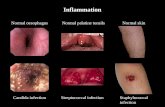
normal carcinome inflammation
Transcript of normal carcinome inflammation

carcinomenormal
inflammation

1964

DKFZ Genomics Activities

MolTools Partners
Anthony BrookesKarolinska Institute
Olli KallioniemiVTT Technical Research Centre
Mike TaussigBabraham Institute
Ed SouthernOGT
Andres, MetspaluEstonian Biocentre
Arvydas JanulaitisFermentas UAB
Ivo GutCentre National de Génotypage Jörg Hoheisel
Peter LichterAnnemarie PoustkaDKFZ
Jörn KochAarhus University
Hans LehrachMax Planck Institute
Ove ÖhmanÅmic
Ulf LandegrenUppsala University
EU
Marc ZabeauMethexis Genomics
Dolores CahillRoyal College of Surgeons
George ChurchHarvard University, USA
Marcus Beierfebit biotech

Basic Level of Functional Analysis
DNA RNA ProteinDNA RNA ProteinTranscription ExpressionMethylation
LocalisationStructure SplicingModificationInteractionOrganisation
RNAiSNPs CombinationStructure
Copy number
DegradationComposition
Enzymatic activity

Epigenetics Platform
• The Human Epigenome Project• Characterisation of DNA-methylation patterns
in tumours• MolTools: Advanced molecular tools for
array-based analyses• Amplification- and label-free SNP-analysis
EU
Nanotechnology
Systematic-Methodological Platform Epigenetics
DKFZ:Jörg Hoheisel (Coordination,
Chip Technologies)Frank Lyko (Epigenetics)Herman-Josef Gröne (Tumour bank)
Bonn UniversityAndreas Waha (Chip Technologies)
University of SaarbrückenJörn WalterInternational University BremenAlbert JeltschMPI for Molecular Genetics, BerlinRichard ReinhardIMB JenaAlbert Platzer

Transcriptional Profiling
gene 1 gene 2 gene 3
cDNA microarray
sample 1 sample 2RNA
scanned image

Transcriptional Profiling Projects
ManMouseDrosophila melanogasterArabidopsis thalianaSaccharomyces cerevisiaeHydractinia echinata
Bacillus subtilisTrypanosoma bruceiNeurospora crassaPseudomonas putidaRhizobium meliloti
EU
EU
EU

In Situ Synthesised Oligonucleotide MicroarrayN. crassa Microarray
Aign & Hoheisel (2003) Fungal Genet. Biol. 40, 225-233.

Mask-Free Control of Oligonucleotide Synthesis
Light
Amidites
Digital micromirror system
Photolabile protection group
1. 2. 3.
1. Molecules bound to glass surface
2. Cleavage of protection groups at illuminated spots
3. Condensation of amiditesresults in oligomer elongation

Chip-Based Analyses
On-chipenzymatic reaction
e.g. primer extension
5´
dTTP dGTP dCTP
GTCGG5´
CGATCGG5´
TGCTAGCC
TGGTA
AC
C
dATP5´
3´3´
hybridisation ofwildtype target
wtwt mt
mt
DNA-polymerase
analysis in solution; on-chip molecule separation
GCTATTGACCA...
tag 15´
ACC
tag 25´
ACCGCTA...
hybridisation ofwildtype target
wtwt mt
mt
Detection via tag sequencesDirect hybridisation
GGCTAGCAT3´
hybridisation ofwildtype target
wtwt mt
mt
GGCTGGCAT3´
CCGATCGTA
3´
5´
CG
GTT
CA
TA
3´
5´
mismatch versus full-matchdiscrimination

ZIP-Code Array vs. Gene-Specific Array
oligo-dTor
gene-specificprimer
template specificprimer extension and labelling
template DNA or RNA
Array ofgene-specificprobes

ZIP-Code Array vs. Gene-Specific Array
Array ofcomplementary
ZIP-code probes
Array ofgene-specificprobes
gene-specificprimer
template specificprimer extension and labelling
template DNA or RNA
ZIP-code
5’

Transcript Profiling on ZIP-Code Array
ZIP-code
5’
primer extension and labelling
AAAAAA-3’
gene-specificprimer
ZIP-code
5’
primer extension and labelling
AAAAAA-3’
sample 1
sample 2
gene-specificprimer
hybridisation toZIP-code array

Transcript Profiling on ZIP-Code Array100000
10000
1000
100
10
34.7
7.0 6.2
-25.9
sign
al in
tens
ity
30
20
10
0
-10
-20
n-fo
ld c
hang
e
standardhybridisation
ZIP-codeanalysis
6.36.4
24.3
-35.7
3.4
20
10
0
-10
-20
-30
n-fo
ld c
hang
e
10000
1000
100
10
sign
al in
tens
ity
yeasthyphe
n-foldchange

Drosophila Exon-Specific PCR-ProductsDrosophila Microarray
Hild et al. (2003)Genome Biol. 5, R3.Boutros et al. (2004)Science 303, 382-385.Hollich et al. (2004)Biotechniques 37, 282-284.

23k Trypanosoma bruceiGenomic Fragment Microarray
T. brucei Microarray
Diehl et al. (2002)Mol. Biochem. Parasit. 123, 115-123.Brems et al. (2005)Mol. Biochem. Parasit. 139, 163-172.

Drosophila Genomic ArrayDrosophila Microarray
X
2L
2R
3R
4
(3L)
Minimal tiling path of 25,000 shotgun clonesmade from 320,000 clones used in sequencing
Hollich et al. (2004) Biotechniques 37, 282-284.

Variation by Preparation
Tumour sample 1vs.
Control
Tumour sample 2vs.
Control
C
2
C
1

Kinetic Parameters
spot size and format stirring vs. non-stirring incubation

In Situ Signal Amplificationsupported by
EU
self-assembly
targethybridisation
seco
ndar
yhy
brid
isat
ion
1600
1200
800
400
010 100 1 10 100 1 10
zmol amol fmol
Fluo
resc
. int
ensi
ty R2 = 0.9857

In Situ Signal Amplificationsupported by
EU
NH
ODMTO
NH
ODMTO
O
P N(Pr)2O
CNEt
Symmetric doubler phosphoramidite
25 nt 25 nt
26 nt
2000 nt
50 nt
20 40 60 80 100
Temperature / C
Rel
ativ
e ab
sorp
tion
550
259
Rel
ativ
e ab
sorp
tion Absorption
Self-assembly76 nt
Meltingbehaviour
200 400 600 800
Wavelength / nm

In Situ Signal Amplificationsupported by
EU Transcriptional profiling on Drosophila microarray
Ambion ArrayControl PCR spots
branchedoligomers control

Concordance Analysis
frequ
ency
[%
]
0.1 1 10
20
15
10
5
0
frequ
ency
[%
]
20
15
10
5
00.1 1 10
One samplelabelled with
Cy3 and Cy5
Two sampleslabelled withCy3 and Cy5
M-CHiPSData Processing Software

Concordance Analysis
One samplelabelled with
Cy3 and Cy5
Two sampleslabelled withCy3 and Cy5
frequ
ency
[%
]
frequ
ency
[%
]
20
15
10
5
00.1 1 100.1 1 10
20
15
10
5
0M-CHiPS
Data Processing Software

Concordance Analysis
One samplelabelled with
Cy3 and Cy5
Two sampleslabelled withCy3 and Cy5
frequ
ency
[%
]
0.1 1 10
20
15
10
5
0
frequ
ency
[%
]
20
15
10
5
00.1 1 10
±σM-CHiPSData Processing Software

Multi-Conditional HybridisationIntensity Processing SoftwareM-CHiPS
Database currently holds data of more than 5,900 experiments.
Database Browser
PostgreSQLDatabase
Selection of arbitrary subset
Matlab-based microarray data mining tools including- Normalisation - Quality filtering- Clustering, Correspondence analysis - Automated analysis of annotations
www.dkfz.de/funct_genome www.mchips.org

Normalisation Based onthe Majority of All Signal Intensities
log regressionlinear regression
logarithmic scalelinear scaleM-CHiPS
Data Processing Software

Normalisation Based onthe Majority of All Signal Intensities
log regressionlinear regression
logarithmic scalelinear scaleM-CHiPS
Data Processing Software

Normalisation Based onthe Majority of All Signal Intensities
linear regression log regression
logarithmic scalelinear scaleM-CHiPS
Data Processing Software

Normalisation
Each data set is correlated with the median of all controlsand fitted by linear or logarithmic regression.
M-CHiPSData Processing Software

Method Output
• Hierarchical Clustering treeEisen et al. (1998) PNAS 95, 14836
• K-Means Clustering set of clustersTavazole et al. (1999) Nat. Genet. 22, 281
• Learning Networks- Neural Networks, e.g. Kohonen Maps set of clusters
Tamayo et al. (1999) PNAS 96, 2907
- Baysian Networks directed graphFriedman et al. (2000) Proc. RECOMB 2000, 127
• Planar Embedding biplot- Multidimensional Scaling
Khan et al. (1998) Cancer Res. 58, 5009
- Principle Components AnalysisHilsenbeck et al. (1999) J. Natl. Cancer Inst. 91, 453
- Correspondence AnalysisFellenberg et al. (2001) PNAS 98, 10781
• Other MethodsBen-Dor et al. (1999) J. Comp. Biol. 6, 281 set of clusters
(incomplete list)
Method Output
• Hierarchical Clustering treeEisen et al. (1998) PNAS 95, 14836
• K-Means Clustering set of clustersTavazole et al. (1999) Nat. Genet. 22, 281
• Learning Networks- Neural Networks, e.g. Kohonen Maps set of clusters
Tamayo et al. (1999) PNAS 96, 2907
- Baysian Networks directed graphFriedman et al. (2000) Proc. RECOMB 2000, 127
• Planar Embedding biplot- Multidimensional Scaling
Khan et al. (1998) Cancer Res. 58, 5009
- Principle Components AnalysisHilsenbeck et al. (1999) J. Natl. Cancer Inst. 91, 453
- Correspondence AnalysisFellenberg et al. (2001) PNAS 98, 10781
• Other MethodsBen-Dor et al. (1999) J. Comp. Biol. 6, 281 set of clusters
(incomplete list)
DataClustering
M-CHiPSData Processing Software

Marker Identification
Hybridisation data from Affymetrix chips
Correspondence Analysis; Fellenberg et al., PNAS 98, 2001
patients
individual genes
M-CHiPSData Processing Software

Drosophila Developmental Transcript ProfileDrosophila Microarray
adult
pupal 3
pupal 2
pupal 1embryonic 12-16 h
larval
embryonic 8-12 h
embryonic 4-8 h
M-CHiPSData Processing Software
embryonic 0-4 h

Pancreatic Cancer vs. Chronic Pancreatitis
carcinomenormal
inflammation
Correspondence Analysis; Fellenberg et al., PNAS 98, 2001
patient
differentiallytranscribed gene
M-CHiPSData Processing Software

Target Selection
carcinomenormal
inflammation
Correspondence Analysis; Fellenberg et al., PNAS 98, 2001
M-CHiPSData Processing Software

Target Selection
carcinomenormal
inflammation
Correspondence Analysis; Fellenberg et al., PNAS 98, 2001
M-CHiPSData Processing Software

Analysis of Gene Ontology (GO) Terms
gene
0% glucose0.01% glucose0.1% glucose1% glucose
GO terms
Effect of glucose on yeast
M-CHiPSData Processing Software Yin et al. (2003) Mol. Microbiol. 48, 713-724.

Analysis of Gene Ontology (GO) Terms
Glu
cone
ogen
esi
Glykolysis
Identification ofaffected pathways
Effect of glucose on yeast
gene
0% glucose0.01% glucose0.1% glucose1% glucose
GO terms
M-CHiPSData Processing Software

Combination of Information Sources
7089
629845005
Mismatch repair(genes MSH2, MSH3, MLH1)
Traversing start control of mitotic cell cycle
(genes CDK10, CDC2, CDC25C)
normal tissue
new tumour entity
cystictumours
GO annotations
ductaladenocarcinoma
M-CHiPSData Processing Software
Busold et al. (2005), Bioinformatics, in press.

Complex Antigen-Antibody InteractionsAntibody Microarrays
illustration
EU
Kusnezow & Hoheisel (2002) BioTechniques 33 (suppl.), 14-23.Kusnezow et al. (2003) Proteomics 3, 254-264.Kusnezow & Hoheisel (2003) J. Mol. Recognit. 16, 165-176.Kusnezow et al. (2004) Protein Microarrays (Schena, M., ed.),
247-284.Kusnezow et al. (2005) Handbook of Immunohistochemistry
(Hayat, M.A., ed.), 23-37.
real experiment

Acknowledgements
• Frank LykoEpigenetics Group, DKFZ
• Helmut FriessChirugie Heidelberg
• Thomas GressMedizinische Klinik Abt. I
• Andres MetspaluInstitute of Molecular and Cell Biology
• Heinrich ArlinghausPhysikalisches Institut, Münster
• Nicole HauserInstitut für Grenzflächen und Bioverfahrenstechnik, Stuttgart
UniversitätHeidelberg
UniversitätUlm
TartuUniversity
Universität.Münster.

Functional Genome Analysis, DKFZ, Heidelberg, Nov. 2002
www.dkfz.de/funct_genome

Joint IGB-DKFZ-MolToolsPractical Course:
Analysis and Interpretation of Complex Transcript Data
27 June 2005 - 30 June 2005DKFZ, Heidelberg
There are no costs involved but for accommodation and travel expenses, which will be the responsibility of the participants.
Emphasis of the course will be hands-on data analysis. The participants are encouraged to bring along their own data for analysis.
For more details see: www. dkfz.de/funct_genome