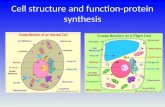
Information about Cell and it's structure and protein synthesis
Protein Synthesis and Structure
-
Upload
susan-dejesus -
Category
Documents
-
view
28 -
download
1
description
Transcript of Protein Synthesis and Structure

Protein Synthesis and StructureSection 2-4

Protein Functions: General Information• Proteins account for almost 50% of the dry mass of most cells
• Proteins are the most structurally sophisticated molecules known
• Each protein has a specific 3-diminsional shape, or “confirmation” that is vital to its function

Protein Functions• Enzymatic• Structural• Storage• Transport• Hormonal• Receptor• Contractile and motor• Defensive

Four Levels of Protein Structure

Primary Structure• The unique sequence of amino acids
that make up a polypeptide
• Amino acids:• An asymmetrical carbon• An amino group (contains Nitrogen)• A carboxyl group• A side chain (R group) which makes each
amino acid unique

Primary Structure• Amino acids are linked together with
dehydration reactions
• Peptide bonds- the bonds between amino acids

Primary Structure• The chain will have two ends:• An amino end- known as the N-
terminus
• A carboxyl end- known as the C-terminus
• Primary structure is achieved through the processes of transcription and translation

Secondary Structure• Refers to the coils and folds in the polypeptide chain due to
hydrogen bonds between repeating areas on the polypeptide backbone

Secondary Structure• Alpha helix (α helix)- a delicate coil held together by hydrogen
bonding between every fourth amino acid
• Beta pleated sheet (β pleated sheet)- two or more regions of a polypeptide chain lying side by side and connected by hydrogen bonds between the two parallel polypeptide back bones

Tertiary Structure• Folding due to interactions
between the amino acid side chains
• Main causes of tertiary structure• Hydrogen bonds between polar
R-groups• Hydrophobic interactions
between nonpolar R groups causes them to clump together and form Van der Wall’s interactions
• Dislufide bridges between two sulfhydryl groups

Tertiary Structure: Main Causes• Hydrogen bonds between polar
R-groups
• Hydrophobic interactions between nonpolar R groups causes them to clump together and form Van der Wall’s interactions
• Dislufide bridges between two sulfhydryl groups

Quaternary Structure• Overall shape of the protein caused by the association of two
or more polypeptide chains• NOT ALL PROTEINS HAVE QUATERNARY STRUCTURE

Protein Synthesis

General Information• Also called gene
expression
• DNA provides the blueprints for the building of proteins

General Information• Involves two processes:
• Transcription- copying DNA into mRNA
• Translation- translates the code from nucleic acid into amino acid at the ribosome

Evolutionary Advantage of Transcription and Translation• DNA is protected inside the nucleus
• Using an RNA intermediate allows multiple copies of a protein to be made at once because many mRNA molecules can be made from one gene, then translated repeatedly

Prokaryotes vs. Eukaryotes
Prokaryotes
• Only one compartment (no nucleus)• Transcription and
Translation occur simultaneously

Prokaryotes vs. Eukaryotes
Eukaryotes
• Transcription occurs in the nucleus
• The primary transcript is then modified (RNA processing) before leaving the nucleus
• Translation occurs in the cytoplasm at the ribosome

The Genetic Code• Triplet Code- the flow if information from gene (DNA) to
protein is written in the DNA as non-overlapping, three-nucleotide segments
• Template Strand:• The mRNA is complimentary to the template strand
• The DNA is read in the 3’ to 5’
• The mRNA is synthesized and read from 5’ to 3’

The Genetic Code• Codons- each three base sequence on the mRNA strand• Each codon codes for a specific amino acid

Redundant but not Ambiguous• Redundant- multiple
codons can code for the same amino acid
• Not Ambiguous- no codon codes for more than one amino acid

Special Codons• AUG= start
• UAA, UAG, UGA= stop

Transcription

Initiation• RNA polymerase binds to the promoter
• The promoter is a specific sequence that tells the RNA polymerase where to bind and determines what DNA strand will serve as the template
• In eukaryotes, specific proteins called transcription factors assist the RNA polymerase in binding and forming the transcription initiation complex

Initiation

Elongation• RNA polymerase adds
nucleotides to the 3’ end of the growing RNA molecule
• Complimentary base pairing occurs
• The new RNA molecule peals away from the DNA template and the DNA reforms

Termination• In prokaryotes, the RNA polymerase detaches after the
termination signal is transcribed
• In eukaryotes, the RNA polymerase transcribes the polyadenylation signal sequence then the mRNA is cut off of the RNA polymerase

RNA Processing
EUKARYOTIC CELLS ONLY

Altering of the Ends of the mRNA• 5’ cap- modified guanine molecule added on the 5’ end
• Poly-A-tail- 50-250 adenine nucleotides are added to the 3’ end
• Functions:• Facilitate export from the nucleus• Protect the mRNA from degradation by enzymes• Assist the ribosomes in attaching in the cytoplasm

RNA Processing

RNA Splicing• Removal of large portions of the mRNA
• snRNPs (“snurps”) recognize and cut out areas of the mRNA
• Introns- the portions of the mRNA that are removed
• Exons- the portions of the mRNA that exit the nucleus

Translation

Transfer RNA, tRNA• Translates nucleotides
into amino acids
• One end has an anticodon, complementary to the mRNA codon
• The other end is bound to an amino acid
• Excellent example of how structure fits function

Ribosomes• Contain three sites for holding tRNA:
• P site- holds the growing polypeptide chain
• A site- holds the tRNA that is carrying the next amino acid in the chain
• E site- where the tRNA leaves the ribosome
• Exit Tunnel= where the polypeptide leaves the ribosome

Translation- The Process

Initiation• Small ribosomal subunit binds the mRNA and the initiator
tRNA
• Subunit scans the mRNA until it reaches the start codon, establishing the correct reading frame as the tRNA hydrogen bonds to the start codon

Initiation• Translation initiation complex forms- the large ribosomal
subunit attaches with the assistance of initiation factors and an expenditure of energy

Elongation

Elongation
• The ribosome reads the mRNA in the 5’ to 3’ direction
• Anticodon of the incoming tRNA hydrogen bonds to the mRNA codon in the A site
• The peptide bond forms between the amino acid on the tRNA of the A site and the growing polypeptide chain in the P site
• Translocation of the tRNA shifts the A site tRNA to the P site and the P site tRNA to the E site so it can exit

Termination• Release factor:• Added when stop codon is reached
• Causes the addition of a water molecule to the end of the polypeptide
• The polypeptide is released

Forming a Functional Protein

Protein Folding• Folding occurs as the protein is being synthesized
• Folding is dependent on• The properties of the peptide chain• The physical and chemical properties of the environment
WHY MIGHT THIS BE A PROBLEM???

Chaperonins• Proteins that assist in the proper folding of other proteins by
shielding them form the cell environment

Post-Translational Modification• Chemical modification by the attachment of sugars, lipids,
phosphate groups, or other components
• Enzymes may remove one or more amino acids from the N-terminus
• Single polypeptides may by cut into two or more smaller pieces

Denaturation• The changes in a protein’s native conformation that renders it
biologically inactive
• Factors that cause denaturation:• Change in the environment• Change in temperature• Change in pH

Changes in Environment• If moved from an aqueous environment to a nonpolar organic
solvent, the protein will turn inside out
• Chemicals can disrupt disulfide and hydrogen bonds that stabilize secondary and tertiary structure

Changes in Temperature• Excessive heat can cause movement to overpower sensitive
hydrogen bonds
• Excessive cold will slow the protein down substantially

Changes in pH• All proteins have an optimal pH at which they function
• Optimal pH is not necessarily 7