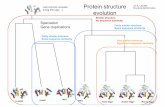
Protein Evolution: Structure, Function, and Human Health
-
Upload
dan-gaston -
Category
Education
-
view
386 -
download
0
description
Transcript of Protein Evolution: Structure, Function, and Human Health

Protein Evolu-on
Structure, Func-on, and Human Health
11/28/2013 Dr. Daniel Gaston, Department
of Pathology 1

So, about this evolu-on thing?
Why should I care? What use is it?

Lots of reasons
• Knowledge for its own sake is good – Otherwise, why do science at all?

Lots of reasons
• Knowledge for its own sake is good – Otherwise, why do science at all?
• Shapes our understanding of ecology and biological diversity

Lots of reasons • Knowledge for its own sake is good
– Otherwise, why do science at all? • Shapes our understanding of ecology and biological diversity
• Prac-cal reasons – An-bio-c resistance – Microbiome: Fecal transplanta-on – Cancer – Predic-ng gene/protein func-on – Predic-ng the impact of muta-ons for poten-al to cause human disease (Genotype:Phenotype)

Evolu-on of Life on Earth
A (Very) Brief Overview

Eubacteria"
ROOT Iwabe et al. 1989 Gogarten et al. 1989
Eukaryota"
Archaebacteria"

Eubacteria"
ROOT Iwabe et al. 1989 Gogarten et al. 1989
Eukaryota"
Archaebacteria"

Eubacteria"
ROOT Iwabe et al. 1989 Gogarten et al. 1989
Eukaryota"
Archaebacteria"


You are here

A Brief History of Cells and Molecules
• Origin of the earth ~4.5 billion years ago • Origin of life: ~3.0-4.0 billion years ago
– Origin of self-replicating entities – The RNA world (?) – Origin of the first genes, proteins & membranes – Gave rise to the first cells – the Last Universal Common Ancestor (LUCA) of all cells
– Probably had 500-1000 genes • First microfossils of bacteria: ~3.5 billion years ago (controversial)
~2.7 billion years ago (for certain) • Oxygenation of the atmosphere: 2.3-2.4 billion years ago (by
photosynthetic bacteria) • Origin of eukaryotes: ~1.0-2.2 billion years ago (probably 1.5) • Origin of animals: ~0.6-1.0 billion years ago

• Homology = descent from a common ancestor – homology is all or nothing: sequences are either
homologous (related) or not homologous (not related)
– Not the same as “similarity” (degrees of similarity are possible)
Some Defini-ons

Some Defini-ons • Divergence = change in two sequences over time
(after splitting from a common ancestor)
• Convergence = similarity due to independent evolutionary events
– On the amino acid sequence level, it is relatively rare & difficult to prove (but see an example later)
T T
Ancestral sequence
Sequence 1 Sequence 2

How does evolutionary change happen in proteins?

Evolu-on: Two Groups of Processes
• Muta-on – Many different processes that generate muta-ons – Muta-ons are the raw materials needed for evolu-on to happen
• Selec-on and DriY – Muta-ons happen in individuals – Evolu-on happens in popula-ons of organisms – Selec-on and Gene-c DriY affect the frequency of muta-ons in a popula-on over -me

Muta-ons

Point Muta-ons
! ! AGGTTCCAATTAA!! ! TCCAAGGTCAATT!
!!AGGTTCCAATTAA ! TCCAAGGTTAATT!!
REPLICATION (meiotic or mitotic division)
Unrepaired mispaired base
Mutant allele Wild-type alleles
Mutant Gamete (for multicellular org.)
Wild-type Gamete (for multicellular org.)
AGGTTCCAGTTAA ! TCCAAGGTCAATT!

AGTCCAAGGCCTTAA -------------> AGTTCAAGGCCTTAA point mutation ���
CCTTA AGTCCAAGGCCTTAA -------------> AGTCCAAGGCCTTACCTTAA
insertion
AAGG AGTCCAAGGCCTTAA -------------> AGTCC-CCTTAA
deletion AGTCCAAGGCCTTAA -------------> AGTCCCCTTCCTTAA
` inversion AGTCCAAGGCCTTAA -------------> AGTCCAAGGCC + translocation + GGTCCTGGAATTCAG GGTCCTGGAATTCAGTTAA AGTCCAAGGCC --------------> AGTCCAAGGCCAGTCCAAGGCC duplication AAGG AGTCCAAGGCCTTAA ---------------> AGTCCAAAGGCTTAA
recombination AGGC

Larger Scale Muta-ons

Exon shuffling and Protein Domains
Exon1 Exon 2 Exon 3

Exon shuffling and Protein Domains
Exon1 Exon 2 Exon 3
Domain 1 Domain 2

Exon shuffling and Protein Domains
Exon1 Exon 2 Exon 3

Exon shuffling and Protein Domains
Exon1 Exon 2 Exon 3
Domain 2 Domain A

Genomic Scale Muta-ons
Gene 1 Gene 2

Genomic Scale Muta-ons
Gene 1 Gene 2

Gene Duplica-on
Gene 1 Gene 2

Gene Duplica-on
Gene 1 Gene 2 Gene 1a

Gene-c DriY and Selec-on

Mutations vs. substitutions
• Mutations happen in individual organisms
• A nucleotide ‘substitution’ occurs IF after many generations, all individuals in the population harbour the ‘mutation’
• This process is called “fixation of mutations”
• substitution = fixed mutation • When comparing homologous protein sequences between
species, looking at amino acid substitutions

Fixation of alleles
N generations
Proportion of = 1.0 (100%) This is the same as saying that was fixed in the population in N generations The ‘mutation’ became a ‘substitution’ after it was fixed in the population
Population with two alleles:
Proportion of = 1/14 (7.1%) Proportion of = 13/14 (93%)

Natural selection and Neutral drift • Positive selection
– Mutation confers fitness advantage (more offspring that survive)
– RARE • Purifying selection (negative selection)
– Mutation confers fitness disadvantage (less offspring or ‘no’ viable offspring - e.g. lethal)
– FREQUENT • Neutral evolution (genetic drift)
– Mutation has very little fitness effect – Will drift in frequency in the population due to random
sampling effects – VERY FREQUENT

Nearly-neutral theory ���

Common Examples of Posi-ve Selec-on
• MHC Genes – Diversity = Good – Very polymorphic in humans
• Envelope (gp120) of HIV – Immune system evasion
• Enzymes involved in human dietary metabolism – Accelerated posi-ve selec-on over last ~10,000 years

Gene-c DriY
Select a marble randomly from a jar and “copy” it in to the next Fixa-on of the plain blue allele in 5 genera-ons

Polymorphism
• Polymorphisms are sites with more than one allele present in a popula-on – Muta-ons that have not yet been fixed

Muta-on and Codons
Not all muta-ons are created equal

Point mutations in protein genes are classified according to the genetic code:
The genetic code is degenerate: more than one codon often specifies a single amino acid. E.g. Serine has 6 codons, Tyrosine has 2 codons and Tryptophan has one codon!

Point mutations in ���protein-coding genes
• synonymous (silent) substitutions: cause interchange between two codons that code for the same amino acid:
e.g. CTG --> CTA = Leu --> Leu Mostly invisible to selection
• non-synonymous (replacement) mutations: cause change between codons that code for different amino acids (missense) or stop codons (nonsense)
e.g. CTG --> ATG = Leu --> Met TGG --> TGA = Trp --> Stop


8 kinds of 1st codon-position synonymous mutation: R-->R and L-->L

126 kinds of 3rd-codon position synonymous mutation:

A Note on Indels
• Ignored because indels are far more likely to be deleterious – More likely to result in frame shiYs
• Can s-ll be non-‐deleterious – Par-cularly if in mul-ples of three – Over evolu-onary -me indels more oYen observed in loops than more constrained structural elements

Evolu-onary Rates
Speed of Evolu-on

Rates of protein evolution���(i.e. rates that individual amino acids are substituted)
• Different regions in proteins have different rates of evolution (functional constraints)
• Different proteins have different overall rates of evolution


Enolase • Ubiquitous glycolytic enzyme, highly conserved throughout evolution
• TIM Barrel family doing an α-proton abstraction
cMLE
MLE
Archaea
Bacteria
Euks
β α γ

All Eukaryotes site rates (63 taxa) mapped on Lobster Enolase
low rates blue high rates red

Site rate categories 1 and 2 (slowest sites)

Site rates Categories 3 and 4

Site rates Categories 5 and 6

Site rates Categories 7 and 8 (fastest sites)

Evolutionary rates as a function of enolase structure/function
• Rates of evolution increase from the centre of the molecule (slow) to the surface (fast)
• The pattern is probably due to: – Distance from the catalytic centre --> catalytic residues don’t change
(slowest), residues that interact with catalytic residues are constrained (slow)
– Geometric constraints - residues in the centre of the molecule have restricted ‘space’ around them that constrains them. At the surface, there are fewer such constraints
– Hydrophobic core in centre – More loops and alpha helices on surface
• NOTE: this pattern seems to work for soluble globular enzymes with catalytic centre in the centre of mass. It does not hold for structural proteins like tubulin, actin etc.

Rates of evolution of sites versus their structural position
• There are no completely general rules! – It depends on what the protein is doing and where.
• Functional sites (catalytic sites) or sites at interfaces (protein-protein interactions) are conserved
• Geometric, chemical, folding and functional constraints (catalysis, binding) determine evolutionary constraints

Detec-ng and Quan-fying Evolu-onary Rela-onships

How do we know if two proteins are homologous?
(A) If sequences > 100 amino long are >25% identical --> they are probably significantly similar and very likely to be homologous -BLAST, FASTA, Smith-Waterman algorithms are likely to find them “significantly similar” (E-value << 1x10-4)
(B) If they are >100 long and 15-25% identical (Twilight Zone) --> probably homologous BUT need to rigourously test it -a number of methods are available: permutation test
(C) If they are <15% identical......difficult to prove homology -test it -if its not significant look for motifs in multiple alignments -look at tertiary structure

15-23%!identity!
}!


Applica-ons
• Evolu-onary methods for studying protein func-on – Annota-ng novel proteins – Func-onal divergence
• Predic-ng pathogenicity of muta-ons Informing protein structure predic-on – Mendelian disease – Cancer

Applica-ons of Evolu-onary Biology to Medicine
Inherited Gene-c Diseases and Cancer

Lynch Syndrome
• Autosomal dominant cancer syndrome • Increased risk for many cancers, mostly colorectal cancer due to mismatch repair defects

Lynch Syndrome
• Autosomal dominant cancer syndrome • Increased risk for many cancers, mostly colorectal cancer due to mismatch repair defects

Mutator Phenotype
• Inac-va-on of mismatch repair (MMR) genes led to mutator phenotypes in E. coli and yeast • Included Microsatellite instability

Mutator Phenotype
• Inac-va-on of mismatch repair (MMR) genes led to mutator phenotypes in E. coli and yeast • Included Microsatellite instability
• Careful research iden-fied human homologs – MLH1 and MSH2 – Defects in these genes cause Lynch Syndrome

Mismatch Repair
• Mismatch Repair -‐> • Microsatellite Instability -‐> • Cancer Most microsatellites spread throughout the genome in non-‐genic regions But some are found in important tumor suppressor genes

Applica-ons of Evolu-onary Biology to Medicine
Predic-ng Pathogenicity and Impact of Human Muta-ons

The Sequencing Revolu-on

Problem
• OYen leY with hundreds to thousands of poten-al muta-ons in a family that “track” with the disease – Needle in a “stack of needles” problem
• Must discriminate neutral missense muta-ons from pathogenic ones

Evolu-on at Work
• Many programs exist to make these predic-ons: – PolyPhen – Muta-on Taster – EvoD – SIFT – PROVEAN – FATHMM – etc

Evolu-on at Work
• Important amino acids have low evolu-onary rates – Higher conserva-on
• The more important the protein the more likely it is to be broadly found among eukaryotes – Also higher overall conserva-on
• However many important proteins in humans only found in primates, mammals, or animals

Evolu-on at Work
…RPLAHTY…! …RPLAHTY…!…RPLVHTY…!…RPIAHTY…!…RPIGHTY…!…RPIICTY…!…RPLACTY…!…RPLLCTY…!!
Reference Sequence Mul-ple Sequence Alignment

Evolu-on at Work
…RPLAHTY…! …RPLAHTY…!…RPLVHTY…!…RPIAHTY…!…RPIGHTY…!…RPIICTY…!…RPLACTY…!…RPLLCTY…!!
Reference Sequence Mul-ple Sequence Alignment
Compute an Evolu-onary Conserva-on Score for Each Posi-on

Evolu-on at Work
…RPLACTY…! …RPLAHTY…!…RPLVHTY…!…RPIAHTY…!…RPIGHTY…!…RPIICTY…!…RPLACTY…!…RPLLCTY…!!
Reference Sequence Mul-ple Sequence Alignment
Conserva-ve changes more likely to be neutral

Evolu-on at Work
…RPLACTP…! …RPLAHTY…!…RPLVHTY…!…RPIAHTY…!…RPIGHTY…!…RPIICTY…!…RPLACTY…!…RPLLCTY…!!
Reference Sequence Mul-ple Sequence Alignment
Radical changes more likely to be deleterious

Applica-ons of Evolu-onary to Protein Func-on
Func-onal Divergence

Func-onal Divergence
Gene 1 Gene 2 Gene 1a
Over evolu-onary -me scales Gene 1 and Gene 1a are known as paralogs, a subset of homologs They can diverge from one another in sequence, as well as func-on.

Types of Func-onal Divergence
• Subfunc-onaliza-on – Paralog specializes and retains only a subset of ancestral func-on
• Neofunc-onaliza-on – Paralog gains a new func-on, and loses old func-on(s)
• Subneofunc-onaliza-on – Paralog undergoes rapid subfunc-onaliza-on but then undergoes neofunc-onaliza-on

Gene A
Family B
Family A
Func-onal Divergence

Func-onal Divergence …A L H… Species 1 …A L H… Species 2 …A L H… Species 3 …A L H… Species 4 …A L H… Species 5 …A L H… Species 6
…R A H… Species 1 …R R H… Species 2 …R C H… Species 3 …R A H… Species 4 …R A H… Species 5 …R Y H… Species 6
Family B
Family A

Glyceraldehyde-‐3-‐Phosphate Dehydrogenase
NAD+ NADH +Pi +H+
NAD+ NADH + Pi + H+
Glyceraldehyde-‐3-‐Phosphate 1,3-‐Biphosphoglycerate
Cytosol: Glycolysis

Glyceraldehyde-‐3-‐Phosphate Dehydrogenase
NADP+ NADPH +Pi +H+
NADP+ NADPH +Pi +H+
Glyceraldehyde-‐3-‐Phosphate 1,3-‐Biphosphoglycerate
Plas-d: Calvin Cycle

GAPDH Evolu-on
Green Plants
Cyanobacteria
‘Chromalveolates’
Cytosolic GapC
Cytosolic GapC

GAPDH Structure

NADPH Binding Necessary for Calvin Cycle Func-on