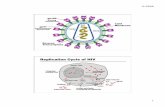
HIV GP120 Envelope Protein
description
Transcript of HIV GP120 Envelope Protein

HIV GP120 Envelope Protein
Objectives :
• Observing consensus sequences of different strains
• Structure analysis

GP120 FunctionGP120 involved in infection of human T-cells

HIV TYPES
HIV Type IHIV Type II
HIV Type III
SIV
Human viruses Simian virus

GP120 Sequence analysisSequence analysis shows a high mutation rate with preserved large consensus regions not only between strains, but also between different types

GP120 Sequence analysis
HIV Type I shows additional sequence of 7 aa in it’s consensus sequence in a consensus region in acids 27-33
consensus LWVTVFYGVPVWKEATTTLFCATDNK-------NTWATHACVPTDPDYQEVELNNVTENF 53
HIV1 LWVTVYYGVPVWHDADPVLFCASDAKahsteahNIWATQACVPTDPSPQEVFLPNVIESF 92

Two structures of GP120-CD4 from HIV 1 were analyzed
Hiv-1 Yu2 isolate Hiv-1 Hxbc2 isolate

Difference between the proteins
Score = 566 bits (1460), Expect = e-161Identities = 276/321 (85%), Positives = 289/321 (89%), Gaps = 8/321 (2%)
The similarity is shown, using protein blast

Structure of conserved region
GP120 Hxbc2 isolatePreserved region marked in yellow
CD4

Docking of GP120
Clustered solution ofDimers
With current approach GP120 exists as atrimers or tetramers on the membrane surface.According to J. Biol. Chem., Vol. 275, Issue 45, 35137-35145, November 10, 2000

Conclusions1. Preserved sequences found
2. GP120-CD4 theoretical docking performed, clustered solutions found
3. The results provide a basis for further research