic
requirements.
IX. References
Bienvenue,J.etal.(2010).AnIntegratedMicrofluidicDeviceforDNAPurificationand
PCRAmplificationofSTRfragments.ForensicSciIntGenet4(3):178‐86.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
43
Fredrickson, C. K. and Z. H. Fan (2004). "Macro‐to‐micro interfaces for microfluidic
devices."LabChip4(6):526‐33.
Giese, H., R. Lam, et al. (2009). "Fast multiplexed polymerase chain reaction for
conventionalandmicrofluidicshorttandemrepeatanalysis."JForensicSci54(6):1287‐96.
Hopwood, A. et al. (2010). Integrated Microfluidic System for Rapid Forensic DNA
Analysis:SampleCollectiontoDNAProfile.AnalChem2010,82:6991‐9.
Jesson,G.etal.(2003).MicroTotalAnalysisSystems,ed.M.A.Northrup,K.F.Jensen
andD.J.Harrison,pp.155–158.
Lauk,C.andJ.Schaaf(2007)."ANewApproachfortheExtractionofDNAfromPostage
Stamps."ForensicScienceCommunications9(1).
Lazzarino,M.F.,A.ColussiandM.M.Lojo (2008)."DNARecovery fromSemenSwabs
withtheDNAIQSystem."ForensicScienceCommunications10(1).
Lee,H.C.andC.Ladd(2001)."Preservationandcollectionofbiologicalevidence."Croat
MedJ42(3):225‐8.
Lee,H.C.,C.Ladd,C.A.ScherczingerandM.T.Bourke(1998)."Forensicapplicationsof
DNA typing:part2: collectionandpreservationofDNAevidence."AmJForensicMedPathol
19(1):10‐8.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.
SampleCollectionSystemforDNAAnalysisofForensicEvidence NetBioInc.NIJAward2008‐DN‐BX‐K010
44
Leemans,P.(2006)."EvaluationandmethodologyfortheisolationandanalysisofLCN‐
DNAbeforeandafterdactyloscopicenhancementoffingerprints."IntCongressSer1288:583‐
585.
Lin, Y.W.,M. J. Huang andH. T. Chang (2003). "Analysis of double‐strandedDNA by
microchip capillary electrophoresis using polymer solutions containing gold nanoparticles." J
ChromatogrA1014(1‐2):47‐55.
Norris,J.V.,K.Manning,S.J.Linke,J.P.FerranceandJ.P.Landers(2007)."Expedited,
chemicallyenhancedspermcellrecoveryfromcottonswabsforrapekitanalysis."JForensicSci
52(4):800‐5.
Voorhees,J.C.,J.P.FerranceandJ.P.Landers(2006)."Enhancedelutionofspermfrom
cottonswabsviaenzymaticdigestionforrapekitanalysis."JForensicSci51(3):574‐9.
This document is a research report submitted to the U.S. Department of Justice. This report has not been published by the Department. Opinions or points of view expressed are those of the author(s) and do not necessarily reflect the official position or policies of the U.S. Department of Justice.