Exploration of a potential FtsZ inhibitors as new scaffolds by Ligand and Structure based drug...
-
Upload
pavan-kumar -
Category
Science
-
view
28 -
download
0
Transcript of Exploration of a potential FtsZ inhibitors as new scaffolds by Ligand and Structure based drug...

1
Dr. Sachchidanand (HOD) Pharmacoinformatics Dept.
NIPER Hajipur.
Dr. Shailendra S. ChaudhaeryLecturer Pharmacoinformatics Dept.
NIPER Hajipur.
Under the supervision of :
PAVAN KUMAR
M.S.(Pharm.) Pharmacoinformatics
4th semester,NIPER, Hajipur.
Presented by :
Exploration of a potential FtsZ inhibitors as new scaffolds by Ligand and Structure based drug design methods for
development of novel anti-tubercular drugs

Tuberculosis (TB) is the chronic infectious disease caused by infection with species belonging to the Mycobacterium tuberculosis (Mtb). Mtb is slow-growing bacterium, which remains in dormant state for long period of time in the host, thus they are resistant to the effect of antibiotics. TB typically attacks the lungs but can also affect other parts of the body.
2M.tuberculosis
Introduction

The classical symptoms of active TB infection are: Chronic cough Blood-tinged sputum
Fever
Night sweats
Weight loss
Fatigue and finger clubbing
Classical Symptoms of TB

Global burden & epidemiology of TB• It is estimated that about 8.7 million new cases of TB (13% co-infected with
HIV) and 1.4 million people died from TB in 2011.• Recent statistics from WHO estimate that there are approximately 9.2
million new tuberculosis (TB) cases every year with a global mortality rate of 23%.

Category Name of drug Mechanism of action
First line drugs
Isoniazid Inhibition of fatty acid synthesis
Rifampicin Inhibition of protein synthesis
Pyrazinamide Disrupt plasma membrane, disrupt energy metabolism.
Ethambutol Inhibition of arabinogalactan synthesis
Streptomycin Inhibition of protein synthesis
Second line drugs
Fluoroquinolones Inhibit Nucleic Acid Synthesis
Cycloserine Inhibition of cell wall synthesis
Capreomycin Inhibition of protein synthesis
Para aminosalicylic acid Inhibition of folic acid and iron metabolism
Ethionamide Inhibition of fatty acid synthesis
Aminoglycosides
(Amikacin/kanamycin)
Inhibition of protein synthesis5
Antitubercular drugs

6
Different targets for Tuberculosis
5. FtsZ (Filamentous temperature-sensitive protein Z ) Drug Target for Tuberculosis.
Serial No.
Drug Target Antibiotic class Drug name Mechanism of action
1. DNA gyrase Fluoroquinolones
Moxifloxacin, gatifloxacin
Inhibition of protein synthesis
2. RNA polymerase Rifamycins Rifapentine Inhibits DNA-dependent RNA polymerase activity
3. Ribosome Oxazolidinones Linezolid,PNU-100480,AZD-5847
Inhibition of protein synthesis
4. ATP synthase Diarylquinoline TMC-207 Depletion of membrane energy
Multi-drug resistant Mtb is a major worldwide health problem. Therefore, it is need to develop new antibiotics with novel modes of action to overcome this emerging resistance problem.

7
1.It is an essential protein for bacterial cell division.
2. This protein is highly conserved, and identified in many bacteria.
3. It is not present in higher eukaryotes organism, So it shows that FtsZ inhibitors should not be toxic to human cells as well as high eukaryotes.
FtsZ is an emergent target

8
Filamentous temperature-sensitive protein Z (FtsZ)
FtsZ is the key protein of bacterial cell division, filament-forming GTPase and a structural homologue of eukaryotic tubulin.
It interacts with membrane-associated proteins FtsA and ZipA
and assembles into a ring like structure at the midcell, this ring is known as Z-ring.
The formation of the Z-ring is facilitated by the ability of FtsZ to bind to GTP, which enables polymerization of FtsZ, resulting in the creation of straight protofilaments.
It is the first protein to move to the division site, and is essential for recruiting other proteins that produce a new cell wall between the dividing cells. So it is an emergent target for new antibiotics.

Cell envelope
DNA
FtsZ ring

10
Mechanism of action of FtsZ

11
Benzimidazoles: A new class of anti-TB drugs
Benzimidazole nucleus is a constituent of bioactive heterocyclic compounds and structural isosters of naturally occurring nucleotides, which allows them to interact easily with the biopolymers of the living system.
The 2,5,6-trisubstituted Benzimidazoles series of compound taken for study having biological activity ranging from 0.06 to 100 µg/ml, from the literature.
Benzimidazole structure

To build 2D-QSAR model to derive important physicochemical properties related to FtsZ inhibitors.
To build Gaussian based 3D-QSAR model for FtsZ inhibitors.
Development of best 3D-Pharmacophore model using Hip-hop based method.
Analysis of important interaction involve in FtsZ protein using docking-based approach.
Analysis of Hits obtained through virtual screening using pharmacophore-based Approach and docking-based Approach.
Aim and objective

• Ligand based drug design Structure based drug design
Methodology
Docking2D-QSAR Pharmacophore
Virtual screening
Novel drug molecule
3D-QSAR

1.QSAR A quantitative structure-activity relationship (QSAR) correlates molecular properties to some specific biological activity in terms of an equation.
Collection of compound from literature
Descriptor Generation
Feature Selection
Model Construct
Model Validation
Model Development flow chart
Ligand based approach

Collection Of Dataset: Total 59 compounds were collected from the literature, and drawn using Marvin Sketch program.Biological Assay:FtsZ polymerization inhibitory assayBiological activity( MIC) ranging from the 0.06 – 100 µg/ml.MIC is converted to pMIC values.Imported to the QSAR Module of Discovery studio Software. Series of compounds
1. The 2,5,6-trisubstituted benzimidazoles.
, october,2011 Journal of Medicinal Chemistry , September 2011
2D-QSAR
Reference: Kumar, K.; Awasthi, D.; Lee, S.-Y.; Zanardi, I.; Ruzsicska, B.;Knudson, S.; Tonge, P. J.; Slayden, R. A.; Ojima, I. Novel trisubstituted benzimidazoles, targeting Mtb FtsZ, as a new class of antitubercular agents. J. Med. Chem. 2011, 54, 374−381.

16
Comp No. R1R2N R3 MIC((µg/ml) pMIC
1 0.63 5.78
2 6.25 4.82
3 100 3.63
4 50 3.96
5 0.06 6.77
Benzimidazole series compounds which are used in QSAR study

17
MLR PLS
Methods
Training set Test set Training set Test set
r2
1. Random method
0.936 0.732 0.870 0.696
2. Diversed method
0.933 0.608 0.849 0.607
Division of Training and Test set
Best 2D-QSAR Model was generated through Random based and MLR method.
Number of molecules in Training set = 48Number of molecules in Test set = 11

Log (1/C) = 0.415 + 0.407 * (ALogP)- 3.268e-003 * ( Molecular_Weight ) + 0.1909 * (Num_H_Donors ) + 0.3852 * (Num_H_Acceptors) - 0.1666 * (Num_RotatableBonds) - 0.5808 * (Num_Rings) - 1.433e-002 * ( Num_AromaticRings) + 10.36 * (Molecular_Fractional Polar Surface Area)
2D-QSAR Model

Graph of 2D-QSAR (Random based -MLR)
19
Training set-48 (r2 = 0.936) Test set-11 (r2 = 0.732 )

20
The value of AlogP = 1.6-5.6
Number of HBA = 2-5
Number of HBD = 2-4
Polar surface area = 0.141-0.292
Number of rotatable bonds = 4-12
Number of Rings = 1-3
Conclusion of 2D-QSAR
Benzimidazole scaffolds

3D-QSAR (Gaussian Based Method)
Alignment of molecule Docking based
Pharmacophore
basedAtom based method
3D-QSAR exploits the three-dimensional properties of the ligands to predict their biological activities using chemometric techniques. It has served as a valuable predictive tool in the design of pharmaceuticals.

22Training set-45 (r2 = 0.844 ) Test set -11 (r2 = 0.682)
Compound (Random based )
r2
Training set (48 comp.) 0.844Test set (11 comp.) 0.682
Total compounds:59
3D-QSAR GRAPH( Field-based method)
Scatter Plot Analysis

Training set-48 (r2 = 0.839) Test set -11 (r2 = 0.667)
Scatter Plot Analysis
3D-QSAR GRAPH(Gaussian-based method
Compound (Random based )
r2
Training set (48 comp.) 0.839Test set (11 comp.) 0.667
Total compounds:59

24
COUNTER MAP OF 3D-QSAR (CoMFA)
Field-Based QSAR-Steric
Field-Based QSAR-Electrostatic
NH1
N
Electropositive
Electronegative
POSITIVER3

25
COUNTER MAP OF 3D-QSAR (CoMSIA)
Gaussian Based QSAR-Steric
POSITIVER3

26
NR3 POSITIVE NEGATIVE
NH1 N Electropositive Electronegative
Gaussian Based QSAR-Hydrophobic
Gaussian Based QSAR-Electrostatic

27
Gaussian Based QSAR-HBD
N
R3
POSITIVE
NEGATIVE
O
R3
NEGATIVE
POSITIVE
Gaussian Based QSAR-HBA

28
HBA
HBD
Steric
Hydrophobic
Electropositive
Electronegative Summary of the 3D-QSAR

2.pharmacophoreA pharmacophore is the ensemble of steric and electronic features that is necessary to ensure the optimal supramolecular interactions with a specific biological target structure and to trigger its biological response.
Model Development flow chart
Input-2D/3D molecules Structure
CHARMm forcefield and Minimization
Diverse Conformation generation
Generation of Hypothesis
validation

Pharmacophore resultFeatures Rank Direct Hit Partial Hit Max Fit
01 YZHH 116.621 1111111111 0000000000 4
02 YZHH 116.121 1111111111 0000000000 4
03 YZDH 115.918 1111111111 0000000000 4
04 RYZH 115.363 1111111111 0000000000 4
05 RYZH 115.363 1111111111 0000000000 4
06 RYZH 115.159 1111111111 0000000000 4
07 RYZH 115.159 1111111111 0000000000 4
08 YZDH 114.740 1111111111 0000000000 4
09 YZHA 114.621 1111111111 0000000000 4
10 YZHA 114.621 1111111111 0000000000 4
Hy-ali
Hy
HBD: Hydrogen bond donar (D)HBA: Hydrogen bond acceptor (H)
HY: Hydrobhobic (Z)Hy-ali: Hydrobhobic aliphatic (Y)
HBA
HBD

31
A. (Most active Compound-5) B. (Least active compound-3)
Alignment of the most potent compound-5 & least active compound-3.

Docking
32
PDB ID= 1RLU Resolution=2.08 R-value=0.182 pH=5.6
LIGAND-C10 H16 N5 O13 P3 S 5'-GUANOSINE-DIPHOSPHATE- MONOTHIOPHOSPHATE
Structure based approach
Preparation of protein
Preparation of ligand

Structure of FtsZ protein
Nucleotide binding site
GTP γ Thiophosphate

34
Active site residues PDB:1RLU, Ligand-Protein Complex
Molecular structure view

35
Ligand Interaction Diagram

36
RMSD VALUE=0.726
DOCKING SCORE= -8.93
Docking of the substrate on the same active site
Co-crystalised ligand Docked ligand

37
Compound NameBiological Activity
(pMIC) Docking Score
Compound-5 6.77 -8.11Compound-39 6.38 -7.32
Compound-43 6.1 -7.07
Compound-14 5.78 -6.27
Compound-30 5.36 -6.61
Compound-37 5.42 -6.81
Compound-7 5.09 -6.62
Compound-38 4.86 -6.33
Compound-48 4.76 -6.27
Compound-18 4.59 -6.22
Compound-21 4.48 -5.50
Compound-3 3.56 -4.98
The important amino acid residue are Glu136, Arg140, Phe 180, Asp184.
Docking of the selected FtsZ Inhibitors

38
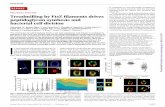
Interaction of Most active compound-5 & Least active compound-3
Compound-5
Compound-3
Compound-3
Compound-5

39
Virtual screening workflow Database ( Asinex database with 100000 molecules)
Filter (Lipinski rule of 5)
Shape based screening (ROCS)
Retrieval of protein information from PDB (PDB ID 1RLU)
Protein preparation
Pharmacophore-based virtual screening
5 Hit Compounds
Receptor grid generation 500 Compounds
78000 Compounds
Docking studies
Novel Hit Molecules

40
Compoundnumber
Compound name Fit value
Hit 1 1-(3-Benzyl-2-butyl-5-methyl-3H-imidazo[4,5-b]pyridin-6-yl)-3-(3-chloro-phenyl)
3.65885
Hit 2 Pentanoic acid {2,5-dimethoxy-4-[3-(tetrahydro-furan-2-ylmethyl)-thioureido]-ph
3.54616
Hit 3 N-[2-(5-Methyl-furan-2-yl)-1H-benzoimidazol-5-yl]-butyramide_35
3.54553
Hit 4 Cyclopropanecarboxylic acid {2,5-diethoxy-4-[3-(tetrahydro-furan-2-ylmethyl)-th
3.53696
Hit 5 1-[2-Butyl-3-(4-chloro-benzyl)-5-methyl-3H-imidazo[4,5-b]pyridin-6-yl]-3-(3-chl
3.51725
Virtual screening Result Analysis
Table : Compound name and fit value of Hits molecules

41
Hit 3Hit 1 Hit 2
Hit 4 Hit 5
Hit molecules from Virtual screening

42
Conclusion
The study of 2D-QSAR of FtsZ inhibitors concluded that the descriptor ALogP, HBA, HBD, PSA are important for antitubercular activity.
3D-QSAR models were interpreted in the form of contour maps, concluded that steric field contribution is higher as compared to other field intensities. The contour map of steric, hydrophobic, electro-negative, electro-positive, HBA at R3 position and electro-positive, HBD at R1 and R2 position responsible for increased in antitubercular activity.
The best hypothesis with four point pharmacophore concluded that the best model generated is the 3rd number hypothesis i.e. compound-5 (YZDH).

43
The docking study concluded that Glu136, Arg140, Phe180 and Asp184 are important residues for ligand binding. Asp184 also provides essential H-bond interactions for favourable ligand binding. And Asp184 should be taken in drug design targeting the FtsZ.
Virtual screening method from Asinex database resulted in identification of five new Hits, as potential Hits for testing of novel inhibitors, these hits may responsible for the antitubercular activity.
So finally we concluded that FtsZ is an emergent target for the development of new antitubercular drugs.

REFERENCES
2. Leung AKW, White EL, Ross LJ, Reynolds RC, DeVito JA, Borhani DW. Structure of Mycobacterium tuberculosis FtsZ reveals unexpected, G protein-like conformational switches. J Mol Biol 2009, 342(3), 953–970.
3. Margalit DN, Romberg L, Mets RB, et al. Targeting cell division: small-molecule inhibitors of FtsZ GTPase perturb cytokinetic ring assembly and induce bacterial lethality. [Erratum to document citedin CA141:271048]. Proc Natl Acad Sci USA 2004, 101(38), 13969.
4. Scheffers D-J, de Wit JG, den Blaauwen T, Driessen AJM. GTP hydrolysis of cell division protein FtsZ: evidence that the active site is formed by the association of monomers. Biochemistry 2002, 41(2), 521–529.
1. K. Kumar, et al. Discovery of anti-TB agents that target the cell-division protein FtsZ, Future Med Chem, 2010, 2 (8), 1305-1323.

45