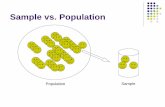
Sample classification using Microarray Data. AB We have two sample entities malignant vs. benign...
-
Upload
shonda-hines -
Category
Documents
-
view
221 -
download
1
Transcript of Sample classification using Microarray Data. AB We have two sample entities malignant vs. benign...
A B
We have two sample entities
• malignant vs. benign tumor
• patient responding to drug vs. patient resistant to drug
• etc...
A
B
What about differences in the profiles?
Do they exist?
What do they mean?
... and the situation looks like this:
A
B
What is the problem with high dimensional dependent data?
Back to the statistical classification problem:
Spent a minute thinking about this in three dimensions
Ok, there are three genes, two patients with known diagnosis, one patient of unknown diagnosis, and separating planes instead of lines
• Problem 1 never exists!• Problem 2 exists almost
always!
OK! If all points fall onto one line it does not always work. However, for measured values this is very unlikely and never happens in praxis.
In summary:
There is always a linear signature separating the entities ... a biological reason for this is not needed.
Hence, if you find a separating signature, it does not mean (yet) that you have a nice publication ...
... in most cases it means nothing.
Take it easy!
There are meaningful differences in gene expression ... we have build our science on them ... and they should be reflected in the profiles.
In general a separating linear signature is of no significance at all.
There are good signatures and bad ones!
Strategies for finding good signatures ...
... statistical learning:
1. Gene Selection
2. Factor-Regression
3. Support Vector Machines (machine learning)
4. Informative Priors (Bayesian Statistics)
5. .....
What are the ideas?
Gene selectionWhy does it help?
When considering all possible linear planes for separating the patient groups, we always find one that perfectly fits, without a biological reason for this.
When considering only planes that depend on maximally 20 genes it is not guaranteed that we find a well fitting signature. If in spite of this it does exist, chances are better that it reflects transcriptional disorder.
If we require in addition that the genes are all good classifiers them selves, i.e. we find them by screening using the t-Score,
finding a separating signature is even more exceptional.
Gene selectionWhat is it?
Choose a small number of genes … say 20 … and then fit a model using only these genes.
How to pick genes:-Screening ( i.e. t-score)
single gene association
-Wrapping
multiple gene association
1 . choose the best classifying single gene g1
2. choose the optimal complementing gene g2
g1 and g2 are optimal for classification
3. etc ….
How many genes
- All linear planes
- All linear planes depending on at most 20 genes
- All linear planes depending on a given set of 20 genes
High probability for finding a fitting signature
Low probability for finding a fitting signature
High probability that a signature is meaningful
Low probability that a signature is meaningful
How you find them
-Wrapping:multiple gene association
-Screening: Single gene association
Tradeoffs
Flexible model
Smooth model
High probability for finding a fitting signature
Low probability for finding a fitting signature
High probability that a signature is meaningful
Low probability that a signature is meaningful
More Generally
Factor-Regression
For example PCA, SVD ...
Use the first n (3-4) Factors only
... does not work well with expression data, at least not with a small number of samples
)( |1 0][ ii i x βYP
Support Vector Machines
Fat planes: With an infinitely thin plane the data can always be separated correctly, but not necessarily with a fat one.
Again if a large margin separation exists, chances are good that we found something relevant.
Large Margin Classifiers
First:Singular Value Decomposition
X
FDAE
Data
Loadings Singular values
Expression levels of super genes, orthogonal
matrix
Keep all super-genes:The Prior Needs to Be
designed in n Dimensions
• n number of samples• Shape?• Center?• Orientation?• Not to narrow ... not to wide
Not to Narrow ... Not to Wide
Auto adjusting model
Scales are hyper parameters with their own priors
What are the additional assumptions
that came in by the prior?
• The model can not be dominated by only a few super-genes ( genes! )
• The diagnosis is done based on global changes in the expression profiles influenced by many genes
• The assumptions are neutral with respect to the individual diagnosis
All models confine the set of possible signatures a priori; however, they do it in different ways.
Gene selection aims for few genes in the signature
SVM go for large margins between data points and the separating hyper-plane.
PC-Regression confine the signature to 3-4 independent factors
The Bayesian model prefers models with small weights
(a la ridge-regression or weight decay)
A common idea behind all models ...
... and a common problem of all models:The bias variance trade
offModel Complexity:
- max number of genes
- minimal margin
- width of prior
-etc
How much regularization do we need?
- The Bayesian answer: What you do not know is a random variable ... regularization becomes part of the model
- Or model selection by evaluations ...
Model Selection with separate data
Training Test
100 50 50
Selection
Split of some samples for Model Selection
Train the model on the training data with different choices for the regularization parameter
Apply it to the selection data and optimize this parameter (Model Selection)
Test how good you are doing on the test data (Model Assessment)
10 Fold Cross-Validation
Train TrainTrain TrainSelect
Train TrainTrain TrainSelect
...
...
Chop up the training data (don‘t touch the test data) into 10 sets
Train on 9 of them and predict the other
Iterate, leave every set out once
Select a model according to the prediction error (deviance)
Leave one out Cross-Validation
Train TrainTrain TrainSelect
Train TrainTrain TrainSelect
...
...
Essentially the same
But you only leave one sample out at a time and predict it using the others
Good for small training sets
1
1
Model Assessment
How well did I do?
Can I use my signature for clinical diagnosis?
How well will it perform?
How does it compare to traditional methods?
The most important thing:
Don‘t fool yourself! (... and others)
This guy (and others) thought for some time he could predict the nodal status of a breast tumor from a profile taken from the primary tumor!
... there are significant differences. But not good enough for prediction
(West et al PNAS 2001)
DOs AND DONTs :
1. Decide on your diagnosis model (PAM,SVM,etc...) and don‘t change your mind later on
2. Split your profiles randomly into a training set and a test set
3. Put the data in the test set away.
4. Train your model only using the data in the training set
(select genes, define centroids, calculate normal vectors for large margin separators,perform model selection ...)
don‘t even think of touching the test data at this time
5. Apply the model to the test data ...
don‘t even think of changing the model at this time
6. Do steps 1-5 only once and accept the result ...
don‘t even think of optimizing this procedure
The selection bias
- You can not select 20 genes using all your data and then with this 20 genes split test and training data and evaluate your method.
-There is a difference between a model that restricts signatures to depend on only 20 genes and a data set that only contains 20 genes
-Your model assessment will look much better than it should