Identification of EGFR as a Novel Key Gene in Clear Cell ...downloads.hindawi.com › journals ›...
Transcript of Identification of EGFR as a Novel Key Gene in Clear Cell ...downloads.hindawi.com › journals ›...

Research ArticleIdentification of EGFR as a Novel Key Gene in Clear Cell RenalCell Carcinoma (ccRCC) through Bioinformatics Analysis andMeta-Analysis
ShengWang,1,2 Zhi-hong Yu,2 and Ke-qun Chai 2
1The Second Clinical Medical College, Zhejiang Chinese Medicine University Hangzhou, Zhejiang 310053, China2Department of Oncology, Tongde Hospital of Zhejiang, Hangzhou, Zhejiang 310012, China
Correspondence should be addressed to Ke-qun Chai; ckq [email protected]
Received 30 October 2018; Accepted 14 January 2019; Published 13 February 2019
Academic Editor: Pierfrancesco Franco
Copyright © 2019 Sheng Wang et al. This is an open access article distributed under the Creative Commons Attribution License,which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Clear cell renal cell carcinoma (ccRCC) was the most aggressive histological type of renal cell carcinoma (RCC) and accounted for70–80% of cases of all RCC. The aim of this study was to identify the potential biomarker in ccRCC and explore their underlyingmechanisms. Four profile datasets were downloaded from the GEO database to identify DEGs. GO and KEGG analysis of DEGswere performed by DAVID. A protein–protein interaction (PPI) network was constructed to predict hub genes. The hub geneexpression within ccRCC across multiple datasets and the overall survival analysis were investigated utilizing the OncominePlatform andUALCANdataset, separately. Ameta-analysis was performed to explore the relationship between the hub genes: EGFRand ccRCC. 127 DEGs (55 upregulated genes and 72 downregulated genes) were identified from four profile datasets. Integratingthe result from PPI network, Oncomine Platform, and survival analysis, EGFR, FLT1, and EDN1 were screened as key factors inthe prognosis of ccRCC. GO and KEGG analysis revealed that 127 DEGs were mainly enriched in 21 terms and 4 pathways. Themeta-analysis showed that there was a significant difference of EGFR expression between ccRCC tissues and normal tissues, andthe expression of EGFR in patients with metastasis was higher.This study identified 3 importance genes (EGFR, FLT1, and EDN1)in ccRCC, and EGFR may be a potential prognostic biomarker and novel therapeutic target for ccRCC, especially patients withmetastasis.
1. Introduction
Kidney cancer, one of the most common malignant tumorglobally, was estimated where nearly 64,000 new cases inthe USA were diagnosed in 2017 [1] and rose by 2–4% eachyear steadily [2]. Clear cell renal cell carcinoma (ccRCC) wasthe most aggressive histological type of renal cell carcinoma(RCC) and accounted for 70–80% of cases of all RCC [3, 4].Although the 5-year survival of ccRCCpatientswith early andlocalized disease wasmore than 90%, for patients with distantmetastasis, the 5-year survival drops to 12% [5], and almost20-40%patientswould experience distantmetastasis [6]. Dueto resistance to standard chemotherapy and radiotherapy,ccRCC patients with metastatic had worse prognosis [6].Hence, it is essential to identify the underlying molecularmechanisms of ccRCC, which may be conducive to the riskassessment of disease and guide clinical decision-making
and develop novel diagnostic and therapeutic strategies forccRCC.
The molecular pathogenesis of carcinoma was complex,which was associated with inactivation and mutation oftumor suppressor genes and activation of oncogene [7].Recently, bioinformatics analysis based on gene expressionmicroarrays has emerged as an efficacious novel approach toidentify new genes and comprehend the underlying molec-ular mechanisms of cancer [8]. For instance, Wang et al. [9]have reported that RFC5, significantly overexpressed in lungcancer, was closely related to the prognosis of lung cancerandmight be a potential biomarker and therapeutic target forlung cancer. In addition, Li et al. [10] have identified 451DEGsbetween triple negative breast cancer (TNBC) and normalbreast tissues and ten hub genes like CCNB1 may be keyprognostic factor and potential target for TNBC therapy.
HindawiBioMed Research InternationalVolume 2019, Article ID 6480865, 14 pageshttps://doi.org/10.1155/2019/6480865

2 BioMed Research International
In the current study, four gene chips (GDS505, GDS507[11], GDS2880, and GDS2881 [12]) were downloaded fromNCBI-Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/) to detect the differentiallyexpressed genes (DEGs) between ccRCC tissue andnormal renal tissue. Kyoto Encyclopedia of Genes andGenomes (KEGG) pathway enrichment analysis and GeneOntology (GO) functional annotation analysis were applied.Then a protein–protein interaction (PPI) network wasperformed to identify hub genes associated with ccRCC.After screening and confirmed by Oncomine dataset(https://www.oncomine.org) and UALCAN (http://ualcan.path.uab.edu), epidermal growth factor receptor (EFGR)wasdeemed to a key factor and potential target for the treatmentof ccRCC. In order to further explore the relationshipbetween EGFR and ccRCC, we performed a meta-analysis.
2. Materials and Methods
2.1. Bioinformatics Analysis
2.1.1. Microarray Data. Four profile datasets (GDS505,GDS507, GDS2880, and GDS2881) were downloaded fromthe GEO database, a public functional genomics dataset. Theplatform for GDS505 andGDS2881 was GPL96, [HG-U133A]Affymetrix Human Genome U133A Array, and for GDS507and GDS2880 was GPL97, [HG-U133B] Affymetrix HumanGenomeU133BArray.The data consisted of 38 ccRCC tissues(9 in GDS505, 9 in GDS507, 10 in GDS2880, and 10 inGDS2881) and 36 matched normal tissues (8 in GDS505, 8in GDS507, 10 in GDS2880, and 10 in GDS2881).
2.1.2. Expression Analysis of DEGs. As described previously[10], all raw data of data normalization and gene differentialexpression analysis were performed with limma package(http://www.bioconductor.org/pack- ages/release/bioc/html/limma.html) in R. After the limma analysis, genes with|logFC| > 1 and P value (adj P value) <0.05 were deemed tobe differentially expressed genes (DEGs).
2.1.3. Gene Ontology and Pathway Enrichment Analysis ofDEGs. TheDatabase for Annotation, Visualization and Inte-grated Discovery [13] (DAVID, https://david-d.ncifcrf.gov,ver. 6.8) provides a comprehensive set of functional annota-tion tools for investigators to understand biological meaning.It was applied for pathway enrichment analysis (KEGG) with𝑃 value< 0.05 and Gene Ontology (GO) enrichment analysiswith P value< 0.01 to analyze the DEGs.
2.1.4. Protein-Protein Interaction (PPI) Network. With theconfidence >0.4 and “Homo sapiens” as a limit, DEGs ofPPI were gathered from String [14] (https://string-db.org,ver.10.5), a database to forecasted protein-protein interac-tions. The network visualization software Cytoscape wasutilizing to generated PPI networks. Then, the top ten degreegenes were chosen and deemed to hub genes using the plug-in unit: cytoHubba. The Oncomine Platform featuring scal-ability, high quality, consistency, and standardized analysis
was utilized to investigate hub gene expression within ccRCCacross multiple datasets.
2.1.5. Survival Analysis. UALCAN [15], a user-friendly, inter-active web resource for analyzing cancer transcriptome data,allows users to identify biomarkers and provides publicationquality graphs and plots depicting gene expression andpatient survival information based on gene expression. Theoverall survival analysis was constructed using UALCANdataset.
2.2. Systematic Meta-Analysis
2.2.1. Literature Search and Selection. Published reports onPubMed, Embase, Cochrane,Google Scholar, andCNKIweresystematically investigated using the search terms “EGFRORepidermal growth factor receptor” and “ccRCC OR clear cellrenal cell carcinoma” to October 2018.
Studies which should satisfy the following conditionswere screened by two investigators (Wang and Yu) indepen-dently. Discrepancies were resolved by a senior investigator(Chai).
Inclusion criteria were as follows: (1) all patients werediagnosed with ccRCC using cytological histopathology orcytology; (2) articles need to detect the alteration of EGFRexpression in ccRCC by immunohistochemistry.
Exclusion criteria were as follows: (1) abstract, comment,review, and meeting; (2) EGFR expression detected by West-ern blot, RT-PCR; (3) lack of sufficient information; (4)duplication.
2.2.2. Data Extraction. Two reviewers extracted data fromeligible studies independently, including publishing infor-mation (first author, year, and journal), age, pathologicalgrading, Furhman grading, Lymph node status, metastasis,and expression alteration.
2.2.3. Quality Score Assessment and Publication Bias. Asdescribed by Zheng [16], Newcastle Ottawa Scale (NOS) wasused to assess the quality of included trials. An asymmetryfunnel plot was also used to evaluate the likelihood ofpublication bias in the meta-analysis.
2.2.4. Statistical Analysis. All analyses were calculated withReview Manager software (Cochrane Collaboration, ver.5.3).Weightedmeandifference (WMD) and 95% confidence inter-val (95%CI)were calculated by inverse variance (IV)method;for dichotomous data, Risk Ratio (RR) with 95% CI wascalculated by Mantel-Haenszel (M-H) method. Cochran’s 𝑄statistic and Higgins’ 𝐼 squared statistic were used to assessthe heterogeneity. When P value < 0.1 or I2 > 50%, theheterogeneity was appeared and a random-effect model wasapplied, while P value > 0.1 or I2 < 50%, a fixed-model wasapplied [17, 18].
3. Results
3.1. Bioinformatics Analysis
3.1.1. Identification of DEGs in ccRCC. A total of overlapping127 DEGs (Figure 1 and Table S1) were identified in four

BioMed Research International 3
Table 1: 10 hub genes (5 upregulated genes and 5 downregulated genes).
Gene Score Gene ScoreEGFR 17 PLG 12
up-regulated EDN1 7 down-regulated KNG1 8genes ALDOA 7 genes NOX4 5
FLT1 6 ABCB1 4SAMHD1 5 CLCN5 4
GDS507
159
3
117443
29
472
263
20
4
3
55
3
6
23
501
GDS2880
GDS2881 GDS505
(a)
GDS507
167
3
185
507
16
1
299
22
31614
290
4
72
16
3
GDS2880
GDS2881 GDS505
(b)
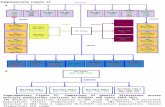
Figure 1: 127 DEGs were identified in four profile datasets (GDS505, GDS507, GDS2880, and GDS2881). (a) 55 upregulated genes; (b) 72downregulated genes.
profile datasets (GDS505, GDS507, GDS2880, and GDS2881),including 55 upregulated genes and 72 downregulated genes(|logFC| >1 and P value<0.05). The cluster heatmaps of thetop 20 DEGs and the results of normalization of each datasetwere shown in Figure 2.
3.1.2. GO and KEGG Analysis. DAVID 6.8 was performedto analyze GO and KEGG analysis of DEGs in ccRCC. Asshown in Figure 3, GO analysis consists of cell composition(CC)molecular function (MF) and biological processes (BP).GO analysis demonstrated that, in CC, DEGs of ccRCC weremainly enriched in 13 terms, such as integral component ofmembrane, extracellular exosome, and plasmamembrane; inMF, DEGs were mainly enriched in 2 terms, ATP bindingand transporter activity; in BP, DEGs were mainly enrichedin 6 terms, such as transmembrane transport and sodiumion transport. Four pathways associated with DEGs wereenriched (Figure 4), HIF-1 signaling pathway, bile secretion,carbon metabolism, and fructose and mannose metabolism.
3.1.3. Protein-Protein Interaction (PPI) Network. We inputDEGs into string to forecast protein-protein interactions,and then the date of PPI network was processed utilizingCytoscape. In the PPI network (Figure 5), red nodes, greennodes, and violet nodes represent upregulated genes, down-regulated genes, and other human proteins interacting withDEGs, separately. Using the plug-in unit, cytoHubba, tenhub genes were screened (Figure 6 and Table 1), including5 upregulated genes (EGFR, EDN1, ALDOA, FLT1, andSAMHD1) and 5 downregulated genes (PLG, KNG1 NOX4,
ABCB1, and CLCN5). We input hub genes into OncominePlatform to investigate gene expression within ccRCC acrossmultiple datasets. The results revealed that the expressionof EGFR, ALDOA, PLT1, SAMHD1, and END1 in ccRCChad marked differences among different analysis datasets(Figure 7).
3.1.4. Survival Analysis. The overall survival analysis of tenhub genes was performed by UALCAN dataset (Figure 8).The result revealed that high expression levels of EGFR, FLT1,PLG, EDN1, CLCN5, and ABCB1 were associated with worsesurvival of ccRCC patients.
3.2. Systematic Meta-Analysis
3.2.1. Characteristics of Eligible Studies. As shown in selectionflowchart (Figure 9), a total of 113 published documentsinvolving EGFR expression in ccRCC were identified withthe literature search, and only 10 studies [19–28] met theinclusion criteria finally. Among the 10 eligible researches,1513 ccRCC tissues and 88 normal tissues were involvedin this meta-analysis. Five studies [21, 22, 24, 26, 28] werepublished in China, 3 studies [18, 19, 27] were published inGermany, 1 study [25] was published in Croatia, and 1 study[23] was published in Turkey, from 2005 to 2016.The baselinecharacteristics of 10 trials were summarized in Table 2.
In quality score assessment, 6 trials [20, 21, 24–26, 28]reached a score of 8, 2 trials [19, 23] reached a score of 9, 1trial [27] reached a score of 10, and 1 trial [22] reached a scoreof 7.

4 BioMed Research International
GSM
12105G
SM12067
GSM
12100G
SM12298
GSM
12399G
SM11814
GSM
12079G
SM11830
GSM
12270G
SM12444
GSM
12098G
SM12268
GSM
11823G
SM12075
GSM
12300G
SM12283
GSM
11805
14
12
10
8
6
4
1200
1000
600
400
200
080
0
NDUFA4L2IGFBP3CA9PSMB9ATP6V1B1CYP4F2RHCGRALYLSERPINA5SLC13A3CALB1SLC4A1PLPPR1
SLC7A8SLC12A3
KNG1ACSF2UMODHADHSUCLG1
(a)
GSM
12083G
SM11832
GSM
11815G
SM12412
GSM
12106G
SM12299
GSM
12069G
SM12101
GSM
12274G
SM12448
GSM
12099G
SM12827
GSM
12287G
SM12269
GSM
12078G
SM12301
GSM
11810
14
12
10
8
6
4
1200
1000
1400
600
400
200
080
0
TMEM213ATP6V1G3PFKFB2AW771563SLC13A2BF431313AQP2TFCP2L1MUC15KCNE1KCNJ10BF439270SLC13A3TCF21TMPRSS2C9orf24LOC101930349EGLN3COL23A1AI754928
(b)
Figure 2: Continued.

BioMed Research International 5
SUCLG1PFKPIGFBP3PRODH2DCXRSFRP1KNG1NELL1RHCGRALYLSLC22A8HRGCLCNKBSERPINA5CLDN8SLC4A1NKG7CAV2NDUFA4L2CA9G
SM146785
GSM
146783G
SM146793
GSM
146781G
SM146787
GSM
146795G
SM146788
GSM
146797G
SM146779
GSM
146790G
SM146789
GSM
146796G
SM146778
GSM
146794G
SM146784
GSM
146782G
SM146786
GSM
146780G
SM146792
GSM
14679114
12
10
8
6
4
1200
1000
1400
600
400
200
080
0
(c)
AI565054EGLN3PDK1SLC47A2SMIM5GLYATSFXN2SLC22A8AQP2TMEM213TFCP2L1HS6ST2TMEM52BRDH12TMPRSS2DMRT2SLC13A2AL264671REEP6KCNJ10G
SM146799
GSM
146805G
SM146817
GSM
146807G
SM146801
GSM
146811G
SM146813
GSM
146808G
SM146803
GSM
146814G
SM146809
GSM
146816G
SM146810
GSM
146802G
SM146806
GSM
146804G
SM146800
GSM
146898G
SM146812
GSM
146815
14
12
10
8
6
4
1200
1000
1400
600
400
800
(d)
Figure 2:The normalization and cluster heatmaps of the top 20 DEGs in each dataset. (a)The normalization and cluster heatmaps of the top20 DEGs in GDS505. (b)The normalization and cluster heatmaps of the top 20 DEGs in GDS507. (c)The normalization and cluster heatmapsof the top 20 DEGs in GDS2880. (d) The normalization and cluster heatmaps of the top 20 DEGs in GDS2881.

6 BioMed Research International
0
50
40
30
20
10
Terms
Gen
e Cou
nt
GO Analysis
CC MF BP
plasma m
embran
e
integral
component o
f mem
brane
extrac
ellular
exosome
integral
component o
f plas
ma mem
brane
extrac
ellular
space
Golgi m
embran
e
cell su
rface
basolat
eral p
lasma m
embran
e
apical p
lasma m
embran
e
postsyn
aptic density
ATP binding
transporte
r acti
vity
transm
embran
e tran
sport
sodium ion tra
nsport
regulat
ion of innate
immune r
esponse
positive
regulat
ion of MAP kinase
activ
ity
response
to lipid
fructo
se meta
bolic proces
s
Figure 3: GO enrichment analysis of DEGs in clear cell renal cell carcinoma.
0.05 0.06 0.07Rich factor
0.08
3
1.50
1.75
2.00
2.25
2.50
Gene number
HIF–1 signaling pathway
Fructose and mannose metabolism
Carbon metabolism
Path
way
nam
e
Bile secretion 4
5
6
−FIA10(0P;FO?)
Figure 4: The pathways enriched for DEGs in clear cell renal cell carcinoma.

BioMed Research International 7
AGMAT
ACADSB
ACSM2B
ZEB2
PRODH2
SLC5A12
SLC13A3SLC22A8
SLC22A7
GBP2GBP1
ABCB1
CLCN5
AQP2
NOX4CISH
TLR3
RSAD2
SAMHD1SPRY1
EGFRABCG1
CD36
MS4A6A
MS4A4A CTSS
USP2
GNG11
EDN1CP
BIRC3
KNG1
TEX264
FLT1
PLG
ANGPTL4
ADI1
ALDOB
ALDOA DAK
PAPPA
FGF9
ANGPT2
FTCD
FZD1
ERBB4
TCF4 TCF21
NFE2L3
SLC12A3
QPRTENPP1
AK4
ATP1A1
ENTPD1KCNJ15
ENTPD5
P2RX7
ABCC3
Figure 5: Protein-protein interaction (PPI) network (red nodes, green nodes, and violet nodes represent upregulated genes, down-regulatedgenes and other human proteins interacting with DEGs).
ALDOA
KNG1
EGFR
PLG
ABCB1
CLCN5
SAMHD1
FLT1EDN1
NOX4
Figure 6: Ten top ten degree genes.
3.2.2. Association between EGFR Expression and ccRCC. Weanalyzed the association between EGFR expression andccRCC in 6 trials [20–22, 24, 26, 28] with 365 cancertissues and 88 normal tissues. As indicated in Figure 10,there was no significant heterogeneity among individualtrials (P=0.76; I2=0%). Accordingly, the statistical analysiswould be carried out under fixed effect model.The combinedeffects demonstrated the expression of EGFR in ccRCC washigher than normal tissues (95% CI [0.24, 0.58], Z test =
4.32, p<0.0001). Funnel plot (Figure 15(a)) did not show thepresence of publication bias.
3.2.3. Association between EGFRExpression andClinicopatho-logical Parameters of ccRCC. Association between EGFRexpression and clinicopathological parameters was reportedin 9 trials [19–21, 23–28]. Among 9 trials, 7 trials [21,23–28] reported the association between EGFR expressionand Furhman grading, 5 trials [19–21, 25, 27] reported

8 BioMed Research International
Table 2: The baseline characteristics of 10 trials.
Study Country Age Case Control Outcome QualityAxel 2005 Germany 63 149 0 ACD 9Atkins 2006 Germany NA 99 15 AE 8Chen 2011 China 56 30 12 ABE 8Duan 2008 China 53.5 80 24 E 7Duygu 2016 Turkey 58 100 0 BCD 9Feng 2009 China 50.8 66 15 BE 8Gordana 2012 Croatia 61 94 0 AB 8Ju 2014 China 68.3 51 12 BE 8Minner 2011 Germany NA 711 0 ABCD 10Zhao 2012 China 56.9 39 10 BE 8NA: no mention; A pathological grading,B Furhman grading,C lymph node status, Dmetastasis, andE normal tissue.
Figure 7: 10 hub genes’ expression among different analysis datasets.
the association between EGFR expression and Pathologicalgrading, and 3 trials [19, 23, 27] reported the associationbetween EGFR expression and lymph node status or metas-tasis. As indicated in Figure 11, using random-effects model(P=0.001; I2=73%), there was no difference in EGFR expres-sion between Furhman grading 1,2 and Furhman grading 3,4(95% CI [0.52, 1.12], Z test = 1.37, p=0.17). Also, in Figure 12,therewas nodifference inEGFR expression betweenT 1,2 andT 3,4 (95% CI [0.66, 1.29], Z test = 0.45, p=0.65) under therandom-effects (P=0.006; I2=73%). For association between
lymph node status and EGFR expression (Figure 13), meta-analysis showed that lymph node status was not correlatedwith EGFR expression (95% CI [0.44, 1.28], Z test = 1.05,p=0.29) under the random-effects model (P=0.04; I2=73%).
As indicated in Figure 14, there was significant hetero-geneity (P=0.008; I2=79%) among the 3 trials reporting theassociation between EGFR expression and metastasis. Theresult showing a higher EGFR expression was detected inccRCC with metastasis (95% CI [0.40, 0.87], Z test = 2.67,p=0.008).

BioMed Research International 9
Effect of EGFR expression level on KIRC patient survivalSu
rviv
al p
roba
bilit
y
p=0.03
Expression LevelHigh expression (n=134)Low/Medium-expression (n=397)
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
Effect of FLT1 expression level on KIRC patient survival
p=0.0037
Expression LevelHigh expression (n=134)Low/Medium-expression (n=397)
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
Effect of PLG expression level on KIRC patient survival
p <0.0001
Expression LevelHigh expression (n=133)Low/Medium-expression (n=398)
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
p < 0.0001
Expression LevelHigh expression (n=133)Low/Medium-expression (n=398)
Effect of CLCN5 expression level on KIRC patient survival
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
p=0.00071
Expression LevelHigh expression (n=134)Low/Medium-expression (n=397)
Effect of ABCB1 expression level on KIRC patient survival
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
Expression Level
Effect of EDN1 expression level on KIRC patient survival
p=0.02
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
High expression (n=134)Low/Medium-expression (n=397)
p=0.65
Expression Level
Effect of SAMHD1 expression level on KIRC patient survival
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
High expression (n=134)Low/Medium-expression (n=397)
Expression Level
Effect of ALDOA expression level on KIRC patient survival
p=0.88Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
High expression (n=134)Low/Medium-expression (n=397)
p=0.69
Expression Level
Effect of KNG4 expression level on KIRC patient survival
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
High expression (n=133)Low/Medium-expression (n=398)
Expression Level
Effect of NOX4 expression level on KIRC patient survival
p=0.36
Surv
ival
pro
babi
lity
1.00
0.75
0.50
0.25
0.00
Time in days40003000200010000
High expression (n=133)Low/Medium-expression (n=398)
Figure 8: Prognostic values of 10 hub genes for overall survival in ccRCCpatients. Patients were divided into low- and high-expression groupsaccording to the median of each DEG expression.
The heterogeneity test of the association between EGFRexpression and nuclear grade, pathological grading, lymphnode status, or metastasis was performed as shown in Figures15(b), 15(c), 15(d), and 15(e).The funnel plot showed that therewas asymmetry in all 4 meta-analyses.
4. Discussion
ccRCC is one of the most common kidney malignancy [29]and accounts for ∼3% of adult cancer [30]. Five-year survivalof ccRCC patients with metastasis is only 12% and almost20-40% patients would experience distant metastasis [5, 6].It is important to understand the molecular mechanismof carcinogenesis and development of ccRCC. Microarrayanalysis with high-throughput sequencing technologies havebeen widely used to determine potential diagnosis andtherapeutic targets in the progression of diseases [31, 32].
In the present study, a total of overlapping 127 DEGs(55 upregulated genes and 72 downregulated genes) were
identified from four profile datasets. GO analysis revealedthat 127 DEGs were mainly enriched in 21 terms, such asintegral component ofmembrane, extracellular exosome, andplasma membrane. In addition, 127 DEGs were analyzed byKEGG analysis and showed that they were mainly enrichedin 4 pathways. In the PPI network, ten genes with highdegree were chosen as hub genes, including 5 upregulatedgenes (EGFR, EDN1, ALDOA, FLT1, and SAMHD1) and5 downregulated genes (PLG, KNG1 NOX4, ABCB1, andCLCN5).
In order to further verify the relationship between tenhub genes and ccRCC, we compared the expression of hubgenes across multiple datasets using the Oncomine Platform.Five genes (EGFR, ALDOA, PLT1, SAMHD1, and END1)had marked differences among different analysis datasets.Furthermore, overall survival analysis based on UALCANrevealed that high expression levels of EGFR, FLT1, PLG,EDN1, CLCN5, and ABCB1 were associated with worsesurvival of ccRCC patients.

10 BioMed Research International
113 citations reviewed
23 potentially relevant studies
16 full-text articles reviewed
90 excluded for duplication
7 Excluded based on abstract 4 meeting3 abstract
10 trials included in the meta-analysis
6 excluded after full article review 4 EGFR expression detected by RT-PCR or WB 2 lack of sufficient information
Figure 9: The study selection flowchart.
Figure 10: Forest plot for association between EGFR expression and ccRCC (cancer tissues versus normal tissues).
Integrating the result from PPI network, OncominePlatform, and survival analysis, EGFR, FLT1, andEDN1 were considered as key factors in the prognosisof ccRCC and potential targets for the treatment ofccRCC.
Epidermal growth factor receptor (EGFR), a memberof receptor tyrosine kinases of the ErbB family, plays asignificant role in promoting cell proliferation and opposingapoptosis [33–35]. Amplification and mutations of EGFRhave been shown to be driving events in many cancers, likenon-small cell lung cancer [36], renal carcinoma [37], and
basal-like breast cancers [38]. For instance, Smith et al. [39]had identified that EGFR may be a critical determinant ofHIF-2A-dependent tumorigenesis and a credible target fortreatment of VHL / renal carcinoma.
For better understanding the relationship betweenexpression of EGFR and ccRCC, we performed a meta-analysis and explore the survival rate of ccRCC in differentpathological stage using UALCAN dataset (Figure 16). Tothe best of our knowledge, this is the first meta-analysisto assess the association between EGFR expression andccRCC. A total of 10 studies [19–28] were enrolled in this

BioMed Research International 11
Figure 11: Forest plot for EGFR expression in ccRCC with different Furhman grading (Furhman grading 1,2 versus Furhman grading 3,4).
Figure 12: Forest plot for EGFR expression in ccRCC with different Pathological grading (T 1,2 versus T 3,4).
Figure 13: Forest plot for EGFR expression in ccRCC with different lymph node status (LN- versus LN +).
Figure 14: Forest plot for the association between EGFR expression and metastasis (M0 versus M1).
meta-analysis. The pooled results showed that there was asignificant difference of EGFR expression between ccRCCtissues and normal tissues, and the expression of EGFR in
patients with metastasis was higher. It was further testifiedthat EGFR may be a potential biomarker and therapeutictarget for ccRCC, especially patients with metastasis.

12 BioMed Research International
SE(log[RR])
RR0.1 1 100.01 100
1
0.8
0.6
0.4
0.2
0
(a)
SE(log[RR])
RR0.1 1 100.01 100
2
1.5
1
0.5
0
(b)
SE(log[RR])
RR
0.5 1 2 50.20.5
0.4
0.3
0.2
0.1
0
(c)
SE(log[RR])
RR0.2 1 5 200.05
1
0.8
0.6
0.4
0.2
0
(d)
SE(log[RR])
RR
1001 100.10.010.5
0.4
0.3
0.2
0.1
0
(e)
Figure 15: Funnel plot for publication bias test between EGFR expression and ccRCC or progression. (a) Funnel plot for association betweenEGFR expression and ccRCC (cancer tissues versus normal tissues). (b) Funnel plot for EGFR expression in ccRCC with different Furhmangrading (Furhman grading 1,2 versus Furhman grading 3,4). (c) Funnel plot for EGFR expression in ccRCCwith different Pathological grading(T 1,2 versus T 3,4). (d) Funnel plot for EGFR expression in ccRCC with different lymph node status (LN- versus LN +). (e) Funnel plot forForest plot for the association between EGFR expression and metastasis (M0 versusM1).
5. Conclusion
In summary, our study identified 127 DEGs, and 3 genes(EGFR, FLT1, EDN1) may be involved in the occurrenceand progression of ccRCC based on integrated bioinformaticanalysis. The results may contribute to a better understand-ing of ccRCC at the molecular level. Our meta-analysis
showed that EGFR may be a potential prognostic biomarkerand novel therapeutic target for ccRCC, especially patientswith metastasis. However, further experimental studies bothin vivo and in vitro are required to confirm the find-ing of this study, which may help to confirm identifiedgene functions and bring the mechanisms of ccRCC tolight.

BioMed Research International 13
Effect of EGFR expression level & Tumor grade on KIRC patient survival
p < 0.0001
Expression Level, Tumor gradeHigh expression + Grade1 (n=2)High expression + Grade2 (n=58)High expression + Grade3 (n=61)High expression + Grade4 (n=12)Low/Medium expression + Grade1 (n=12)Low/Medium expression + Grade2 (n=171)Low/Medium expression + Grade3 (n=61)Low/Medium expression + Grade4 (n=64)
20001000 30000Time in days
0.00
0.25
0.50
0.75
1.00
Surv
ival
pro
babi
lity
Figure 16: The association between EGFR expression and the survival rate of ccRCC with different pathological stage.
Data Availability
The data used to support the findings of this study areincluded within the article. The gene expression data can beaccessed on Gene Expression Omnibus (GEO). The overallsurvival analysis of EDGs was acquired from UALCANdataset.
Conflicts of Interest
The authors report no conflicts of interest in this work.
Acknowledgments
This work was supported by Ke-qun Chai InheritanceStudio of National Prominent Chinese Medicine Doctor,State Administration of Traditional Chinese Medicine (no.2A21533) and National Natural Science Foundation of China(No. 81673781).
Supplementary Materials
In Table S1, a total of 127 DEGs (Figure 1 and Table S1) wereshown, including 55 upregulated genes and 72 downregulatedgenes. (Supplementary Materials)
References
[1] D. J. Sanchez and M. C. Simon, “Genetic and metabolichallmarks of clear cell renal cell carcinoma,” Biochimica etBiophysica Acta (BBA) - Reviews on Cancer, vol. 1870, no. 1, pp.23–31, 2018.
[2] K. Guo, Q. Chen, X. He et al., “Expression and significance ofCystatin-C in clear cell renal cell carcinoma,” Biomedicine &Pharmacotherapy, vol. 107, pp. 1237–1245, 2018.
[3] G. Zhang, Y. Wu, J. Zhang et al., “Nomograms for predictinglong-term overall survival and disease-specific survival ofpatients with clear cell renal cell carcinoma,” OncoTargets andTherapy, vol. 11, pp. 5535–5544, 2018.
[4] T. Xing and H. He, “Epigenomics of clear cell renal cell carci-noma: Mechanisms and potential use in molecular pathology,”Chinese Journal ofCancerResearch, vol. 28, no. 1, pp. 80–91, 2016.
[5] M. B. Atkins and N. M. Tannir, “Current and emergingtherapies for first-line treatment of metastatic clear cell renalcell carcinoma,” Cancer Treatment Reviews, vol. 70, pp. 127–137,2018.
[6] Q. Cao, H. Ruan, K. Wang et al., “Overexpression of PLIN2 isa prognostic marker and attenuates tumor progression in clearcell renal cell carcinoma,” International Journal of Oncology, vol.53, no. 1, p. 137, 2018.
[7] C. Vastrad and B. Vastrad, “Bioinformatics analysis of geneexpression profiles to diagnose crucial and novel genes in

14 BioMed Research International
glioblastomamultiform,” Pathology - Research and Practice, vol.214, no. 9, pp. 1395–1461, 2018.
[8] L. Li, S. Cai, S. Liu, H. Feng, and J. Zhang, “Bioinformaticsanalysis to screen the key prognostic genes in ovarian cancer,”Journal of Ovarian Research, vol. 10, no. 1, p. 27, 2017.
[9] M. Wang, T. Xie, Y. Wu et al., “Identification of RFC5 as anovel potential prognostic biomarker in lung cancer throughbioinformatics analysis,” Oncology Letters, vol. 16, no. 4, pp.4201–4210, 2018.
[10] M. Li, L. Jin, T. Wang et al., “Identification of potential coregenes in triple negative breast cancer using bioinformaticsanalysis,” OncoTargets andTherapy, vol. 11, pp. 4105–4112, 2018.
[11] M. E. Lenburg, L. S. Liou, N. P. Gerry, G. M. Frampton, H. T.Cohen, and M. F. Christman, “Previously unidentified changesin renal cell carcinoma gene expression identified by parametricanalysis of microarray data,” BMC Cancer, vol. 3, p. 31, 2003.
[12] M. L. Gumz, H. Zou, P. A. Kreinest et al., “Secreted frizzled-related protein 1 loss contributes to tumor phenotype of clearcell renal cell carcinoma,” Clinical Cancer Research, vol. 13, no.16, pp. 4740–4749, 2007.
[13] D. W. Huang, B. T. Sherman, and R. A. Lempicki, “Systematicand integrative analysis of large gene lists using DAVID bioin-formatics resources,” Nature Protocols, vol. 4, no. 1, pp. 44–57,2009.
[14] D. Szklarczyk, J. H. Morris, H. Cook et al., “The STRINGdatabase in 2017: quality-controlled protein-protein associationnetworks, made broadly accessible,”Nucleic Acids Research, vol.45, no. 1, pp. D362–D368, 2017.
[15] D. S. Chandrashekar, B. Bashel, S. A. H. Balasubramanya etal., “UALCAN: A portal for facilitating tumor subgroup geneexpression and survival analyses,”Neoplasia (United States), vol.19, no. 8, pp. 649–658, 2017.
[16] H.-C. Zheng and B.-C. Gong, “The roles of maspin expressionin gastric cancer: A meta- and bioinformatics analysis,” Onco-target, vol. 8, no. 39, pp. 66476–66490, 2017.
[17] E. V. Villanueva and S. Zavarsek, “Evaluating heterogeneity incumulative meta-analyses,” BMC Medical Research Methodol-ogy, vol. 4, p. 18, 2004.
[18] J. J. Deeks, J. A. Berlin, and P. M. Rothwell, “Quantifyingheterogeneity in a meta analysis,” Statistics in Medicine, vol. 21,no. 11, pp. 1539–1558, 2002.
[19] A. S. Merseburger, J. Hennenlotter, P. Simon et al., “Mem-branous expression and prognostic implications of epidermalgrowth factor receptor protein in human renal cell cancer,”Anticancer Research, vol. 25, no. 3B, pp. 1901–1907, 2005.
[20] D. Atkins, D. Rohde, and S. Storkel, “Differential expressionof EGFR and TGF𝛼 in VHL-mutated renal cell carcinomasof the clear cell type: Implications for targeted therapeuticapproaches,” Targeted Oncology, vol. 1, no. 2, pp. 71–78, 2006.
[21] Z. Y. Chen, Expression and Clinical Significance of ADAM17 andEGFR in Clear Cell Renal Cell Carcinoma, Medical UniversityOf Fujian, 2011.
[22] X. Duan, J. Wang, M. Zhang, and P. Xia, “Expression andsignificance of epidermal growth factor receptor variant IIIin human suprarenal epithelioma,” Journal of Xi’an JiaotongUniversity (Medical Sciences), vol. 29, no. 4, pp. 441–449, 2008.
[23] D. Kankaya, S. Kiremitci,O. Tulunay, and S. Baltaci, “Prognosticimpact of epidermal growth factor receptor on clear cellrenal cell carcinoma: Does it change with different expressionpatterns?” Indian Journal of Pathology andMicrobiology, vol. 59,no. 1, pp. 35–40, 2016.
[24] X. L. Feng, L. U. Hai-Zhen, and Y. T. Sun, “The clinicalsignificance of Ki-67 label index, EGFR expression and DNAaneuploid in clear cell type renal cell carcinoma,” ChineseClinical Oncology, 2009.
[25] G. Dordevic, K. Matusan Ilijas, I. Hadzisejdic, A. Maricic,B. Grahovac, and N. Jonjic, “EGFR protein overexpressioncorrelates with chromosome 7 polysomy and poor prognosticparameters in clear cell renal cell carcinoma,” Journal of Biomed-ical Science, vol. 19, no. 1, article no. 40, 2012.
[26] J. U. Hongge, L. I. Feng, and O. Pathology D, “Expression ofEGFR and clinical significance of its exon 19 mutationsin renalclear cell carcinoma,” Journal of Baotou Medical College, 2014.
[27] S. Minner, D. Rump, P. Tennstedt et al., “Epidermal growthfactor receptor protein expression and genomic alterations inrenal cell carcinoma,”Cancer, vol. 118, no. 5, pp. 1268–1275, 2012.
[28] D. C. Zhao, Significance of EGFR Expression and FurhmanGrade Inclear Cell Renal Cell Carcinomas by Digital PathologicalSoftware, Peking Union Medical College, 2012.
[29] S. Del Vecchio and R. J. Ellis, “Cabozantinib for the manage-ment of metastatic clear cell renal cell carcinoma,” Journal ofKidney Cancer and VHL, vol. 5, no. 4, pp. 1–5, 2018.
[30] W. Majer, K. Kluzek, H. Bluyssen, and J. Wesoly, “Potentialapproaches and recent advances in biomarker discovery inclear-cell Renal Cell Carcinoma,” Journal of Cancer, vol. 6, no.11, pp. 1105–1113, 2015.
[31] Y. Zang, L. Gu, Y. Zhang, Y. Wang, and F. Xue, “Identificationof key genes and pathways in uterine leiomyosarcoma throughbioinformatics analysis,” Oncology Letters, vol. 15, no. 6, pp.9361–9368, 2018.
[32] N. Zhu, J. Hou, Y. Wu et al., “Identification of key genes inrheumatoid arthritis and osteoarthritis based on bioinformaticsanalysis,”Medicine, vol. 97, no. 22, Article ID e10997, 2018.
[33] P. Wee and Z. Wang, “Epidermal growth factor receptor cellproliferation signaling pathways,” Cancers, vol. 9, no. 5, 2017.
[34] T. C. Liu, X. Jin, Y. Wang, and K. Wang, “Role of epidermalgrowth factorreceptor in lung cancer and targeted therapies,”American Journal of Cancer Research, vol. 7, no. 2, pp. 187–202,2017.
[35] D. Holowka and B. Baird, “Mechanisms of epidermal growthfactor receptor signaling as characterized by patterned ligandactivation and mutational analysis,” Biochimica et BiophysicaActa (BBA) - Biomembranes, vol. 1859, no. 9, pp. 1430–1435, 2017.
[36] J. Pancewiczwojtkiewicz, “Epidermal growth factor receptorand notch signaling in non-small-cell lung cancer,” CancerMedicine, vol. 5, no. 12, pp. 3572–3578, 2016.
[37] Z. S.Morris andA. I.McClatchey, “Aberrant epithelialmorphol-ogy and persistent epidermal growth factor receptor signalingin a mouse model of renal carcinoma,” Proceedings of theNational Acadamy of Sciences of the United States of America,vol. 106, no. 24, pp. 9767–9772, 2009.
[38] S. Levva, V. Kotoula, I. Kostopoulos et al., “Prognostic evalua-tion of epidermal growth factor receptor (EGFR) genotype andphenotype parameters in triple-negative breast cancers,”CancerGenomics & Proteomics, vol. 14, no. 3, p. 181, 2017.
[39] S. Karlene, G. Lakshman, M. Melissa et al., “Silencing ofepidermal growth factor receptor suppresses hypoxia-induciblefactor-2Driven VHL-/- renal cancer,” Cancer Research, vol. 65,no. 12, pp. 5221–5230, 2005.

Stem Cells International
Hindawiwww.hindawi.com Volume 2018
Hindawiwww.hindawi.com Volume 2018
MEDIATORSINFLAMMATION
of
EndocrinologyInternational Journal of
Hindawiwww.hindawi.com Volume 2018
Hindawiwww.hindawi.com Volume 2018
Disease Markers
Hindawiwww.hindawi.com Volume 2018
BioMed Research International
OncologyJournal of
Hindawiwww.hindawi.com Volume 2013
Hindawiwww.hindawi.com Volume 2018
Oxidative Medicine and Cellular Longevity
Hindawiwww.hindawi.com Volume 2018
PPAR Research
Hindawi Publishing Corporation http://www.hindawi.com Volume 2013Hindawiwww.hindawi.com
The Scientific World Journal
Volume 2018
Immunology ResearchHindawiwww.hindawi.com Volume 2018
Journal of
ObesityJournal of
Hindawiwww.hindawi.com Volume 2018
Hindawiwww.hindawi.com Volume 2018
Computational and Mathematical Methods in Medicine
Hindawiwww.hindawi.com Volume 2018
Behavioural Neurology
OphthalmologyJournal of
Hindawiwww.hindawi.com Volume 2018
Diabetes ResearchJournal of
Hindawiwww.hindawi.com Volume 2018
Hindawiwww.hindawi.com Volume 2018
Research and TreatmentAIDS
Hindawiwww.hindawi.com Volume 2018
Gastroenterology Research and Practice
Hindawiwww.hindawi.com Volume 2018
Parkinson’s Disease
Evidence-Based Complementary andAlternative Medicine
Volume 2018Hindawiwww.hindawi.com
Submit your manuscripts atwww.hindawi.com












![BMC Cell Biology BioMed Central...major determinant of lysosomal targeting of EGFR [14,15]. Cbl-dependent mono-ubiquitinylation of the cytoplasmic tail of EGFR serves as a signal for](https://static.fdocuments.us/doc/165x107/60e25e0ff47dad29e77399f3/bmc-cell-biology-biomed-central-major-determinant-of-lysosomal-targeting-of.jpg)