Dedifferentiation of Human Epidermal Keratinocytes Induced ...
RCC with Sarcomatoid Dedifferentiation: New … 6, 2015 RCC with Sarcomatoid Dedifferentiation: New...
Transcript of RCC with Sarcomatoid Dedifferentiation: New … 6, 2015 RCC with Sarcomatoid Dedifferentiation: New...
Jose A. Karam, MD, FACSAssistant Professor
Department of Urology
November 6, 2015
RCC with Sarcomatoid Dedifferentiation: New Insights
Epidemiology of sRCC
• ~5% of all renal tumors are sarcomatoid
• More than 75% of patients with sRCC present with metastatic disease
• Median overall survival of less than a year
Not improved despite the advent of targeted therapies
Shuch B. The Oncologist. 2012
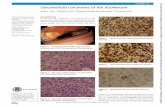
Histology 2 Components
• Epithelial component
– Clear cell, papillary, chromophobe, collecting duct, MTSCC, other
• Sarcomatoid component
Delahunt B. Am J Surg Path. 2013
Outline-Questions
• Can we identify sRCC on preoperative imaging?
• Can we identify sRCC on preoperative biopsy?
• Is sRCC different from non-sRCC on
– RNA level?
– DNA level?
– immune markers?
MRI-1
• Retrospective (2003-2009)• 9 patients • 2 radiologists reviewed preoperative MRI in patients who
later had nephrectomy– 5 clear cell
• Irregular or infiltrative morphology and heterogeneous T2 signal intensity and enhancement
• Internal necrosis in all cases Non-specific
Rosenkrantz AB. Clinical Imaging. 2011
MRI-2
• Retrospective (2004-2012)
• 11 patients with clear cell RCC with sarcomatoid elements
• Correlate preoperative MRI with pathology from nephrectomy
Takeuchi M. Clinical Imaging. 2013
MRI Findings
• sRCC showed significantly lower enhancement than the ccRCC part in each MRI phase
• Pseudocapsule disruption all cases
• Issue non-blinded, need comparator arm
Takeuchi M. Clinical Imaging. 2013
Current Ongoing Research
• Identification of sRCC using preoperative imaging
– PIs: Rivka Colen (Radiology) and Jose Karam (Urology), MD Anderson
FNA/Core biopsy (MD Anderson)
• 166 patients (1991-2007)
• FNA/core biopsy of the kidney before CRN
• 34 patients had sarcomatoid RCC at CRN
Only 4 patients (11.7%) had sarcomatoid elements detected on preoperative biopsy
Abel EJ. J Urology. 2010
FNA/Core biopsy (MD Anderson)
• 405 FNA or biopsies from 378 patients (1991-2007)– 239 from mets– 166 from primary tumors
• 76 patients had sarcomatoid RCC at time of surgery Only 7 patients (9.2%) had sarcomatoid elements
detected on preoperative biopsy
Abel EJ. BJU Int. 2012
FNA/Core biopsy (University of Wisconsin)
• 122 biopsies (cT2 or greater renal masses)– 46 standard– 76 multiquadrant (at least 4 separate areas)
• 2009 to 2014• 96 were RCC• Sensitivity to detect sarcomatoid RCC
– Standard: 2 of 8 (25.0%) (p=0.0062) – Multiquadrant: 13 of 15 (86.7%)
• 6 patients had some cores without sarcomatoid elements
Abel EJ. J Urol. 2015
Current Ongoing Research
• Identification of molecular signature specific for sRCC using needle core biopsy tissue (MD Anderson)
Experimental Plan
• 8 patients with ccRCC with sarcomatoid component (FFPE)
– RNA-seq (N=5)
– RT-qPCR
– IHC
Pal S. Mol Cancer Res. 2015
Experiment
Discovery cohort (cDNA microarrays)• 36 sarcomatoid RCC
– Epithelial component (clear cell) [E]
– Sarcomatoid component [S]
• 22 non-sarcomatoid RCC– All clear cell [E*]
Validation cohort (RNA-seq)• 7 sarcomatoid RCC
– Epithelial component (clear cell) [E]
– Sarcomatoid component [S]
• 15 non-sarcomatoid RCC– All clear cell [E*]
Sircar K. The Journal of Pathology: Clinical Research. 2015
Sarcomatoid RCC vs. non-sarcomatoid RCC
cDNA microarray (unsupervised) RNA-seq
Sircar K. The Journal of Pathology: Clinical Research. 2015
Sarcomatoid RCC vs. Grade 4 non-sarcomatoid RCC
cDNA microarray (unsupervised) RNA-seq
Sircar K. The Journal of Pathology: Clinical Research. 2015
Fox Chase Cancer Center
• SNP-based microarrays CNA
– 9 sRCC (3 ccRCC, 2 pRCC, 4 unclassified RCC)
– 71 non-sRCC (39 ccRCC, 26 pRCC, 6 chrRCC)
• Unique for sRCC
– Loss 9q (88%), 15q (77%), 18q (66%), 22 (77%)
– Gain 1q (55%) and 8q (66%)
Ito T. J Clin Oncol 33, 2015 (suppl 7; abstr 478)
MSKCC
• 7 patients with sRCC with ccRCC epithelioid• Whole exome sequencing
• Mutations:– Similar VHL gene mutations (in 6/7, in both S and E)– PBRM1 mutations (in 4/7, 3 in both S and E)– SETD2 mutations (in 3/7, 2 in S component only)– TERT promoter mutations (in 2/7, in both S and E)– No mutations were found in KDM5C, PTEN, MTOR or TP53
• Chromosome changes:– Chromosome 3p loss (in 6/7 E, in 3/7 S)– Chromosome 14q24 loss (in 3/4, in both S and E)– Chromosome 9p21 loss (in 4/5 samples, in both S and E)– Chromosome 17q23-24 gain (in 3/7 S, none in E)
Mano R. AUA 2015. MP47-09
MD Anderson
• Primary Objective
– Identify genomic alterations in sRCC using a genomic profiling assay
Malouf G. Submitted for publication
Outline of the Project
• Step 1: GP of 3 sRCC patients using matched epithelial (ccRCC) and sarcomatoid components of sRCC
• Step 2: GP of 26 sRCC cases • Step 3: Comparison with GP of 56 non-
sarcomatoid cases (internal validation)• Step 4: Comparison with TCGA data (external
validation)
Malouf G. Submitted for publication
Methods
• FFPE
• Targeted sequencing done at Foundation Medicine Inc.
– 3,230 exons of 236 cancer-related genes
– 37 introns from 19 genes
Malouf G. Submitted for publication
Step 1: GP of 3 sRCC patients using matched epithelial (ccRCC) and sarcomatoid components
Patient Stage Histology MEDIAN
EXON
DEPTH
KNOWN SOMATIC
SHORT-VARIANTS
LIKELY SOMATIC SHORT-VARIANTS AMPL HOMOZYG
OUS
DELETIONS
A14-1 IV Epithelial/Clear cell 884PTEN_c.277C>T_p.H93Y
TP53_c.473G>T_p.R158L
PTEN_c.280A>T_p.N94Y,
PTEN_c.1027-1G>T_p.splicenone VHL
Sarcomatoid (70%) 1006LRP1B_c.10638G>C_p.E3546D
TP53_c.395A>T_p.K132MPTEN_c.209+1G>C_p.splice JAK2 VHL
A14-2 IV Epithelial/Clear cell 844 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
Sarcomatoid (60%) 892 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
A5 IV Epithelial/Clear cell 934 none VHL_c.513_513delG_p.K171fs*31 none none
Sarcomatoid (20%) 354 none VHL_c.513_513delG_p.K171fs*31 none none
Malouf G. Submitted for publication
Step 1: GP of 3 sRCC patients using matched epithelial (ccRCC) and sarcomatoid components
Patient Stage Histology MEDIAN
EXON
DEPTH
KNOWN SOMATIC
SHORT-VARIANTS
LIKELY SOMATIC SHORT-VARIANTS AMPL HOMOZYG
OUS
DELETIONS
A14-1 IV Epithelial/Clear cell 884PTEN_c.277C>T_p.H93Y
TP53_c.473G>T_p.R158L
PTEN_c.280A>T_p.N94Y,
PTEN_c.1027-1G>T_p.splicenone VHL
Sarcomatoid (70%) 1006LRP1B_c.10638G>C_p.E3546D
TP53_c.395A>T_p.K132MPTEN_c.209+1G>C_p.splice JAK2 VHL
A14-2 IV Epithelial/Clear cell 844 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
Sarcomatoid (60%) 892 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
A5 IV Epithelial/Clear cell 934 none VHL_c.513_513delG_p.K171fs*31 none none
Sarcomatoid (20%) 354 none VHL_c.513_513delG_p.K171fs*31 none none
Malouf G. Submitted for publication
Step 1: GP of 3 sRCC patients using matched epithelial (ccRCC) and sarcomatoid components
Patient Stage Histology MEDIAN
EXON
DEPTH
KNOWN SOMATIC
SHORT-VARIANTS
LIKELY SOMATIC SHORT-VARIANTS AMPL HOMOZYG
OUS
DELETIONS
A14-1 IV Epithelial/Clear cell 884PTEN_c.277C>T_p.H93Y
TP53_c.473G>T_p.R158L
PTEN_c.280A>T_p.N94Y,
PTEN_c.1027-1G>T_p.splicenone VHL
Sarcomatoid (70%) 1006LRP1B_c.10638G>C_p.E3546D
TP53_c.395A>T_p.K132MPTEN_c.209+1G>C_p.splice JAK2 VHL
A14-2 IV Epithelial/Clear cell 844 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
Sarcomatoid (60%) 892 VHL_c.473T>C_p.L158P PBRM1_c.100A>T_p.K34* none none
A5 IV Epithelial/Clear cell 934 none VHL_c.513_513delG_p.K171fs*31 none none
Sarcomatoid (20%) 354 none VHL_c.513_513delG_p.K171fs*31 none none
Malouf G. Submitted for publication
Step 2: GP of 26 sRCC cases
• 7 females and 19 males
• Stage
– III = 7
– IV = 18
– n/a = 1
Malouf G. Submitted for publication
Step 2: GP of 26 sRCC cases
• Epithelial histology– Clear cell = 12– Unclassified = 9– Collecting duct = 2– Papillary = 1– MTSCC = 1– n/a = 1
• Tumor assayed– Primary renal tumor in 23 cases– Metastatic site in 3 cases (lymph node, liver metastasis, and peritoneal
nodule)
Malouf G. Submitted for publication
Step 3: Comparison with GP of 56 non-sarcomatoid cases (internal validation)
• Compared our 26 sRCC cases with 56 advanced stage clear cell RCC cases • Evaluated by the same CGP• Grade at diagnosis
– Grade 2 = 21 – Grade 3 = 19 – Grade 4 (non-sarcomatoid) = 16
• Stage at diagnosis– Stage I = 1– Stage II = 4– Stage III = 12– Stage IV = 39
Malouf G. Submitted for publication
Step 3: Comparison with GP of 56 non-sarcomatoid cases (internal validation)
• Mutations in these 56 clear cell RCC (non-sarcomatoid)
• VHL = 73%
• PBRM1 = 47%
• SETD2 = 31%
• BAP1 = 13%
• TP53 only 9%
• NF2 only 2%
• KRAS, NRAS or HRAS None
Malouf G. Submitted for publication
Step 4: Comparison with TCGA data (external validation)
TP53 (%) NF2 (%)
05
1015
20
25
30
354045
50
TCGA. Nature. 2013TCGA. Cancer Cell. 2014Durinck S. Nat Genet. 2015TCGA. NEJM. 2015
05
1015
20
25
30
354045
50
PD-1 and PD-L1 (Mayo Clinic, AZ and Caris)
• Sarcomatoid (40 26) ?cc• ccRCC non-sarcomatoid (91 29)
– Only 6 of 91 (7%) were Grade 4
• Whole sections• PD-1:
– BD Pharmingen, Clone 561273 – ≥1 PD-1+ TIL/mm2
• PD-L1: – 2 Abs (R&D systems, Clone 130021 and Spring Bioscience, Clone SP142)– ≥5% staining – Score of 2+ or 3+
Joseph RW. Cancer Immunol Res. 2015
PD-1 and PD-L1 (Mayo Clinic, AZ and Caris)
• PD-1+ TILs
– Sarcomatoid 25 (96%)
– ccRCC non-Sarc 18 (62%)
– P=0.003
• PD-L1
– Sarcomatoid 14 (54%)
– ccRCC non-Sarc 5 (17%)
– P=0.006
Joseph RW. Cancer Immunol Res. 2015
PD-L1 and PD-L2 (Asan Medical Center)
• Sarcomatoid RCC 54• Clear cell non-sarc RCC 150• TMA• PD-L1:
– Cell Signaling Technology, Clone E1L3N– moderate expression in ≥5 % of tumor cells– strong expression in ≥5 % of tumor cells
• PD-L2:– R&D systems, Clone 176611– Similar to PD-L1
Shin SJ. Ann Surg Onc. 2015
PD-L1 and PD-L2 (Asan Medical Center)
• PD-L1
– Sarcomatoid 16 (29.6%)
– ccRCC non-Sarc 11 (7.3%)
– P<0.001
• PD-L2
– Sarcomatoid 24 (44.4%)
– ccRCC non-Sarc 46 (30.7%)
– P=0.067
Shin SJ. Ann Surg Onc. 2015
Sarcomatoid RCC (MD Anderson)
• 118 patients (94 clear cell)
• FFPE
• Re-reviewed by GU pathology
• Epithelial and sarcomatoid components
• Whole slides and TMA
Kawakami F. Submitted
Control Group: ccRCC
• All clear cell RCC
– No sarcomatoid components
• 92 patients
– Grade 2 = 14
– Grade 3 = 58
– Grade 4 = 20
Kawakami F. Submitted
PD-1 and PD-L1
• PD-1– Staining manually evaluated
– Positive: PD-1 cell numbers≥1/HPF
• PD-L1– Digital analysis
– Semiquantitative method (H-score=0-300)
• Results will be presented at ASCO GU 2016
Kawakami F. Submitted
Future Directions
• Identifying sRCC on imaging• Identifying sRCC using molecular signature on
biopsy• Validation of DNA sequencing findings• Mechanistic work• Animal models• Novel therapeutics
Acknowledgements
• Urology– Christopher Wood– Surena Matin– Mehrad Adibi
• GU Medical Oncology– Eric Jonasch– Nizar Tannir
• Interventional Radiology– Kamran Ahrar– Sabir Sharjeel
• Translational Molecular Pathology– Fumi Kawakami– Ignacio Wistuba– Jaime Rodriguez-Canales
• Pathology– Pheroze Tamboli– Kanishka Sircar
• Urology Fellows/Residents– Megan Merrill– Patrick Kenney– Kara Babaian– Arun Thomas– Zachary Compton
• Radiology– Rivka Colen
• Statistics– Rebecca Slack
• Bioinformatics and Computational Biology– Xiaoping Su
• Foundation Medicine– Siraj Ali– Kai Wang
• City of Hope– Sumanta Pal
• Pitie-Salpetriere– Gabriel Malouf