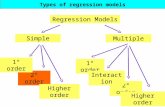
Regression models I
Transcript of Regression models I

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Faculty of Health Sciences
Regression modelsQuantitative covariate, Quantitative outcome, 23-4-2012
Lene Theil SkovgaardDept. of Biostatistics
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Quantitative covariate, Quantitative outcome
PKA & LTS, Sect. 4.1, 4.1.1Simple linear regression
I The assumption of linearityI Estimation and testingI Confidence and prediction limitsI Model checks and diagnosticsI Transformation
Home pages: http://biostat.ku.dk/~pka/regrmodels12E-mail: [email protected]
2 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Quantitative covariates
I AgeQuantitative outcome: Systolic blood pressure
I Body mass indexQuantitative outcome: Vitamin D status
3 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Quantitative covariate, no grouping
In principle:A separate mean value for all distinct values of the covariate xIn practice:A more parsimoneous model,combining the mean values in a smooth way
Smoothing:I Take local averagesI Combine these with a smooth curve,
to give a hint to the form of the relationship (if any)Every smooth curve can be approximated by a straight line– at least locally
4 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
The straight line
Mathematical formulation: y = a + bx
InterpretationI Intercept a:
The expected outcome, when the covariate x is zeroI Slope b: The expected difference in y corresponding to a
one unit difference in x5 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Choice of scale
for the linearity assumption...depends on the nature of the outcome
The scale should ideally be unlimited, with no boundariesTraditional scales / link functions:
I Quantitative: Mean value of outcomeidentity link
I Binary: logit of the probability of some eventlogit link
I Survival times: logarithm of hazard ratecloglog link
6 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Vitamin D example
Scatterplot of vitamin D concentration versus body mass index forIrish women.
Does this look like a straight line?
7 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Model for vitamin D vs. BMI
yi : the vitamin D concentration for the ith individualxi : the corresponding body mass index
Model:E(yi) = mi = a + bxi
We call this a simple linear regressionI simple, because there is only one covariateI linear, because the covariate has a linear effect
8 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Method of Least Squares
derived from the likelihood principle:Minimize the residual sum of squares:
SSres =n∑
i=1(yi − yi)2 =
n∑i=1
(yi − a − bxi)2,
residuals here being the vertical distance from the observation yito the line, yi = a + bxi , i.e.
ri = yi − (a + bxi)
9 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Residuals in simple linear regression
10 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Estimation of slope
Maximum likelihood estimate:
b = −2.392
with estimated uncertainty SD(b) = 0.690
Good precision, whenI the residual variation sy|x is smallI the sample (n) is largeI the variation in the covariate (sx) is large
11 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Confidence interval for slope
For large samples, b will have an approximate Normal distribution.An approximate 95% confidence interval for b is therefore
b ± 1.96 · SD(b)
For moderate-sized samples, we usually replace 1.96 by theappropriate t-distribution quantile (≈ 2).Here, n = 41, so df = 41− 2 = 39, so the t-quantile is 2.023and CI=(-3.788, -0.996)
12 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Results for vitamin D
Intercept a = 111.05(18.40) uninteresting here (it often is)
Slope b = −2.392(0.690)Residual standard deviation sy|x = 17.91
a and sy|x are measured in the units of the outcome variable(nmol/l)b is measured in units of “outcome per explanatory variable”((nmol/l)/(kg/m2)).
13 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Test of zero slope
I Walds Test: W = (−2.392/0.690)2 = 12.02 ∼ χ2(1)P = 0.0005
I T -test: t = (−2.392/0.690) = −3.47 ∼ t(39)P = 0.0013
Strong evidence of a relationship between body mass index andvitamin D status
Causality? not necessarily...
14 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Reparametrization
Reparametrization of body mass index x, to x∗ = x − 25gives the very same model
E(yi) = a∗ + bx∗i = a + bxi
with the same slope, but with a new intercept
a∗ = a + 25b,
now interpreted as the expected level of vitamin D for an individualwith a body mass index of 25.Here, we get a∗ = 51.244(2.948), leading to a 95% confidenceinterval of (45.280, 57.207).
15 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Predicted values
The predicted value of vitamin D for the ith individual is given bythe straight line
yi = a + bxi
Confidence limits show the uncertainty in the estimated regressionline
Prediction limits show the (future) variation in the outcome, forgiven covariate (reference regions)
16 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Confidence limits for line
I Tell us where the line may also beI Limits become narrower when sample size is increased
17 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Prediction limits for line
I Tell us where future subjects will lieI Limits have approximately same width no matter the sample
size
18 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Check of model assumptions
I Linearity:Plot residuals vs. covariate, curves?
I Variance homogeneity:Plot residuals against predicted values, trumpet shape?
I Normality:Histogram, skewness?
I Quantile plot, hammock shape?
19 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
20 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
If assumptions fail
I Linearity:Transform or do non-linear regression
I Variance homogeneity:Transform
I Normality:Transform
Linearity is the most important assumption,unless the task is to construct prediction intervals!
Transformations in a little while....
21 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
The idea in Diagnostics
Assess the influence by leaving out one observation at a time
22 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Measures of influence
I Omit the ith individual from the analysisI Obtain new estimates, a(−i) and b(−i)I Compute deletion diagnostics:
dev(a)i = a − a(−i)dev(b)i = b − b(−i)
both normalized by the standard deviation of the estimateI Combine the squared deletion diagnostics into a single
diagnostic, Cook’s distance Cook(a, b)i .
23 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Deletion diagnostics
24 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Cooks distance
25 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Example: Cell concentration of tetrahymena
The unicellar organism tetrahymena grown intwo different media, with and without glucose
Research question:How does cell concentration x (number of cells in 1 ml of thegrowth media) affect the cell size y (average cell diameter,measured in µm).
Quantitative covariate : concentration xQuantitative outcome : diameter y
26 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Scatter plot
27 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Residual plot for naive linear regression
Note the curved shape indicating that linearity between celldiameter and concentration is (obviously) not appropriate.
28 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Power relationship
Suggested relationship between diameter (y) and concentration(x):
y = axb
Interpretation of the parameters:I a is a parameter denoting the cell size for a concentration of
x = 1, an extrapolation to the extreme lower end of theconcentration range as seen from the scatter plot
I b is ....When the concentration x is doubled, the diameter willincrease with a factor 2b
29 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Logarithmic transformation
Transforming the diameter (y) with a logarithm (here base 10)yields the theoretical relationship
log10(y) = log10(a) + b log10(x).
or in terms of observations:
E(y∗i ) = a∗ + bx∗i
where y∗i = log10(yi), x∗i = log10(xi),a∗ = log10(a) is the intercept and b the slope.
30 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Scatter plot on double logarithmic scale
looks pretty linear
31 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Model check for logarithmic analysis
32 / 34

u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Estimates for the multiplicative model
a∗ = 1.635(0.0202), CI=(1.5921, 1.6774)b = −0.0597(0.0041), CI=(−0.0684,−0.0510)
Back-transformingThe effect of a doubling of the concentration is estimated to2b = 2−0.0597 = 0.959, a 4.1% reduction of diameter.
Confidence limits: (2−0.0684, 2−0.0510) = (0.954, 0.965), i.e.between a 3.5% and a 4.6% reduction
33 / 34
u n i v e r s i t y o f c o p e n h a g e n d e pa rt m e n t o f b i o s tat i s t i c s
Estimated relation on original scale
34 / 34