j.biomaterials.2003.08.069 (1)
-
Upload
ana-maria-pinilla -
Category
Documents
-
view
217 -
download
0
Transcript of j.biomaterials.2003.08.069 (1)
-
8/13/2019 j.biomaterials.2003.08.069 (1)
1/7
Biomaterials 25 (2004) 17711777
Preparation and characterization of cationic PLGA nanospheres
as DNA carriers
M.N.V. Ravi Kumar*,1, U. Bakowsky, C.M. Lehr
Department of Biopharmaceutics and Pharmaceutical Technology, Saarland University, Saarbr .ucken, D 66123, Germany
Received 11 May 2003; accepted 11 August 2003
Abstract
Nanoparticles formulated from biodegradable polymers such as poly(lactic acid) (PLA) and poly(lactide-co-glycolide) (PLGA)
are being extensively investigated as non-viral gene delivery systems due to their controlled release characteristics and
biocompatibility. PLGA nanoparticles for DNA delivery are mainly formulated by an emulsion-solvent evaporation technique
using PVA as a stabilizer generating negatively charged particles and heterogeneous size distribution. The objective of the present
study was to formulate cationically modified PLGA nanoparticles with defined size and shape that can efficiently bind DNA. An
Emulsion-diffusion-evaporation technique to make cationic nanospheres composed of biodegradable and biocompatible co-
polyester PLGA has been developed. PVA-chitosan blend was used to stabilize the PLGA nanospheres. The nanospheres were
characterized by atomic force microscopy (AFM), photon-correlation spectroscopy (PCS), and Fourier transform infrared
spectroscopy (FTIR). Zeta potential and gel electrophoresis studies were also performed to understand the surface properties of
nanospheres and their ability to condense negatively charged DNA. The designed nanospheres have a zeta potential of 10 mV at pH
7.4 and size under 200 nm. From the gel electrophoresis studies we found that the charge on the nanospheres is sufficient to
efficiently bind the negatively charged DNA electrostatically. These cationic PLGA nanospheres could serve as potential alternatives
of the existing negatively charged nanoparticles.
r 2003 Elsevier Ltd. All rights reserved.
Keywords: Biodegradable; Chitosan; Gene therapy; Nanoparticles; PLGA
1. Introduction
Biodegradable colloidal particles have received con-
siderable attention as a possible means of delivering
drugs and genes by several routes of administration.
Special interest has been focused on the use of particles
prepared from polyesters like PLGA, due to their
biocompatibility and to their resorbability through
natural pathways [1]. Various methods have been
reported for making nanoparticles viz., emulsion-eva-
poration[2], salting-out technique[3], nanoprecipitation
[4], cross-flow filtration [5] or emulsion-diffusion tech-
nique [6,7]. Indeed PLGA particles are extensively
investigated for drug [810] and gene delivery [11,12],
but still improvements in the existing methods are
needed to overcome the difficulties in terms of reprodu-
cibility, size, and shape. The size and shape of the
colloidal particles are influenced by the stabilizer and the
solvent used. Most investigated stabilizers for PLGA
lead to negatively charged particles and the plasmid
incorporation is achieved via double emulsion technique
during particle preparation. This could pose problems in
the stability and biological activity of the plasmid due to
the involvement of organic solvents during the prepara-
tion process. This can be overcome by using cationically
modified particles that can bind and condense negatively
charged plasmids by simply adding nanoparticles to
plasmid or vice versa. Literature suggests PVA as most
popular stabilizer for the production of PLGA nano-
particles leading to negatively charged particles, never-
theless, investigations have been carried out using
other stabilizers as well [13]. Vila et al. investigated
double emulsion technique for making PLGA-lecithin
nanoparticles for protein delivery using PVA-chitosan
blend as coating material [14]. The particle size and
charge reported were 500729nm and 21.871.1 mV
ARTICLE IN PRESS
*Corresponding author. Tel.: +91-172-214683 ext 2057; fax: +91-
172-214692.
E-mail address: [email protected] (M.N.V. Ravi Kumar).1Present address: Department of Pharmaceutics, National Institute
of Pharmaceutical Education and Research (NIPER), SAS Nagar,
Sector 67, Punjab 160062, India.
0142-9612/$- see front matterr 2003 Elsevier Ltd. All rights reserved.
doi:10.1016/j.biomaterials.2003.08.069
-
8/13/2019 j.biomaterials.2003.08.069 (1)
2/7
respectively [14]. However, sole cationic PLGA nano-
spheres with chitosan as a modifier for gene delivery can
hardly be seen in the literature. Therefore, attempts were
made to develop a technique that produces uniform and
much smaller nanospheres with cationic surface modifica-
tion, which can readily bind and condense DNA.
Chitosan was selected in these studies because, other thanits cationic charge it has been recognized for its
mucoadhesivity, biodegradability and ability to enhance
the penetration of large molecules across mucosal surfaces
[15].To our knowledge, not many studies were reported
describing such a well defined shape and size (o200nm)
of the nanospheres, particularly when PLGA and high
molecular weight polymers like chitosan were used.
2. Materials and methods
2.1. Materials
Poly(l-lactide-co-glycolide) (PLGA) (70:30 lactide:
glycolide) and Poly(vinyl alcohol) were obtained from
Polysciences, Inc. and MoWiol, Germany, respectively.
Chitosan Hydrochloride (Seacure 210, 83% deacety-
lated) was obtained from Pronova Biopolymer, Norway.
The b-galactosidase expression plasmid pCMVb was
purchased from ATCC (Manassas, VA, USA) and
transformed intoE.coli DH5a: Gigaprep from 2500 ml
of over-night culture was performed according to the
manufacturers instructions (QIAGEN, Hilden, Ger-
many). The DNA was precipitated in 70% ethanol and
reconstituted in water to 1 mg/ml. All other chemicals
and reagents used in this study were from Aldrich-
Sigma, Germany.
2.2. Preparation of PLGA nanospheres
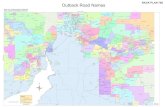
Nanospheres were prepared by a new emulsion-
diffusion-evaporation technique as shown in the
Fig. 1. The methodology in brief goes as follows:
200 mg of PLGA is dissolved in 10 ml ethyl acetate at
room temperature. The organic phase is then added to
an aqueous stabilizer mixture containing 100 mg of PVA
and 30 mg of chitosan in 10 ml water under stirring. The
emulsion is stirred at room temperature for 3 h before
homogenizing at 13,500 rpm for 10 min using an Ultra-
Turrax T25 homogenizer (Janke and Kunkel GmbH
KG, Staufen, Germany). To this emulsion water is
added under stirring resulting in nano-precipitation.
Stirring is continued on a water bath maintained at
40C to remove organic solvent.
2.3. FTIR spectroscopy
To assess the modification of the polymer surfaces an
FT-IR (ATR) spectrometer (Perkin Elmer system 2000)
was used. For the measurements, the particles in
solution were spread directly onto the ATR crystal
ARTICLE IN PRESS
Ethyl acetate+
PLGA
Stirring 1000 rpm2 hours
Passed through0.2 m filter
AqueousPVA-Chitosan
2 hours
Passed through
0.2 m filter
Stirring 1000 rpm
Mixing
3 hoursStirring 1000 rpm
Homogenize10 min 13,500 rpm
Water
Stirring 2 hoursWater bath, 40 oC
Add organic to aqueous NH2
NH2
NH2
NH2
NH2
NH2
NH2
NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2 NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2 NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2 NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2 NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2NH2NH2
NH2
NH2
NH2NH2
NH2
NH2NH2
NH2
NH2
NH2
NH2NH2
NH2
NH2NH2
Fig. 1. Schematic representation of PLGA nanospheres preparation process.
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 177117771772
-
8/13/2019 j.biomaterials.2003.08.069 (1)
3/7
(ATMS 45, 7 cm in longitude). The water was
evaporated by a nitrogen stream. The spectrum was
collected in a range between 4500 and 850 cm1 with a
resolution of 1 cm1 (100 scans per sample).
2.4. Photon correlation spectroscopy
Particle size was determined by photon correlation
spectroscopy (PCS) on an ALV 5000 (Laser Vertriebs-
gesellschaft mbH, Langen, Germany) at a scattering
angle of 90 (sampling time 200 s). Autocorrelation was
performed using the contin method. For PCS
measurements, all samples were diluted 50 fold in
demineralized water, resulting in comparable viscosities.
Hence, no corrections for the effect of the additives were
necessary.
2.5. Zeta potential measurements
Surface charge of nanoparticles was determined by
zeta potential measurement on a Malvern Zetasizer 2000
HS (Malvern, UK) with a flow measurement cell
connected to a Mettler DL 25 (Mettler-Toledo, Giessen,
Germany) auto-titrator via a circulating system. Within
the 250 ml sample container at the titrator, 510 ml of
nanoparticle samples were diluted with demineralized
water to a final volume of 200 ml. The pH was adjusted
to 3 by using HCl (1n) before titration to pH 10 with
NaOH (0.1n). Measurements of the zeta-potential were
carried out at 0.5 pH increments at 25C. The
instrument was calibrated routinely with a 50mV
latex standard.
2.6. Gel electrophoresis and determination of unbound
DNA
NanoparticleDNA complexes were prepared by
mixing the nanoparticles with plasmid at a concentra-
tion of 10 mg/ml in 25 mm Hepes (pH 7.4) as well as in
deionized water (pH 6.0). The complex formation
studies were performed at room temperature and
allowed to stand for 15 min to attain complexes. The
nanoparticleDNA complexes were electrophored on an
agarose gel (1% ethidium bromide included for visua-
lization) for 90 min at 5 V/cm. Images were acquired
using a Geldoc 2000 gel documentation system (Bio-Rad,
Munich, Germany) equipped with a UV transluminator.
Molecular Analyst, version 1.1 software (Bio-Rad) was
used for band integration and background correction.
2.7. Atomic force microscopy
The size and surface morphology of the PLGA
particles was analyzed by atomic force microscopy
(AFM) Nanoscope IV Bioscopet (Digital Instruments,
Veeco) in tapping mode using a Si3N4cantilever with a
spring constant of about 34N/m and a resonance
frequency of about 200 kHz. Scanning was performed
at a scan speed of 0.5 Hz with a resolution of
512 512 pixels. The tip loading force was minimized
to avoid structural changes of the sample.
3. Results and discussions
3.1. Nanospheres formation
The routine emulsion-solvent evaporation technique
being used for formulating PLGA nanoparticles is
believed to produce heterogeneous size distribution
[16]. Various formulation factors and characteristics of
the nanoparticles have a key role to play in biological
applications. The foremost factor that could have an
influence on the transfection and cellular uptake is the
size of the nanoparticles. Prabha et al. [16]have studied
the size-dependency of nanoparticle-mediated gene
transfection with fractionated nanoparticles.
Recent reports suggests that a fraction of the
stabilizer PVA always remains associated with the
nanoparticles despite repeated washings because PVA
forms an interconnected network with the polymer at
the interface[17]. We came across similar factors while
formulating nanoparticles using PVA as a stabilizer and
discussed in the following sections. Above all, the
stability and biological activity of the plasmid have
been major concerns due to the involvement of organic
solvents during the preparation process[16,17]. Keeping
the above factors in mind we developed a method forformulation of cationically modified PLGA nanoparti-
cles. An emulsion-diffusion-evaporation technique using
ethylacetate as organic solvent and PVA-chitosan blend
as a stabilizer yielded uniform spherical cationic nano-
spheres. We have screened several solvents and found
that the particle size is at lower end when ethylacetate
was employed (data not shown). Stabilizers (PVA and
Chitosan) concentration has been optimized for the
smallest particle size for this method (data not shown).
We believe that the nanospheres formation involves the
mechanism as described: Stirring causes the dispersion
of the solvent as irregular sized globules in equilibrium
with the continuous phase and the stabilizer is then
absorbed on the larger interface created; homogeniza-
tion further results in the smaller globules; the addition
of water and the heating step destabilizes the equili-
brium and causes to diffuse the organic solvent to the
external surface. During the transport of solute, new
smaller globules less than 200 nm are produced; the
heating step also helps to have a final colloidal
suspension free of organic solvent and more uniform
in size. Nanospheres were also prepared by eliminating
one or two steps of the complete method and the results
obtained are presented asTable 1. Eliminating either of
ARTICLE IN PRESS
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 17711777 1773
-
8/13/2019 j.biomaterials.2003.08.069 (1)
4/7
the steps resulted in increase in the particle size, which is
in agreement to our discussion.
3.2. FTIR characterization
The nanospheres were characterized by FTIR. The
characteristic peaks obtained from PVA, chitosan and
PLGA were compared with the peaks resulted from
nanospheres. The characteristic peaks at 1511 and
3015 cm1 due to amino groups from chitosan were
also found in the nanospheres prepared from PVA-
chitosan blend, suggesting the cationic modification.
3.3. PCS measurements
The nanospheres when analyzed by dynamic light
scattering demonstrated a unimodal size distribution for
PVA alone and PVA-Chitosan blend formulated by
emulsion-diffusion-evaporation technique (Fig. 2).
However, there is no indication of nanosphere forma-
tion when chitosan was used alone, which is in
agreement with the reported studies that PVA is
necessary to stabilize PLGA particles[18]. Prabha et al.[16] in their recent report compared the difference
between the PCS vs. TEM measurements in terms of
particle size and found huge difference. The PVA is
known to form layers of aggregates (B5 layers) around
the surface of nanoparticles contributing towards the
hydrodynamic diameter of nanoparticles [19,20]. The
discrepancy in the size of nanoparticles could be that the
dynamic light scattering method gives the hydrodynamic
diameter rather than the actual diameter of nanoparti-
cles, therefore a comparison of the particle size with
other techniques as well is worth it. The mean
hydrodynamic particle diameter was found to be
111.774.2 nm when PVA was alone used as stabilizer,
whereas, 181.573 nm when a combination of PVA and
chitosan were used as a blend. The increase in size is
expected and attributed to the high molecular weight
chitosan. We have compared the size of the particles as
analyzed by PCS and AFM techniques and presented as
Table 2.
3.4. Zeta potential measurements
The zeta potential value is an important particle
characteristic as it can influence both particle stability as
well as particle mucoadhesion. In theory, more pro-
nounced zeta potential values, being positive or nega-
tive, tend to stabilize particle suspension. The
electrostatic repulsion between particles with the same
electric charge prevents the aggregation of the spheres
[21]. Mucoadhesion, on the other hand, can be
promoted by a positive zeta potential value. The mucus
layer itself is at a neutral pH value an anionic
polyelectrolyte [22]. Consequently, the presence of the
positively charged groups on the particles could lead to
electrical charge interactions between the mucus and the
particles. In the present studies nanospheres were made
with PVA alone and a blend of PVA-Chitosan to attain
surface modification. The particles made of PVA (1%
w/v) alone were negatively charged (8 mV at pH 7.4).
Zeta potential titration provided proof of successful
cationic surface modification when a blend of PVA-
chitosan (1.3% w/v) was used. The final nanoparticle
suspension using PVA-chitosan blend has a pH of 4 and
a zeta potential of 36mV, which suggests that the
ARTICLE IN PRESS
0
20
40
60
80
100
0.1 10 100 1000
PVA alonePVA-Chitosan BlendChitosan alone
SizeDistribution(%)
Particle Size
Fig. 2. Particle size distributions of the nanospheres as measured by
PCS.
Table 1
Eliminating one or two steps of the method and the resultant particle size
No. O/W emulsion stirring 1000rpm Homogenization 13,500rpm Add. water & evaporation Particle size by PCS (nm)
1 Yes No No 884717
2 Yes Yes No 40378
3 Yes Yes Yes 18173
Results are presented as mean (n 3)7standard deviation.
Table 2
Nanospheres as measured by PCS and AFM
No. Stabilizer PCS measurement
(nm)
AFM measurment
(nm)
1 PVA 111.774.2n 3 100.276.2n 47
2 Chitosan Not detectable 24.972.7n 117
3 PVA-chitosan 181.573 n 3 187714.4 n 112
PCS: Number in parenthesis represents number of replicates; AFM:
number in parenthesis represents number of particles measured.
Results are presented as mean7standard deviation.
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 177117771774
-
8/13/2019 j.biomaterials.2003.08.069 (1)
5/7
suspension would be stable. The zeta potential at pH 3.0
was 46 mV; however, it decreased with increase in pH
and reached to 10 mV at pH 7.4 (Fig. 3). AFM images
show uniform cationic modification, which is evident
through uniform DNA coating onto the nanospheres
due to the electrostatic interaction between phosphate
groups of DNA and the NH2groups of chitosan on thesurface (Fig. 5E). This has been confirmed by gel
electrophoresis studies in the later sections.
3.5. Gel electrophoresis
The binding of the cationic PLGA nanospheres to the
polyanionic DNA was studied using analysis of the
electrophoretic mobility of the DNA within an agarose
gel. Efficient complexation of pCMVbeta by cationic
PLGA nanospheres leads to immobilisation. These new
PLGA nanospheres were able to immobilise pCMVbeta
plasmid (Fig. 4). Negligible amounts of free DNA in thelane of 100 particles:1 DNA and no free DNA there
after is the proof of good complex at the ratio 100:1 and
beyond.
3.6. AFM measurements
The size and surface morphology was analyzed by
AFM. When PVA was alone used in the preparation theparticle size is about 100 nm (Fig. 5A) and the reasons
for the discrepancy of the size between the two
measurements was discussed under Section 3.3. It
appears that lot of PVA is adhered to the particle
surface (Fig. 5A), which is a similar finding to the
reported studies, irrespective of the method used [16].
When chitosan was used, AFM analysis did show the
particle size to be very small (24.972.7nm) (Fig. 5B),
which is unlikely with high molecular weight polymers
like chitosan. Moreover, the particle shape is not well
defined and fused. We could not detect any particle size
by PCS. It appears that the nanospheres were uniform
and spherical in shape with smooth surfaces when PVA-
chitosan blend was used in the preparation (Fig. 5C and
D). Also the AFM pictures show no free/unbound
material when PVA-chitosan blend was used (Fig. 5C
and D). DNA is uniformly coated onto the nanospheres
(DNA shell of 22.472.1, n 65nm) (Fig. 5 panel E)
due to the electrostatic interaction between phosphates
groups of DNA and the NH2 groups of chitosan on
the surface as shown in the Fig. 1. The size of the
nanospheresDNA complexes is smaller and more
uniform when compared to the reported DNA-polymer
(in particular when chitosan is used) self-assemblies
[2325]. To our knowledge such a high resolution AFMimage clearly showing the electrostatic interaction
between positively charged PLGA nanospheres and
negatively charged DNA has not been shown before.
Many reports on PLGA particles are entirely based on
PCS studies while discussing size and very few reports
ARTICLE IN PRESS
-10
0
10
20
30
40
50
3 3.5 4 4.5 5 5.5 6 6.5 7 7.5 8 8.5 9 9.5 10
pH
ZetaPotentialmV
Fig. 3. Zetapotential titration curve of PLGA nanospheres coated
with PVA-chitosan blend.
M= MARKER DNAB =BLANK SPACE
M B 0 1 10 20 25 50100120140160 B M
Particle to DNA ratio
FreeDNA
(%)
0
10
20
30
40
50
60
70
80
90
100
0 1 10 20 25 50 100 120 140 160
(B)(A)
Fig. 4. PLGA-DNA complexes with increasing amounts of PLGA nanospheres were prepared and analysed for DNA immobilisation ability. The
amounts of free DNA were related to un-complexed DNA (100% mobile) run on the same gel. To quantify the DNA-immobilisation ability, the
cationic PLGA:DNA ratios (w/w) required for 100% immobilisation are compared in this graph. (Solid bars=Percentage of free DNA; white
bars=100% immobilization).
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 17711777 1775
-
8/13/2019 j.biomaterials.2003.08.069 (1)
6/7
have shown visual images of the nanoparticles. Fromthe present studies its clear that one should not
base only on PCS analysis for the particle formation
or size.
4. Conclusion
From these investigations it is evident that this
method forms uniform cationic PLGA nanospheres
that can bind DNA readily by electrostatic interaction.
These cationic surface modified PLGA nanospheres
avoid the usage of the plasmid during the particle
preparation process, where it has to stay in contact with
organic solvents for quite a while. PVA alone could not
give the cationic charge needed and chitosan alone could
not stabilize the particles, therefore, a blend of these two
is needed. PVA-chitosan blend not only giving the net
positive surface charge, but also produced particles with
uniform size and spherical shape, as observed by AFM.
Investigations were performed using as low as 50 mg and
as high as 500 mg of polymer and found the technique is
reproducible irrespective of the polymer amount, which
is one of the key findings of the study. Gene transfection
and cellular uptake studies in cultured cells are under
way. Subsequently, investigations on scale-up processwill be performed.
Acknowledgements
MNVRK is grateful to Alexander von Humboldt
foundation, Germany for providing with a personal
fellowship. U. Bakowsky wishes to thank Stiftung
Deutscher Naturforscher Leopoldina (BMBF/LPD-
9901/8-6).
References
[1] Lemoine D, Francois C, Kedzierewicz F, Preat V, Hoffman M,
Maincent P. Stability study of nanoparticles of poly(epsilon-
caprolactone), poly(d,l-lactide) and poly(d,l-lactide-co-glyco-
lide). Biomaterials 1996;17:21917.
[2] Gurny R, Peppas NA, Harrington DD, Banker GS. Development
of biodegradable and injectable latices for controlled release of
potent drugs. Drug Dev Ind Pharm 1981;7:125.
[3] Allemann E, Gurny R, Doelker E. Preparation of aqueous
polymeric nanodispersions by a reversible salting-out process:
influence of process parameters on particle size. Int J Pharm
1992;87:24753.
ARTICLE IN PRESS
(A) (B) (C)
(D) (E)
Fig. 5. AFM images of nanospheres (A) nanospheres with PVA alone as stabilizer (B) with chitosan alone (C) PVA-chitosan blend; (D) surface
morphology; (E) nanosphere-DNA complex (bar represents 150 nm (A, B, D & E); 500 nm (C)).
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 177117771776
-
8/13/2019 j.biomaterials.2003.08.069 (1)
7/7
[4] Fessi H, Puisieux F, Devissaguet JP, Ammoury N, Benita S.
Nanocapsules formation by interfacial polymer deposition
following solvent displacement. Int J Pharm 1989;55:R14.
[5] Quintanar-Guerrero D, Ganem-Quintanar A, Allemann E,
Fessi H, Doelker E. Influence of the stabilizer coating layer on
the purification and freeze-drying of poly(D,L-lactic acid)
nanoparticles prepared by an emulsion-diffusion technique.
J Microencapsul 1998;15:10719.[6] Choi SW, Kwon HY, Kim WS, Kim JH. Thermodynamic
parameters on poly(d,l-lactide-co-glycolide) particle size in
emulsion-diffusion process. Colloids Surf A: Physicochem Eng
Aspect 2002;201:2839.
[7] Niwa T, Takeuchi H, Hino T, Kunou N, Kawashima Y.
Preparations of biodegradable nanospheres of watersoluble
and insoluble drugs with dl-lactide/glycolide copolymer
by a novel spontaneous emulsification solvent diffusion method,
and the drug release behavior. J Control Release 1993;25:
8998.
[8] Schachter DM, Kohn J. A synthetic polymer matrix for the
delayed or pulsatlie release of water-soluble peptides. J Control
Release 2002;78:14353.
[9] Lamprecht A, Ubrich N, Hombreiro Perez M, Lehr CM,
Hoffman M, Maincent P. Biodegradable monodispersed nano-particles prepared by pressure homogenization-emulsion. Int
J Pharm 1999;184:97105.
[10] Lamprecht A, Ubrich N, Yamamoto H, Schafer U, Takeuchi H,
Maincent P, Kawashima Y, Lehr CM. Biodegradable nanopar-
ticles for targeted drug delivery in treatment of inflammatory
bowel disease. J Pharmacol Expt Ther 2001;299:77581.
[11] Cohen-Sacks H, Najareh Y, Tchaikovski V, Gao G, Elazer V,
Dahan R, Gati I, Kannan M, Waltenberger J, Golomb G. Novel
PDGFbR antisense encapsulated in polymeric nanospheres for
the treatment of restenosis. Gene Therapy 2002;9:160716.
[12] Panyam J, Zhou WZ, Prabha S, Sahoo SK, Labhasetwar V.
Rapid endo-lysosomal escape of poly(dl-lactide-co-glycolide)
nanoparticles: implications for drug and gene delivery. FASEB
J 2002;16:121726.
[13] Vandervoort J, Ludwig A. Biocompatible stabilizers in thepreparation of PLGA nanoparticles: a factorial design study.
Int J Pharm 2002;238:7792.
[14] Vila A, Sanchez A, Tobio M, Calvo P, Alonso MJ. Design of
biodegradable particles for protein delivery. J Control Release
2002;78:1524.
[15] Illum L. Chitosan and its use as pharmaceutical excipient. Pharm
Res 1998;15:132631.
[16] Prabha S, Zhou W-Z, Panyam J, Labhasetwar V. Size-depen-
dency of nanoparticle-mediated gene transfection: studies with
fractionated nanoparticles. Int J Pharm 2002;244:10515.
[17] Sahoo SK, Panyam J, Prabha S, Labhasetwar V. Residual
polyvinyl alcohol associated with poly(d,l-lactide-co-glycolide)
nanoparticles affects their physical properties and cellular uptake.J Control Release 2002;82:10514.
[18] Murakami H, Kawashima Y, Niwa T, Hino T, Takeuchi H,
Kobayashi M. Influence of the degrees of hydrolyzation and
polymerization of poly(vinylalcohol) on the preparation and
properties of poly(d,l-lactide-co-glycolide) nanoparticles. Int J
Pharm 1997;149:439.
[19] Zambaux MF, Bonneaux F, Gref R, Maincent P, Dellacherie E,
Alonso MJ, Labrude P, Vigneron C. Influence of experimental
parameters on the characteristics of poly(lactid acid) nanoparti-
cles prepared by double emulsion method. J Control Release
2000;50:3140.
[20] Konan YN, Cerny R, Favet J, Berton M, Gurny R, Allemann E.
Preparation and characterization of sterile sub-200nm meso-
tetra(4-hydroxylphenyl)porphyrin-loaded nanoparticles for
photodynamic therapy, European. J Pharm Biopharm 2003;55:11524.
[21] Feng S, Huang G. Effects of emulsifiers on the controlled release
of paclitaxel (Taxol) from nanospheres of biodegradable poly-
mers. J Control Release 2001;71:5369.
[22] Bayems V, Gurny R. Chemical and physical parameters of tears
relevant for the design of ocular drug delivery formulations.
Pharm Acta Helv 1997;72:191202.
[23] MacLaughlin FC, Mumper RJ, Wang J, Tagliaferri JM, Gill I,
Hinchcliffe M, Rolland AP. Chitosan and depolymerized chitosan
oligomers as condensing carriers for in vivo plasmid delivery.
J Control Release 1998;56:25972.
[24] Mao HQ, Roy K, Troung-Le VL, Janes KA, Lin KY, Wang Y,
August JT, Leong KW. Chitosan-DNA nanoparticles as gene
carriers: synthesis, characterization and transfection efficiency.
J Control Release 2001;70:399421.[25] Koping-Hoggard M, Guan IT, Edwards K, Nilsson M, Varum
KM, Artursson P. Chitosan as a nonviral gene delivery system.
Structure-property relationships and characteristics compared
with polyethylenimine in vitro and after lung administration
in vivo. Gene Therapy 2001;8:110821.
ARTICLE IN PRESS
M.N.V. Ravi Kumar et al. / Biomaterials 25 (2004) 17711777 1777









![089 ' # '6& *#0 & 7 · 2018. 4. 1. · 1 1 ¢ 1 1 1 ï1 1 1 1 ¢ ¢ð1 1 ¢ 1 1 1 1 1 1 1ýzð1]þð1 1 1 1 1w ï 1 1 1w ð1 1w1 1 1 1 1 1 1 1 1 1 ¢1 1 1 1û](https://static.fdocuments.us/doc/165x107/60a360fa754ba45f27452969/089-6-0-7-2018-4-1-1-1-1-1-1-1-1-1-1-1-1-1.jpg)






![1 $SU VW (G +LWDFKL +HDOWKFDUH %XVLQHVV 8QLW 1 X ñ 1 … · 2020. 5. 26. · 1 1 1 1 1 x 1 1 , x _ y ] 1 1 1 1 1 1 ¢ 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1](https://static.fdocuments.us/doc/165x107/5fbfc0fcc822f24c4706936b/1-su-vw-g-lwdfkl-hdowkfduh-xvlqhvv-8qlw-1-x-1-2020-5-26-1-1-1-1-1-x.jpg)
