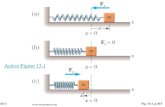
Fig. 13-1, p. 202. Fig. 13-1a, p. 202 Fig. 13-1b, p. 202.
-
Upload
roland-hancock -
Category
Documents
-
view
272 -
download
3
Transcript of Fig. 13-1, p. 202. Fig. 13-1a, p. 202 Fig. 13-1b, p. 202.

Fig. 13-1, p. 202

Fig. 13-1a, p. 202

Fig. 13-1b, p. 202

Fig. 13-2, p. 204

Fig. 13-2, p. 204
RR S
A Mice injected with live cells of harmless strain R do not die. Live R cells are in their blood.
B Mice injected with live cells of killer strain S die. Live S cells are in their blood.
C Mice injected with heat-killed S cells do not die. No live S cells are in their blood.
D Mice injected with live R cells plus heat-killed S cells die. Live S cells are in their blood.

Fig. 13-2, p. 204
C Mice injected with heat-killed S cells do not die. No live S cells are in their blood.
Stepped Art
B Mice injected with live cells of killer strain S die. Live S cells are in their blood.
S
A Mice injected with live cells of harmless strain R do not die. Live R cells are in their blood.
R
D Mice injected with live R cells plus heat-killed S cells die. Live S cells are in their blood.
R

Fig. 13-3, p. 205

Fig. 13-3a, p. 205

Fig. 13-3b, p. 205

Fig. 13-3b, p. 205
35S remains outside cellsVirus particle
coat proteins labeled with 35S
DNA being injected into bacterium
B In one experiment, bacteria were infected with virus particles labeled with a radioisotope of sulfur (35S). The sulfur had labeled only viral proteins. The viruses were dislodged from the bacteria by whirling the mixture in a kitchen blender. Most of the radioactive sulfur was detected in the viruses, not in the bacterial cells. The viruses had not injected protein into the bacteria.

Fig. 13-3c, p. 205

Fig. 13-3c, p. 205
Virus DNA labeled with 32P
32P remains inside cells
Labeled DNA being injected into bacterium
C In another experiment, bacteria were infected with virus particles labeled with a radioisotope of phosphorus (32P). The phosphorus had labeled only viral DNA. When the viruses were dislodged from the bacteria, the radioactive phosphorus was detected mainly inside the bacterial cells. The viruses had injected DNA into the cells—evidence that DNA is the genetic material of this virus.

Fig. 13-3, p. 205
Virus particle coat proteins labeled with 35S
DNA being injected into bacterium
35S remains outside cells
Virus DNA labeled with 32P
Labeled DNA being injected into bacterium
32P remains inside cells
Stepped Art

p. 205

Fig. 13-4, p. 206

Fig. 13-4a, p. 206

Fig. 13-4a, p. 206
adenine (A)
deoxyadenosine triphosphate, a purine

Fig. 13-4b, p. 206

Fig. 13-4b, p. 206
guanine (G)
deoxyguanosine triphosphate, a purine

Fig. 13-4c, p. 206

Fig. 13-4c, p. 206
thymine (T)deoxythymidine triphosphate, a pyrimidine

Fig. 13-4d, p. 206

Fig. 13-4d, p. 206
cytosine (C)deoxycytidine triphosphate, a pyrimidine

p. 207

Fig. 13-5, p. 207

Fig. 13-5a, p. 207

Fig. 13-5b, p. 207

Fig. 13-5b, p. 207
2-nanometer diameter
0.34 nanometer between each base pair
3.4-nanometer length of each full twist of the double helix
The numbers indicate the carbon of the ribose sugars (compare Figure 13.4). The 3’ carbon of each sugar is joined by the phosphate group to the 5’ carbon of the next sugar. These links form each strand’s sugar–phosphate backbone.
The two sugar–phosphate backbones run in parallel but opposite directions (green arrows). Think of one strand as upside down compared with the other.

Fig. 13-6, p. 208

Fig. 13-6, p. 208
A A DNA molecule is double-stranded. The two strands of DNA stay zippered up together because they are complementary: their nucleotides match up according to base-pairing rules (G to C, T to A).
B As replication starts, the two strands of DNA are unwound. In cells, the unwinding occurs simul- taneously at many sites along the length of each double helix.
C Each of the two parent strands serves as a template for assembly of a new DNA strand from free nucleotides, according to base-pairing rules (G to C, T to A). Thus, the two new DNA strands are complementary in sequence to the parental strands.
D DNA ligase seals any gaps that remain between bases of the “new” DNA, so a continuous strand forms. The base sequence of each half-old, half-new DNA molecule is identical to that of the parent DNA molecule.

Stepped ArtFig. 13-6, p. 208
D DNA ligase seals any gaps that remain between bases of the “new” DNA, so a continuous strand forms. The base sequence of each half-old, half-new DNA molecule is identical to that of the parent DNA molecule.
C Each of the two parent strands serves as a template for assembly of a new DNA strand from free nucleotides, according to base-pairing rules (G to C, T to A). Thus, the two new DNA strands are complementary in sequence to the parental strands.
B As replication starts, the two strands of DNA are unwound. In cells, the unwinding occurs simul- taneously at many sites along the length of each double helix.
A A DNA molecule is double-stranded. The two strands of DNA stay zippered up together because they are complementary: their nucleotides match up according to base-pairing rules (G to C, T to A).

p. 208

Fig. 13-7, p. 209

Fig. 13-8a, p. 209

Fig. 13-8a, p. 209
A Each DNA strand has two ends: one with a 5’ carbon, and one with a 3’ carbon. DNA polymerase can add nucleotides only at the 3’ carbon. In other words, DNA synthesis proceeds only in the 5’ to 3’ direction.

Fig. 13-8b, p. 209

Fig. 13-8b, p. 209
The parent DNA double helix unwinds in this direction.
Only one new DNA strand
is assembled continuously.
5’
The other new DNA strand is
assembled in many pieces.
3’
3’
Gaps are sealed by DNA ligase.
5’ 3’3’ 5’
B Because DNA synthesis proceeds only in the 5’ to 3’ direction, only one of the two new DNA strands can be assembled in a single piece.
The other new DNA strand forms in short segments, which are called Okazaki fragments after the two scientists who discovered them. DNA ligase joins the fragments into a continuous strand of DNA.

Fig. 13-9, p. 210

Fig. 13-9a, p. 210

Fig. 13-9a, p. 210
A A cow egg is held in place by suction through a hollow glass tube called a micropipette. The polar body (Section 10.5) and chromosomes are identified by a purple stain.

Fig. 13-9b, p. 210

Fig. 13-9b, p. 210
B A micropipette punctures the egg and sucks out the polar body and all of the chromosomes. All that remains inside the egg’s plasma membrane is cytoplasm.

Fig. 13-9c, p. 210

Fig. 13-9c, p. 210
C A new micropipette prepares to enter the egg at the puncture site. The pipette contains a cell grown from the skin of a donor animal.
skin cell

Fig. 13-9d, p. 210

Fig. 13-9d, p. 210
D The micropipette enters the egg and delivers the skin cell to a region between the cytoplasm and the plasma membrane.

Fig. 13-9e, p. 210

Fig. 13-9e, p. 210
E After the pipette is withdrawn, the donor’s skin cell is visible next to the cytoplasm of the egg. The transfer is complete.

Fig. 13-9f, p. 210

Fig. 13-9f, p. 210
F The egg is exposed to an electric current. This treatment causes the foreign cell to fuse with and empty its nucleus into the cytoplasm of the egg. The egg begins to divide, and an embryo forms. After a few days, the embryo may be transplanted into a surrogate mother.

Fig. 13-10, p. 211

Fig. 13-11a, p. 211

Fig. 13-11b, p. 211

Fig. 13-12, p. 213

p. 213

p. 213