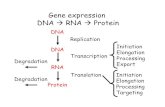
Cell(Division,(DNA(&(RNA,(Mendelian( Gene-cs,(Human(Gene ...
From Gene to Phenotype DNA molecule Gene 1 Gene 2 Gene 3 DNA strand (template) TRANSCRIPTION mRNA...
-
Upload
charlotte-peters -
Category
Documents
-
view
217 -
download
0
Transcript of From Gene to Phenotype DNA molecule Gene 1 Gene 2 Gene 3 DNA strand (template) TRANSCRIPTION mRNA...
From Gene to Phenotype
DNAmolecule
Gene 1
Gene 2
Gene 3
DNA strand(template)
TRANSCRIPTION
mRNA
Protein
TRANSLATION
Amino acid
A C C A A A C C G A G T
U G G U U U G G C U C A
Trp Phe Gly Ser
Codon
3 5
35
Lecture Outline 11/4/05
• The central dogma: – DNA->RNA->protein– Control of gene expression in prokaryotes vs
eukaryotes
• Transcription– Initiation, Elongation,Termination
• mRNA processing– Introns and exons
• Other types of RNA
In prokaryotes
– Transcription and translation occur simultaneously
TRANSLATION
TRANSCRIPTION DNA
mRNA
Ribosome
Polypeptide
Figure 17.3a
– RNA transcripts are modified before becoming true mRNA
– Transcription and translation occur in separate compartments of the cell
In Eukaryotes
Figure 17.3b
TRANSCRIPTION
RNA PROCESSING
TRANSLATION
mRNA
DNA
Pre-mRNA
Polypeptide
Ribosome
Nuclearenvelope
One Gene -> One Enzyme
EXPERIMENT
Class IMutants
Class IIMutants
Class IIIMutantsWild type
Minimal medium(MM)(control)
MM +Ornithine
MM +Citrulline
MM +Arginine(control)
Beadle and Tatum studied mutants of the bread mold Neurospora crassa and showed that each gene specified a particular enzyme
ArginineOrnithine CitrullinePrecursorArginine is an essential amino acid, required for growth
Normal cells can synthesisze arginine from precursors in
the minimal medium
Each mutant blocked a particular step of the pathway
Specific enzymes (arrows) catalyze each step
Which mutants can grow with which supplement?
Mut 1 Mut 2 Mut 3none 0 0 0Ornithine 1 0 0Citrulline 1 1 0Arginine 1 1 1
“One Gene -> One Enzyme”
“One Gene -> One Enzyme”
ArginineOrnithine CitrullinePrecursor
Which mutants can grow with which supplement?
Mut 1 Mut 2 Mut 3none 0 0 0Ornithine 1 0 0Citrulline 1 1 0Arginine 1 1 1
Mutant 2 can grow only if supplemented with citrulline or arginine
Therefore mutant 2 must not make the enzyme to convert Ornithine to Citrulline
One strand of DNA is copied to make messenger RNA
5’ . . . ATGAATGTC . . . 3’ coding3’ . . . TACTTACAG . . . 5’ template5’ . . . augaauguc-> . . 3’ RNA copy
This is the “sense” or “coding” strand, because it reads the same as the mRNA
This is the strand that is actually copied
Synthesis of an RNA Transcript
– Initiation
– Elongation
– Termination
Figure 17.7
PromoterTranscription unit
RNA polymerase
Start point
53
35
35
53
53
35
53
35
5
5
Rewound
RNA
RNA
transcript
3
3Completed RNA transcript
Unwound
DNA
RNA
transcript
Template strand of DNA
DNARNA polymerase binds to a promoter sequence
mRNA copy of gene is synthesized 5’ to 3’
Termination sequence causes transcription to stop
RNA PolymeraseElongation
RNApolymerase
Non-templatestrand of DNA
RNA nucleotides
3 end
C A E G C AA
U
T A G G T TA
AC
G
U
AT
CA
T C C A AT
TG
G
3
5
5
Newly madeRNA
Direction of transcription(“downstream) Template
strand of DNA
RNA synthesis
• RNA synthesis is similar to DNA synthesis except: – Does NOT need a primer– No proofreading
• Why not? mRNA has high turnover. An error in one molecule will not be inherited
• Both strands of DNA can serve as the template– Some genes are on one strand, other genes are on the other
Initiation in eukaryotesTRANSCRIPTION
RNA PROCESSING
TRANSLATION
DNA
Pre-mRNA
mRNA
Ribosome
Polypeptide
Eukaryotic promoters
T A T AAA A
ATAT T T T
TATA box Start point TemplateDNA strand
53
35
Several transcriptionfactors
Additional transcriptionfactors
Transcriptionfactors
53
35
1
2
3
Promoter
53
355
RNA polymerase IITranscription factors
RNA transcript
Transcription initiation complex
Several transcription factors must bind to promoter sequences upstream of the gene
Then RNA polymerase can bind
TATA box
RNA processing in eukaryotes
A modified guanine nucleotideadded to the 5 end
50 to 250 adenine nucleotidesadded to the 3 end
Protein-coding segment Polyadenylation signal
Poly-A tail3 UTRStop codonStart codon
5 Cap 5 UTR
AAUAAA AAA…AAA
TRANSCRIPTION
RNA PROCESSING
DNA
Pre-mRNA
mRNA
TRANSLATIONRibosome
Polypeptide
G P P P
5 3
1. Add 5’ cap2. Add poly A tail to 3’ end
A modified GTP is added, backwards, on the 5’ end
About 200 A’s added at 3’ end
RNA processing in eukaryotes
TRANSCRIPTION
RNA PROCESSING
DNA
Pre-mRNA
mRNA
TRANSLATION
Ribosome
Polypeptide
5 CapExon Intron
1
5
30 31
Exon Intron
104 105 146
Exon 3Poly-A tail
Poly-A tail
Introns cut out andexons spliced together
Codingsegment
5 Cap1 146
3 UTR5 UTR
Pre-mRNA
mRNA
3. Splice out introns
SpliceosomesRNA transcript (pre-mRNA)
Exon 1 Intron Exon 2
Other proteinsProtein
snRNA
snRNPs
Spliceosome
Spliceosomecomponents
Cut-outintron
mRNA
Exon 1 Exon 2
5
5
5
1
2
3
Special “small nuclear RNA” molecules do the splicing
Mature mRNA
Pre- mRNA
Most Eukaryotic genes have introns
13 kb
6 exons5 intronsExample: Red/Green colorblindness
Example: beta globin 3 exons2 introns
1.6 kb
Alternative splicing Make different mRNAs (and proteins) from
same gene by splicing out certain exons
Cell-type specific RNA splicing
Correspondence between exons and protein domains
GeneDNA
Exon 1 Intron Exon 2 Intron Exon 3
Transcription
RNA processing
Translation
Domain 3
Domain 1
Domain 2
Polypeptide
Four types of RNA
• mRNA– Messenger RNA, encodes the amino acid sequence of a
polypeptide
• rRNA– Ribosomal RNA, forms complexes with protein called ribosomes,
which translate mRNA to protein
• tRNA– Transfer RNA, transports amino acids to ribosomes during protein
synthesis
• snRNA– Small nuclear RNA, forms complexes with proteins used in
eukaryotic RNA processing
3ACCACGCUUAA
GACACCU
GC
*GUGU
CUGAG
GU
A
AA G
UC
AGACC
CGAGAG G
G
GACUCAU
UUAGGCG5
Hydrogenbonds
Anticodon
*
*
**
*
**
*
* **
RNA will fold to specific shapes
• Because RNA is single-stranded, parts of the molecule can base pair with other parts of the same molecule, causing it to fold into defined shapes.
• Some RNA molecules can even act as enzymes (ribozymes)