Phosphate Uptake in the Cyanobacterium Synechococcus R-2 PCC
ELECTRON TRANSPORT PROTEINS OF SYNECHOCOCCUS SP. …
Transcript of ELECTRON TRANSPORT PROTEINS OF SYNECHOCOCCUS SP. …
The Pennsylvania State University
The Graduate School
Department of Biochemistry, Microbiology, and Molecular Biology
ELECTRON TRANSPORT PROTEINS OF
SYNECHOCOCCUS SP. PCC 7002
A Thesis in
Biochemistry, Microbiology, and Molecular Biology
by
Christopher T. Nomura
Submitted in Partial Fulfillment
of the Requirements
for the Degree of
Doctor of Philosophy
May 2001
Date of SignatureWe approve the thesis of Christopher T. Nomura
Donald A. BryantErnest C. Pollard Professor of
BiotechnologyProfessor of Biochemistry and Molecular
BiologyChair of Committee
John H. GolbeckProfessor of Biochemistry and Biophysics
Ola SodeindeAssistant Professor of Biochemistry and
Molecular Biology
Paul BabitzkeAssociate Professor of Biochemistry and
Molecular Biology
Juliette T. J. LecomteAssociate Professor of Chemistry
Robert A. SchlegelProfessor of Biochemistry and Molecular
BiologyHead of the Department of Biochemistry
and Molecular Biology
iii
ABSTRACT
Cyanobacteria are photosynthetic, oxygen-evolving prokaryotes that have adapted
to a wide range of ecological niches. In particular, cyanobacteria represent interesting
organisms to study electron transport in because they have both photosynthetic and
respiratory proteins involved in electron transport on the same membrane: the thylakoid
membrane. Of particular interest to our lab is the identification and characterization of the
minimal conserved number of genes responsible for coding proteins used in electron
transport in cyanobacteria. In order to address this issue, a Synechococcus sp. PCC 7002
cosmid library was screened with heterologous probes made from the completely
sequenced genome of the freshwater cyanobacterium, Synechocystis sp. PCC 6803.
These heterologous probes were also used to screen Synechococcus sp. PCC 7002 partial
genomic libraries in the cases where positive hybridizations could not be identified within
the cosmid libraries. In this study, 35 open reading frames in the marine cyanobacterium
Synechococcus sp. PCC 7002 have been identified and sequenced which, either, encode
electron transport proteins or encode accessory proteins necessary for the assembly of
these electron transport proteins based on BLAST algorithm searches. These open
reading frames are represented by the following genes: ndhA, ndhI, ndhG, ndhE, ndhB,
ndhC, ndhK, ndhJ, ndhD1, ndhD2, ndhD3, ndhD4, ndhF1, ndhF2, ndhF3, ndhF4, ndhF5,
iv
ndhH, ndhL, ndbA, ndbB, hypE, hoxE, hoxF, hoxU, hoxY, hypD, hoxH, hoxW, hypA,
hypB, hypF, hypC, hypD, petJ1, petJ2, bcpA, ctaCI, ctaDI, ctaEI, ctaCII, ctaDII, and
ctaEII. These genes putatively encode subunits for the type I NADH dehydrogenase,
two type II NADH dehydrogenases, the subunits and accessory proteins for a bi-
directional hydrogenase, and three putative mobile electron carriers: cytochrome c6,
cytochrome c62, and BcpA, and the subunits for two different members of the heme-
copper cytochrome oxidase family. Most of these genes bear the highest homology to
their respective counterparts in the freshwater cyanobacterial strain Synechocystis sp.
PCC 6803 and a comparison between these minimal conserved sets of genes may
represent the smallest number of genes necessary for these organisms to carry out
electron transport.
Attempts were made to inactivate several of the electron transfer protein genes
identified in this study. These include the ndhB gene, which, encodes a subunit of the
type I NADH dehydrogenase, the petJ1 gene that encodes cytochrome c6, the petJ2 gene
that encodes a putative second c-type cytochrome, the bcpA gene which encodes a
putative blue-copper protein, the hoxH gene which encodes the large Ni containing
subunit of the hydrogenase enzyme, the hoxF gene which encodes the FMN containing
subunit of the hydrogenase enzyme, the ctaDI gene, which encodes the large subunit of
the type I cyanobacterial cytochrome oxidase, and the ctaDII gene, which encodes the
large subunit of a secondary heme-copper oxidase in Synechococcus sp. PCC 7002. This
study found that the ndhB and petJ1genes are essential for the viability of Synechococcus
sp. PCC 7002 and therefore could not be inactivated. The hoxH, hoxF, petJ2, bcpA,
v
ctaDI, and ctaDII genes are all non-essential to Synechococcus sp. PCC 702 under
normal growth conditions and could be inactivated.
Physiological studies of the hoxH, hoxF, petJ2, and bcpA inactivated strains of
Synechococcus sp. PCC 7002 revealed that there were no significant phenotypes under
the conditions tested. However, evidence has been found in this study that the heme-
copper oxidase enzymes encoded by the ctaCIDIEI and ctaCIIDIIEII gene clusters have
significant roles in respiration, high-light tolerance and oxidative stress responses in
Synechococcus sp. PCC 7002.
Table of Contents
List of Figures xiii
List of Tables xviii
Acknowledgments xx
Chapter 1 INTRODUCTION 1
1.1 Electron Transport Proteins 1
1.1.1 Type I NADH dehydrogenase 4
1.1.2 Type II NADH dehydrogenase 7
1.1.3 Hydrogenase 9
1.1.4 Mobile electron carriers 12
1.1.4.1 Cytochrome c6 13
1.1.4.2 Plastocyanin 16
1.1.5 Terminal oxidases 17
1.2 Purpose of the present work 22
Chapter 2 MATERIALS AND METHODS 24
2.1 Bacterial strains and growth conditions 24
2.1.1 Synechococcus sp. PCC 7002 24
2.1.2 Synechocystis sp. PCC 6803 26
2.1.3 Escherichia coli 27
2.2 Standard laboratory methods 28
2.3 Isolation and manipulation of DNA 29
vii
2.3.1 Plasmid and cosmid isolation from Escherichia coli 29
2.3.2 Total DNA isolation from Synechococus sp. PCC 7002 30
and Synechocystis sp. PCC 6803
2.4 Transformation Procedures 31
2.4.1 E. coli transformation procedures 31
2.4.2 Cyanobacterial transformation procedures 32
2.5 Southern blot transfer and hybridization 34
2.6 Cloning and sequencing 35
2.6.1 General cloning strategy 35
2.6.2 Construction of a Synechococcus sp. PCC 7002 genomic library 36
2.6.3 Probes 36
2.6.4 Colony hybridization 38
2.6.5 DNA sequencing and analysis 39
2.7 RNA isolation, Northern-blot hybridization, and RT-PCR analysis 40
2.7.1 RNA isolation 40
2.7.2 Northern-blot hybridization analyses 41
2.7.3 Reverse transcriptase-polymerase chain reaction (RT-PCR) 43
2.8 Overproduction of the rBcpA protein in Escherichia coli 44
2.8.1 Generation of E. coli expression plasmids 44
2.8.2 Protein overproduction in E. coli and isolation of inclusion bodies 45
2.8.3 Purification of rBcpA from E. coli 46
2.9 Cytochrome c6 purification from Synechococcus sp. PCC 7002 47
and amino terminal sequencing
2.10 SDS-polyacrylamide gel electrophoreses, TMBZ staining and 48
immunoblot analysis
viii
2.10.1 SDS PAGE 48
2.10.2 3, 3', 5, 5'-tetramethylbenzidine (TMBZ) staining 49
2.10.3 Immunoblot analysis 50
2.11 Determination of chlorophyll a and carotenoid contents 51
2.12 Oxygen evolution and consumption rates 52
2.13 P-700+ kinetic measurements 52
2.14 77K fluorescence emission measurements 54
2.15 Pulse Amplitude Modulated (PAM) fluorescence measurements 54
2.16 Superoxide dismutase (SOD) activity measurements 55
2.17 Detection of hydroperoxides 56
2.18 Catalase activity 57
2.19 Peroxidase activity 57
2.20 Cell viability 58
Chapter 3 RESULTS 60
Cloning of electron transport protein genes 60
3.1. Type I NADH dehydrogenase 66
3.1.1 ndhAIGE 67
3.1.2 ndhB 70
3.1.2.1 Attempted mutagenesis of the Synechococcus sp 73
PCC 7002 ndhB gene by interposon mutagenesis
3.1.3 ndhD1, ndhD2, ndhD3, ndhD4 78
3.1.4 ndhF1, ndhF2, ndhF3, ndhF4, ndhF5 83
3.1.5 Gene organization and phlyogenetic analysis of the ndhD 89
and ndhF genes in cyanobacteria
ix
3.1.6 ndhCKJ 95
3.1.7 ndhH 97
3.1.8 ndhL 100
3.2 Type II NADH dehydrogenases 103
3.3 Bi-directional hydrogenase 107
3.3.1 hypE 109
3.3.2 hoxE 109
3.3.3 hoxF 109
3.3.4 hoxU 110
3.3.5 hoxY 111
3.3.6 hyp3 111
3.3.7 hoxH 111
3.3.8 hoxW 112
3.3.9 hypA 112
3.3.10 hypB 113
3.3.11 hypF 113
3.3.12 hypC 116
3.3.13 hypD 116
3.3.14 Interposon mutagenesis of the hoxH and hoxF genes in 117
Synechococcus sp. PCC 7002
3.3.15 Chlorophyll and carotenoid contents of the hoxH- and hoxH-/hoxF- 123
strains
3.3.16 Growth analysis of hoxH- and hoxH-/hoxF- double mutants 124
3.4 Mobile electron carriers 127
x
3.4.1 Cloning and sequencing of the Synechococcus sp. PCC 7002 127
cytochrome c6 gene
3.4.2 Attempted deletion of the petJ1 gene by interposon mutagenesis 138
3.4.3 Attempted functional substitution of the Synechococcus sp. 142
strain PCC 7002 petJ gene with the petE and petJ genes from
Synechocystis sp. strain PCC 6803
3.4.4 petJ2 157
3.4.5 Interposon mutagenesis of the petJ2 gene in Synechococcus sp. 159
PCC 7002
3.4.6 Growth analysis of petJ2::aphII 162
3.4.7 cytM 165
3.4.8 bcpA 169
3.4.9 Interposon mutagenesis of the bcpA gene from Synechococcus sp. 172
PCC 7002
3.4.10 Growth rate analysis of bcpA- strains of Synechococcus sp. 176
PCC 7002
3.4.11 Overproduction of the BcpA protein in E. coli 176
3.4.12 Immunoblot analysis of rBcpA 180
3.5 Heme-Copper Oxidases in Synechococcus sp. PCC 7002 186
3.5.1 Screening and Cloning of the ctaI and ctaII gene clusters from 186
Synechococcus sp. PCC 7002
3.5.2 Insertional Mutagenesis of ctaDI and ctaDII from 190
Synechococcus sp. PCC 7002
xi
3.5.3 Expression of the cta gene clusters in Synechococcus sp. PCC 193
7002
3.5.4 Respiratory activity and oxygen evolution activity in 203
Synechococcus sp. PCC 7002 wild type, ctaDI-, and
ctaDII- strains
3.5.5 Growth analysis of Synechococcus sp. PCC 7002 wild type, ctaDI-, 206
ctaDII-, and ctaDI- ctaDII- strains
3.5.6 Chlorophyll and carotenoid contents of wild type, ctaDI-, ctaDII- 211
ctaDI- ctaDII- strains
3.5.7 Photoinhibition of whole chain electron transport activity in 213
Synechococcus sp. PCC 7002 wild type and ctaDII-, and
ctaDI- ctaDII- strains
3.5.8 Fluorescence emission at 77K of wild-type, ctaDI-, ctaDII- and 215
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
3.5.9 P700+ Reduction Kinetics 219
3.5.10 Pulse Amplitude Modulated Fluorescence Measurements 222
3.5.11 Oxidative Stress and Cell Viability of Synechococcus sp. 223
PCC 7002 wild type and ctaD mutant strains
3.5.12 Superoxide Dismutase (SOD) Activity Assays 229
3.5.13 Hydroperoxide levels in Synechococcus sp. PCC 7002 231
3.5.14 Catalase and peroxidase activity in Synechococcus sp. PCC 7002 233
Chapter 4 DISCUSSIONS AND CONCLUSIONS 235
4.1 Genes and gene organization in cyanobacteria 235
4.1.1 Future directions of the gene sequencing project in 237
xii
Synechococcus sp. PCC 7002
4.2 Type I NADH dehydrogenase 238
4.2.1 Future directions for research of the type I NADH dehydrogenease 241
genes in Synechococcus sp. PCC 7002
4.3 Type II NADH dehydrogenases 242
4.3.1 Future directions for research of the type II NADH dehydrogenease 243
genes in Synechococcus sp. PCC 7002
4.4 Bi-directional hydrogenase 244
4.4.1 Future directions for research of bidirectional hydrogenase and 245
hyp genes in Synechococcus sp. PCC 7002
4.5 Mobile electron carriers 247
4.5.1 Future directions for research of the mobile electron carrier genes 252
in Synechococcus sp. PCC 7002
4.6 Terminal Oxidases Present in Synechococcus sp. PCC 7002 253
4.6.1 Effects of Cytochrome Oxidase on Electron Flow Around PSII 257
4.6.2 Cytochrome Oxidase and its Role in High Light Stress 258
4.6.3 Tolerance of Methyl Viologen and High Light Stress in ctaDII- 259
strains of Synechococcus sp. PCC 7002
4.6.4 Future research directions for cta- mutants in Synechococcus 261
sp. PCC 7002
4.7 Concluding Remarks 267
References 270
Appendix A: Restriction maps of electron transport genes 296
Appendix B: ClustalW alignments of electron transport proteins 316
xiii
List of Figures
Figure 1 Depiction of the cyanobacterial thylakoid membrane 2
Figure 2 Schematic illustration of the similarities and difference of 20
four subclasses of the heme-copper oxidase superfamily
Figure 3 Gene organization of ndhAIGE gene cluster in cyanobacteria 68
Figure 4 Gene organization around the ndhB gene in cyanobacteria 72
Figure 5 The ndhB gene of Synechococcus sp. PCC 7002 is 77
required for cell viability
Figure 6 ClustalW alignment of NdhD amino acid sequences from 81
Synechococcus sp. PCC 7002
Figure 7 ClustalW alignment of NdhF amino acid sequences from 86
Synechococcus sp. PCC 7002
Figure 8 Gene organization of ndhF and ndhD genes 91
Figure 9 Phylogenetic analysis of NdhF and NdhD proteins 94
Figure 10 Gene organization of ndhCJK gene clusters 96
Figure 11 Gene organization around the ndhH gene in cyanobacateria 99
Figure 12 Gene organization around the ndhL gene in cyanobacteria 102
Figure 13 Gene organization around the ndbA gene in cyanobacteria 104
xiv
Figure 14 Gene organization around the ndbB gene in cyanobacteria 106
Figure 15 Hydrogenase gene cluster organization 108
Figure 16 HypF and acylphosphatase alignments 115
Figure 17 Interposon mutagenesis of hoxH of Synechococcus sp. PCC 7002 119
Figure 18 Interposon mutagenesis of hoxF of Synechococcus sp. PCC 7002 122
Figure 19 Temperature-shift effects on wild-type and hoxH::aphII 126
Figure 20 SDS PAGE analysis of cytochrome c6 from Synechococcus sp. 130
PCC 7002
Figure 21 Probe design and Southern blot hybridization of petJ from 132
Synechococcus sp. strain PCC 7002
Figure 22 Gene organization around petJ1 in cyanobacteria 134
Figure 23 ClustalW alignment of PetJ1 amino acid sequences 137
Figure 24 Attempted interposon mutagenesis of the petJ1 gene 140
Figure 25 Northern blot analysis of the petJ1 gene 141
Figure 26 Insertion of the Synechocystis sp. PCC 6803 petEgene into 145
the ∆petJ::aphII merodiploid strain of Synechococcus sp
PCC7002
Figure 27 Attempted mutagenesis of the petJ1 gene in the ∆petJ::aphII 147
merodiploid strain Synechococcus sp.PCC 7002/Synechocystis sp.
PCC 6803 petE platform vector strain.
xv
Figure 28 Insertion of the Synechocystis sp. PCC 6803 petJgene into 149
the petJ::aphII merodiploid strain of Synechococcus sp.
PCC 7002.
Figure 29 The petJ gene from Synechocystis sp. PCC 6803 cannot 151
functionally replace the petJ gene from Synechococcus sp.
PCC 7002.
Figure 30 RNA blot analysis of the Synechocystis sp. PCC 6803 153
petE gene in the petJ::aphII merodiploid strain of
Synechococcus sp. PCC 7002
Figure 31 RNA blot analysis of the Synechocystis sp. PCC 6803 155
petJ gene in the ∆petJ::aphII merodiploid strain of
Synechococcus sp. PCC 7002
Figure 32 Western blot analysis of the Synechocystis sp. PCC 6803 petE 156
protein produced in the petJ::aphII merodiploid strain of
Synechococcus sp. PCC7002
Figure 33 ClustalW alignment of PetJ2 with PetJ amino acid sequences 158
Figure 34 Interposon mutagenesis of the petJ2 gene 161
Figure 35 Growth analysis of the petJ2::aphII strain of Synechococcus sp. 164
PCC 7002
Figure 36 Gene organization around the cytM gene in cyanobacteria 167
Figure 37 ClustalW alignment of CytM amino acid sequences 168
Figure 38 A. BcpA ClustalW alignment 171
B. BcpA/PetE ClustalW alignment
xvi
Figure 39 Transcript analysis of petJ1, petJ2, and bcpA 173
Figure 40 Interposon mutagenesis of the Synechococcus sp. 175
PCC 7002 bcpA gene
Figure 41 Growth analysis of the bcpA::aphII strain of Synechococcus sp. 178
PCC 7002
Figure 42 Overproduction of the rBcpA protein 183
Figure 43 Immunoblot analysis of rBcpA 185
Figure 44 Gene organization of the ctaCIDIEI genes from cyanobacteria 188
Figure 45 Gene organization of the ctaCIIDIIEII genes from cyanobacteria 189
Figure 46 Interposon mutagenesis of the ctaDI gene of Synechococcus sp. 192
PCC 7002
Figure 47 Interposon mutagenesis of the ctaDII gene of Synechococcus sp. 195
PCC 7002
Figure 48 Northern blot analysis of the ctaCIDIEI gene cluster of 198
Synechococcus sp. PCC 7002
Figure 49 RT-PCR analysis of the ctaCIIDIIEII gene cluster of 202
Synechococcus sp. PCC 7002
Figure 50 Effect of extreme high light intensity on cell viability wild type 208
and ctaDI- strains of Synechococcus sp. PCC 7002
Figure 51 Growth analysis of analysis of the wild type, ctaDI-, ctaDII-, 210
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002 grown
xvii
under 250 µE m -2 s-1 and 4.5 mE m-2 s-1
Figure 52 Effects of high light intensity on whole chain electron transport 214
in wild type, ctaDI-, ctaDII-, and ctaDI- ctaDII- strains of
Synechococcus sp. PCC 7002
Figure 53 Fluorescence emission spectra at 77K of whole cells of Synechococcus 218
sp. PCC 7002 wild-type and cytochrome oxidase mutant strains-
Figure 54 Methyl viologen tolerance and sensitivity in Synechococcus 225
sp. PCC 7002
Figure 55 Growth analysis of analysis of the wild type, ctaDI-, ctaDII-, 227
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002 in the
presence of methyl viologen
Figure 56 Oxidase Mg2+ and CuA binding motif alignments 255
Figure 57 Model for toxicity in ctaD mutant strains grown under 265
4.5 mE m-2 s-1 constant illumination.
xviii
List of Tables
Table 1 Nomenclature and properties of type I NADH dehydrogenases 5
Table 2 Primers for Synechocystis sp. PCC 6803 electron transfer protein 37
genes
Table 3 Electron transport proteins from Synechococcus sp. PCC 7002 63
Table 4 Percent identity and similarity of Synechococcus sp. PCC 7002 65
electron transport proteins with homologs from other bacteria and
the chloroplast
Table 5 Percent identity and similarity of Synechoccus sp. PCC 7002 NdhD 83
proteins compared to one another
Table 6 Percent identity and similarity of Synechoccus sp. PCC 7002 NdhF 88
proteins compared to one another
Table 7 Protein refolding conditions for rBcpA 183
Table 8 Primers for Synechococcus sp. PCC 7002 ctaCIDIEI and 196
ctaCIIDIIEII genes
Table 9 Oxygen uptake activity of wild type, ctaDI-, ctaDII-, and 204
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Table 10 Doubling times, oxygen evolution rates and Fv’/Fm’ for wild type, 205
ctaDI-, ctaDII-, and ctaDI- ctaDII strains of Synechococcus sp. PCC
7002 grown under various light intensities
xix
Table 11 Chlorophyll a and carotenoid contents for wild type, ctaDI-, 212
ctaDII-, and ctaDI- ctaDII strains of Synechococcus sp. PCC 7002
grown under various light intensities.
Table 12 P700+ redox kinetics of wild type, ctaDI-, ctaDII-, and 220
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Table 13 Cell viability of wild type, ctaDI-, ctaDII-, and ctaDI- ctaDII- 228
strains of Synechococcus sp. PCC 7002
Table 14 Superoxide dismutase activities of wild type, ctaDI-, ctaDII-, 230
and ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Table 15 Hydroperoxide concentrations of wild type, ctaDI-, ctaDII-, 232
and ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Table 16 Catalase and peroxidase activities of wild type, ctaDI-, ctaDII-, 234
and ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
xx
ACKNOWLEDGMENTS
Thanks to everyone who helped make this reality-
Family: Mom, Dad, Mary, Grandma and Grandpa Ito, Grandma Miki and Grandpa
Henry, Uncle Eric and Auntie Hatsumi, Amy, Adam, Celeste, Joachim, The Ito Family,
The Nomura Family, The Nakadegawa Family.
Old School: (WEST COAST) Douglas Jímenez, Wayne Szeto, Kim Gasuad, Lupe
Garcia, Scottie Henderson, Matt Mills and Miné Berg, Bernice Frankl, Dr. Leo Ortiz,
Carlos Morel, John Lee Hooker, Chris Lopez, Leandra, MBRS, MARC, Kwame
Asamoah, “Chocalate” Terry G, Kiersten and Tony. (EAST COAST) Noah, Jason, Elan,
Ranon, Alex, Jess.
New School: Søren and Suzanne, NUF and Yumiko, Gaozhong Shen, Lena and Ilya,
Juergen, Katja, Toshio, Kaori, Tanja Gruber, Vicki Stirewalt, Wendy Schlucter, Soohee
Chung, Zhao, Frank and Jessica, Joel, Fetchko, Denise, Pete, Nick, Celina, Matt, Laurie,
Crazy Bob, all of the undegrads who do all the work in our lab Melissa, Anne, Ginny,
Matt, Kirstin, John, Mel and everyone else I don’t have room for.
The Committee: Dr. John Golbeck, Dr. Ola Sodeinde, Dr. Paul Babitzke, Dr. Juliette
Lecomte, and of course my advisor, Dr. Don Bryant.
Many thanks for all your support, patience, and encouragement. Sorry if I left
you out (thanks to you too!!)
Chapter 1
INTRODUCTION
1.1 Electron transport proteins in cyanobacteria
Cyanobacteria are photosynthetic, oxygen-evolving prokaryotes that have adapted
to a wide range of ecological niches (Stanier and Cohen-Bazire, 1977). Cyanobacteria
represent interesting organisms in which to study electron transport because they have
both photosynthetic and respiratory proteins involved in electron transport on a single
membrane: the thylakoid membrane (Figure 1). Photosynthetic electron transport in
cyanobacteria is similar to that of higher plants (Scherer, 1990). They have a
plastoquinone pool that is reduced by PSII, a cytochrome b6f complex which acts as a
plastoquinol-plastocyanin/cytochrome c6 oxidoreductase and is homologous in function
to the cytochrome bc complex of mitochondria, and a soluble cytochrome c6 that reduces
PSI (Scherer, 1990). Unlike plants, in which the light harvesting apparatus is made up of
integral membrane protein complexes that contain chlorophyll a and chlorophyll b that
direct energy to the photosystems, cyanobacteria use light-harvesting complexes called
phycobilisomes (Bryant et al., 1990).
2
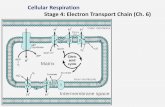
Figure 1. Depiction of the thylakoid membrane of cyanobacteria adapted from D.A.
Bryant (1994). Photosynthetic and respiratory complexes are identified by the text in the
figure. Light energy is represented by hν. The arrows with solid lines represent electron
flow and arrows with dashed lines represent proton movement.
STROMA
FeS
SIV
Cyt fLP
HP
NADP+ + H+
NADPH
h
Cyt 553
Cyt b6
H+
Photosystem ICytochrome b6/f complex
Photosystem II Photosystem II ATPsynthase
Thylakoid membrane
LUMEN
r
o
F
FdA B
Ch
h h
h
QBQA
PheoD1
D2
NADH
P680
Z
933H2O
2H++1/2O2
H+
H+
Phycobilisome
PQ
PQH2
NAD+ + H+
NDH c
a
cc c CF0
b' b
ADP + Pi ATP + H2O
CF1
H+e-
NADH dehydrogenase
Cyt ox
2H++1/2O2
H+
Cytochromeoxidase
P700
A0
A1
FX
H2O
H+
FNR A
D
FE F
h h933
Mn4
D2
D1
Cyt b559 CP43
CP47
PsaA PsaBP
saE
F
PsaD
or
Flvd
Mn4
ABCP47
CP43
FNR
Cyt b559
Phycobilisomes are composed of water-soluble phycobiliproteins that harvest light and
transfer light energy to the photosynthetic reaction centers (Bryant et al., 1990). Light
energy is captured by phycobilisomes and chlorophyll and ultimately transferred to a
special pair of chlorophyll a molecules (P680) from which the electron is passed through
a series of electron acceptors within PSII to plastoquinone QB. The oxidation of water by
the Mn center of the PSII complex provides electrons to reduce the oxidized P680
reaction center. Electrons from reduced plastoquinone are passed through the
3
cytochrome b6f complex, which then transfers electrons to an oxidized mobile electron
carrier, either plastocyanin or cytochrome c6, depending on the species of cyanobacteria
or environmental conditions (Scherer, 1990; Zhang et al., 1992). The mobile electron
carrier then transfers its electron to reduce the oxidized special-pair (P700) chlorophyll a
molecules of PSI. Electrons are transferred in the PSI reaction center through a series of
cofactors to soluble ferredoxin. Ferredoxin transfers its electrons to ferredoxin:NADP+
oxidoreductase, which in turn forms NADPH by reduction of NADP+. The net result of
these reactions is reducing power to fix carbon from carbon dioxide, the production of
oxygen from the oxidation of water, and transfer of protons into the lumenal space of the
thylakoid. This proton transport is facilitated by the cytochrome b6f complex, type I
NADH dehydrogenase, and cytochrome oxidase and generates an electrochemical proton
gradient and transfer of the protons back into the cytoplasmic space through ATP
synthase results in ATP synthesis (Schmetterer, 1994). Although the role of these
photosynthetic proteins has been defined by many studies, the roles of many of the other
potential electron transfer proteins within cyanobacteria remains open for research.
The recent ability to sequence and identify open reading frames from entire
genomes has opened up new avenues to perform comparative analyses between many
different organisms (Blattner et al., 1997; Deckert et al., 1998; Kaneko et al., 1996;
Nelson et al., 1999). The genome of the freshwater, unicellular cyanobacterium
Synechocystis sp. PCC 6803 has been completely sequenced and open reading frames
corresponding to putative electron transport proteins have been identified (Kaneko et al.,
1996). Our lab is interested in determining the similarities and differences in the number
4
and types of genes that may be involved in electron transport in Synechococcus sp. PCC
7002 and other cyanobacterial species. Of particular interest to our lab is the
identification and characterization of the minimal conserved number of genes responsible
for coding proteins used in electron transport in cyanobacteria.
1.1.1 Type I NADH Dehydrogenase
The NADH-ubiquinone oxidoreductase (complex I) of mitochondria is an enzyme
that consists of more than 40 different subunits (Anderson et al., 1982; Arizmendi et al.,
1992a; Arizmendi et al., 1992b; Walker, 1992a; Walker et al., 1992). The genes for most
of the subunits are located in the nucleus and the gene products must be imported into the
mitochondria for assembly on the inner membrane (Walker, 1992a; Walker et al., 1992;
Weiss et al., 1991). In vertebrates, 7 of these subunits are encoded by the mitochondria.
These subunits are ND1, ND2, ND3, ND4, ND4L, ND5 and ND6. Complex I is the first
major enzyme in the respiratory electron transport chain of mitochondria and is required
to generate the proton motive force used for ATP synthesis by the translocation of
protons (Weiss et al., 1991).
Homologues of the mitochondrial-encoded complex I genes as well as some of
the complex I subunits encoded by the nucleus are also found in plastid DNA suggesting
that a NADH dehydrogenase may be located within the chloroplast (Hiratsuka et al.,
1989). The genes for the NADH dehydrogenase of higher plant chloroplasts that are
homologous to the mitochondrial genes are named ndhA, ndhB, ndhC, ndhD, ndhE,
5
ndhF, ndhG, ndhH, ndhI, ndhJ, and ndhK (Table 1) (Friedrich et al., 1995; Friedrich and
Weiss, 1997; Schmitz-Linneweber et al., 2000).
Table 1. Nomenclature and properties of homologous of type I NADH dehydrogenase
subunits from different organisms adapted from Friedrich and Weiss (1997). The NADH
dehydrogenase subunits were identified for E. coli (Weidner et al., 1993), Bos taurus
(Walker, 1992b; Walker et al., 1992), Oryza sativa (Shimada and Sugiura, 1991), and
Synechocystis sp. PCC 6803 (Kaneko et al., 1996).
E. coli B. taurus O. sativa 6803 cofactor(s)/predicted functionNuoA ND3 NdhC NdhCNuoB PSST NdhK NdhK 1X[4Fe-4S]NuoC 30(IP) NdhJ NdhJNuoD 49(IP) NdhH NdhHNuoE 24(FP) not found not found 1X[2Fe-2S]NuoF 51(FP) not found not found NADH-binding; FMN; 1X[4Fe-4S]NuoG 75(IP) not found not found 1X[4Fe-4S]; 1 (2*)X[2Fe-2S]NuoH ND1 NdhA NdhA Ubiquinone-bindingNuoI TYKY NdhI NdhI 2X[4Fe-4S]NuoJ ND6 NdhG NdhGNuoK ND4L NdhE NdhENuoL ND5 NdhF NdhFNuoM ND4 NdhD NdhDNuoN ND2 NdhB NdhB6803 is Synechocystis sp. PCC 6803, * indicates that there is an additional Fe-S cluster on
subunit NuoG of E. coli.
6
Several prokaryotes also possess type I NADH dehydrogenases (Dupuis et al., 1995;
Weidner et al., 1993). The type I NAD(P)H dehydrogenase, or NDH-1, found in
prokaryotes is a multi-subunit complex with a minimum of 14 subunits in E. coli
(Weidner et al., 1993) and Rhodobacter sphaeroides (Dupuis et al., 1995). The NDH-1
complex translocates protons across the membrane, has a flavin mononucleotide and
iron-sulfur clusters as the prosthetic groups, and is inhibited by rotenone in a manner
similar to mitochondrial complex I (Weidner et al., 1993). Thus, biochemically, the
functions of the prokaryotic NADH dehydrogenase are similar to those of the
mitochondrial enzyme.
The nomenclature used for the chloroplast ndh genes is also used for type I
NADH dehydrogenase gene homologs within cyanobacteria. Cyanobacteria also have
another open reading frame, ndhL, associated with the type I NADH dehydrogenase
(Kaneko et al., 1996; Ogawa, 1991; Sugita et al., 1995). The ndhL gene was originally
isolated by Ogawa (Ogawa, 1991) and it was designated ictA (for inorganic carbon
transport gene A) because a mutation which inactivated ictA was found to be defective in
inorganic carbon transport. The NdhL protein is unique to cyanobacteria. In
Synechocystis sp. PCC 6803, the NDH-1 complex was found on both the thylakoid and
cytoplasmic membranes (Schmetterer, 1994). Genes for 16 single-copy ndh genes have
been identified in the Synechocystis sp. PCC 6803 genome. There are multiple copies of
the ndhF and ndhD genes in Synechocystis sp. PCC 6803 (Kaneko et al., 1996).
However, as predicted in chloroplast NDH-1 complexes, there are no genes predicted to
encode subunits involved in NAD(P)H/H+ binding (Kaneko et al., 1996) that are encoded
7
by the nuoE, nuoF, and nuoG genes in E. coli (Weidner et al., 1993). This suggests that
cyanobacteria and chloroplast type I NADH dehydrogenases may use an electron donor
or acceptor other than NAD(P)H/H+ as a donor/acceptor of electrons. Other proteins may
have taken over this function in cyanobacteria (see below).
Inactivation of genes encoding individual subunits of the NDH-1 complex in
cyanobacteria has shown that the NDH-1 complex is active in both photosynthetic and
respiratory processes (Klughammer et al., 1999; Marco et al., 1993; Ogawa, 1991;
Schluchter et al., 1993). Inactivation of the ndhB genes in Synechocystis sp. PCC 6803
(Ogawa, 1991) and in Synechococcus sp. PCC 7942 (Marco et al., 1993). A similar
phenotype was also observed when the ndhD3 and ndhF3 genes of Synechococcus sp.
PCC 7002 were inactivated (Klughammer et al., 1999). However, inactivation of the
ndhF1 gene in Synechococcus sp. PCC 7002 showed that it is involved in both cyclic
electron transport around PSI as well as respiratory electron transport (Schluchter et al.,
1993). These observations suggest that there are multiple types of NDH-1 complexes,
possibly consisting of different subunits, in cyanobacteria.
1.1.2 Type II NADH dehydrogenases
A second type of NADH dehydrogenase, the type II NADH dehydrogenase or
NDH-2, is found in prokaryotes. The NDH-2 protein is a single subunit, has no iron-
sulfur clusters, and does not appear to have the ability to translocate protons across the
membrane. The enzyme also has a flavin adenine dinucleotide (FAD) as a cofactor and
8
unlike NDH-1, is not inhibited by rotenone. This enzyme is found in E. coli (Blattner et
al., 1997), Bacillus sp. YN-1 (Xu et al., 1991), Bacillus megaterium (Thiaglingam and
Yang, 1993) and Thermus thermophilus (Yagi et al., 1988). Recently, three open reading
frames (slr0851, slr1743, sll1484) were identified in Synechocystis sp. PCC 6803 that
exhibited low sequence similarity to the NDH-2 proteins from E. coli, Bacillus sp. YN-1
(Howitt et al., 1999; Kaneko et al., 1996; Xu et al., 1991) and these genes have been
denoted ndbA, ndbB, and ndbC. All three putative proteins have characteristic FAD and
NADH binding motifs and, in contrast to the E. coli NDH-2 protein, all three predicted
NDH-2 proteins from Synechocystis sp. PCC 6803 appear to be hydrophilic (Howitt et
al., 1999). The predicted protein from the sll1484 reading frame is the only predicted
protein product containing a hydrophobic stretch of amino acids that would be long
enough to span a membrane. Expression plasmids were made with slr0851 and slr1743
to see if they could complement a strain of E. coli that lacks a functional NDH-2 or
NDH-1 (Howitt et al., 1999). It was shown that slr1743 was able to complement the
mutant E. coli strain lacking a functional type II NADH dehydrogenase, thus showing
that cyanobacteria contain a functional type II NADH dehydrogenase (Howitt et al.,
1999). Howitt et al. (1999) also made interposon mutations in all three of the NDH-2
reading frames resulting in strains of cyanobacteria that were deplete in one, two, or all
three of the NDH-2 proteins. They also deleted these genes in Synechocystis sp. PCC
6803 strains that lacked PSI. They discovered a very unusual phenotype in that PSI-
deficient strains that were also Ndb- were able to grow in high light whereas the parental
PSI-deficient strain was unable to grow under this condition (Howitt et al., 1999). The
9
results of this study imply that the type II NADH dehydrogenase may function as an
important redox sensor of the membrane plastoquinone pool and the soluble fraction of
NADH within the cell.
1.1.3 Hydrogenase
Hydrogenase enzymes are found in a wide variety of microorganisms, and they
catalyze the reaction: 2H+ + 2 e- Ö H2. The physiological function of most prokaryotic
hydrogenases is the oxidation of hydrogen gas coupled to the energy-conserving
reduction of electron acceptors (Wu and Mandrand, 1993). Another function of
hydrogenase enzymes is the production of hydrogen in a non-energy-conserving manner
in order to maintain intracellular pH homeostasis and redox potential balance (Adams et
al., 1980).
Hydrogenases can be divided into several classes according to Wu and Mandrand
(1993). Class I consists of the membrane-bound NiFe proteins. These enzymes are
heterodimers with a large subunit that has a Ni-Fe active site, and a small subunit that
carries multiple Fe-S clusters interacting with the redox, electron transport partners of the
hydrogenase. These so-called uptake hydrogenases are found in bacteria such as
Rhodobacter capsulatus (Colbeau et al., 1993), Bradyrhizobium japonicum (Zuber et al.,
1986), and Azotobacter vinelandii (Seefeldt and Arp, 1986), Anabaena sp. PCC 7120
(Carrasco et al., 1995) and Nostoc sp. PCC 73102 (Oxelfelt et al., 1998; Tamagnini et al.,
1997). The structure has been solved for the NiFe-containing hydrogenase from
10
Desulfovibrio gigas (Volbeda et al., 1995) and the structure reveals that the NiFe cluster
responsible for hydrogenase activity is covalently coordinated to the protein through four
cysteinyl ligands. A mechanism for hydrogen binding and cleavage was proposed to
involve an intermediate state in which hydrogen is bridged between both the Fe and Ni at
the catalytic site (Volbeda et al., 1995).
Class II consists of the NiFe(Se) hydrogenases that also have two subunits. The
large subunit has a Ni-Fe-Se active site and the small subunit, as in Class I, has multiple
Fe-S clusters. The heterolytic cleavage of molecular hydrogen seems to be mediated by
the nickel center and the selenocysteine residue. The selenium ligand might also protect
the nickel atom from oxidation (Garcin et al., 1999). These enzymes are found in sulfur-
metabolizing bacteria such as D. gigas (Malki et al., 1995) and Desulfovibrio
fructosovorans (Rousset et al., 1990).
Class III hydrogenases are the anaerobic Fe-only hydrogenases; these are soluble
enzymes that are made up of one to four subunits. All Class III hydrogenases have a
catalytic subunit composed of 420-580 amino acids (Meyer and Gagnon, 1991). The
large, catalytic subunit has a binuclear Fe center at the active site. This type of enzyme is
found in anaerobic bacterial species such as Clostridium pasteurianum, where the
structure has been determined (Peters et al., 1998). The enzyme has three distinct [4Fe-
4S] clusters, a [2Fe-2S] cluster, and the active site (H-cluster) is an unusual six iron atom
cluster consisting of a [4Fe-4S] cubane cluster that is covalently bridged by a cysteinate
thiol to a [2Fe] subcluster (Peters et al., 1998). The 76 residues at the N-terminus that
form a [2Fe-2S] cluster have a fold similar to plant-type ferredoxins which may shuttle
11
electrons from the redox partners (c-type cytochromes) to the hydrogenase active site and
(Kummerle et al., 1999).
Class IV hydrogenases are the F420-MV NAD reducing NiFe(Se) enzymes found
in archaebacteria such as Methanococcus voltae (Muth et al., 1987) and
Methanobacterium fervidus (Steigerwald et al., 1990). These enzymes can reduce either
the 8-hydroxy-5-deazaflavin cofactor F420 or methyl viologen. The enzymes comprise 3
subunits; α, β, and γ, with molecular masses of 47 kDa, 31 kDa and 26 kDa, respectively.
These subunits form a complex with a subunit stoichiometry of (α1β1γ1)8 (Alex et al.,
1990).
Class V consists of the reversible/bi-directional hydrogenases which are found in
Alcaligenes eutrophus (now Ralstonia eutropha) (Tran-Betcke et al., 1990), Nocardia sp.
(Schmitz et al., 1995), Synechocystis sp. PCC 6803 (Appel and Schulz, 1996; Kaneko et
al., 1996), Anacystis nidulans (Synechococcus sp. PCC 6301) (Boison et al., 1996;
Boison et al., 1998), and Anabaena sp. PCC 7120 (Houchins and Burris, 1981; Schiefer
and Happe, 1999). In R. eutropha, the enzyme is a heterotetramer that can be divided
into two heterodimers that catalyze specific reactions. One heterodimer is responsible for
the hydrogenase activity and the other heterodimer is responsible for the diaphorase
activity.
Cyanobacteria are known to have two types of hydrogenases: uptake
hydrogenases and bi-directional or reversible hydrogenases. The uptake hydrogenase is
located on the thylakoid membrane of cyanobacteria and catalyzes the oxidation of
hydrogen in the presence of phenazine methosulfate (PMS) or methylene blue (MB) as
12
electron acceptors. However, the uptake hydrogenase of cyanobacteria does not support
the methyl viologen-dependent evolution of hydrogen. This type of enzyme is found in
Anabaena sp. (Carrasco et al., 1995), and Nostoc sp. (Oxelfelt et al., 1998; Tamagnini et
al., 1997). The second type of hydrogenase found in cyanobacteria catalyzes both the
oxidation and evolution of hydrogen and thus is called the bi-directional or reversible
hydrogenase. In cyanobacteria, the bi-directional hydrogenases consist of 4-5 subunits.
The enzyme may also be divided into a heterodimeric hydrogenase moiety and a
heterodimeric or heterotrimeric diaphorase moiety. The hydrogenase subunits are
encoded by conserved gene clusters in cyanobacteria and they share sequence similarity
with hydrogenases from other bacteria such as A. eutrophus (Massanz et al., 1998) and
Escherichia coli (Jacobi et al., 1992).
1.1.4 Mobile Electron Carriers
Mobile electron carriers are responsible for the oxidation of the cytochrome b6f
complex and the transfer of electrons to either oxidized P700 in PSI or to cytochrome
terminal oxidases in cyanobacteria. Two soluble proteins which have been reported to
facilitate this electron transfer in cyanobacteria: cytochrome c6 (PetJ) and the blue copper
protein, plastocyanin (PetE) (Ghassemian et al., 1994; Ho and Krogmann, 1984;
Sandmann and Boger, 1981; Zhang et al., 1992).
13
1.1.4.1 Cytochrome c6
Cytochrome c6 (formerly cytochrome c-553) is a small, soluble iron-heme protein
from a unique class of cytochromes, the c-type cytochromes. These c-type cytochromes
are characterized by a heme cofactor that is covalently bound to the CXXCH-consensus
amino acid motif. The fifth and sixth ligands to the Fe of the heme cofactor are a
conserved methionine and histidine in cytochrome c6. Examples of c-type cytochromes
include the c-type soluble cytochromes of mitochondria, mobile cytochrome c-555 in
anoxygenic green photosynthetic bacteria, and cytochrome c2 in purple photosynthetic
bacteria (Dickerson, 1980).
Cytochrome c6 is used in cyanobacteria as a mobile carrier of electrons between
the cytochrome b6f complex and P700 of the PSI reaction center (Zhang et al., 1992).
Many eukaryotic algae and cyanobacteria use either cytochrome c6 (PetJ) or the blue
copper protein, plastocyanin (PetE) as the mobile carrier of electrons between
cytochrome b6f complex and the P-700 reaction center of PSI (Ho and Krogmann, 1984;
Zhang et al., 1992). Whether the organism (cyanobacterium or eukaryotic alga) uses
plastocyanin or cytochrome c6 as an electron donor to PSI depends on the chemical
environment of the cells during growth. If the cells are grown in the presence of copper,
the cells will synthesize plastocyanin for electron transport from cytochrome b6f to PSI;
however, in the absence of copper, the cells synthesize cytochrome c6 (Ho and
Krogmann, 1984; Zhang et al., 1992). Cytochrome c6 undergoes a reversible oxidation-
reduction reaction similar to that of plastocyanin during photosynthetic electron transport.
14
Cytochrome c6 is similar in size, pI and midpoint potential (335 to 390 mV) to
plastocyanin (Kerfeld and Krogmann, 1998).
In addition to its role in photosynthetic electron transport, cytochrome c6 may also
play a role in respiratory electron transport in cyanobacteria. Lockau has presented
evidence in Anabaena variabilis that both cytochrome c6 and plastocyanin have dual
roles as electron carriers for photosynthesis and respiration (Lockau, 1981). Previous
experiments have demonstrated that cytochrome c6-depleted membranes of Nostoc
muscorum could be made competent in transferring electrons to either PSI or cytochrome
oxidase by the addition of cytochrome c6 (Stürzl et al., 1982).
The gene encoding cytochrome c6 (petJ) has been insertionally inactivated in
Synechococcus sp. strain PCC 7942 (Laudenbach et al., 1990) or deleted in the case of
Synechocystis sp. strain PCC 6803 (Zhang et al., 1994). These
interposon/insertion/deletion mutants were still able to grow at wild-type rates. It was
presumed that other physiological electron donors present within these organisms could
replace the function of cytochrome c6. In Synechocystis sp. strain PCC 6803, the primary
electron donor is plastocyanin (Briggs et al., 1990; Zhang et al., 1992); cytochrome c6 is
expressed only under copper depleted conditions (Zhang et al., 1992). However, it
appears that even under copper deplete conditions, Synechocystis sp. strain PCC 6803 is
still able to produce a minimal amount of functional plastocyanin, since the Synechocystis
sp. strain PCC 6803 can still grow under copper-depleted conditions with the petJ gene
inactivated (Zhang et al., 1994). The observation that the petJ- strain was still able to
grow was attributed to a possible, third mobile electron carrier. However, if this were the
15
case, a double mutation to inactivate the petJ and petE genes encoding the cytochrome c6
and plastocyanin proteins, respectively, should have been easy to achieve. However,
Manna and Vermaas (1997) found that the petJ and petE genes could not be
simultaneously inactivated within the same strain.
Subsequent to the report of Laudenbach, et al. (1990) it has been discovered that
Synechococcus sp. strain PCC 7942 has a petE gene encoding a functional plastocyanin
(Clarke and Campbell, 1996) and as in Synechocystis sp. strain PCC 6803, this mobile
electron carrier was probably responsible for the wild-type phenotype seen in the petJ
interposon mutant of this organism.
In other cyanobacteria, it has been observed that some species have more than one
isoform of cytochrome c6, suggesting that there may be multiple copies of the gene
encoding this protein in cyanobacteria (Ho and Krogmann, 1984). This is similar to the
situation in purple bacteria such as R. sphaeroides (Jenney and Daldal, 1993; Jenney et
al., 1996) and R. capsulatus (Jenney and Daldal, 1993; Jenney et al., 1996). It has been
reported that some cyanobacteria (Nostoc muscorum and Spirulina platensis) have no
detectable plastocyanin (Ho and Krogmann, 1984). In these cases, it is assumed that
cytochrome c6 is the sole mobile electron carrier between the cytochrome b6f complex and
either PSI or cytochrome oxidase.
16
1.1.4.2 Plastocyanin
Plastocyanin is a small (97-99 amino acids) soluble electron carrier that transfers
electrons from cytochrome b6f complex to Photosystem I (PSI) via the reversible
oxidation (Cu+ → Cu
2++ 1 e) and reduction (Cu
2+ + 1 e-→Cu
+) of its active copper center
in plants as well as in eukaryotic algae and some cyanobacteria (Ho and Krogmann,
1984). Plastocyanin folds into an eight-stranded, β-sandwich cylinder (Chapman et al.,
1977) and the copper containing active center is close to the surface of the molecule. The
copper ion is coordinated to a surface exposed histidinyl imidazole as well as a cysteinyl
thiolate, a methioninyl thioether and another histidinyl imidazole (Redinbo et al., 1993).
The geometry of the active site is an irregular tetrahedron. This distortion is imposed by
the folding of the protein and is responsible for stabilizing the high midpoint potential
(+370 mV) of the protein. As mentioned in the previous section, plastocyanin and
cytochrome c6 are interchangeable in function in many cyanobacteria and algae, even
though the structures of the individual protein are very different from one another.
Plastocyanin is mostly comprised of β-sheet secondary structure and cytochrome c6 is
mostly alpha-helical in structure (Kerfeld et al., 1995). Ho and Krogmann (1984) noted
that the isoelectric points of plastocyanin and cytochrome c6 from the same organism are
more similar to one another than they are to the pIs of plastocyanin or cytochrome c6
proteins from different organisms. This observation suggests that the similarity of
isoelectric points within the same organism is the result of convergent evolution of
17
plastocyanin and cytochrome c6 within each organism in response to changes in a
common, interacting reaction partners.
1.5 Terminal oxidases
All aerobic bacterial species examined to date have multiple respiratory oxidases
that allow them to modify their respiratory systems according to their environmental
challenges. These respiratory oxidases may fall into either the heme-copper respiratory
oxidase super-family or into the unrelated cytochrome bd oxidase family. Heme-copper
respiratory oxidases can be further divided into two subgroups: (I) heme-copper oxidases
that are reduced by cytochrome c, which includes the mitochondrial cytochrome c
oxidase, as well as the bacterial oxidases of the aa3, ba3, caa3, cao3, and bo3 type; and
(II) those heme-copper oxidases that are directly reduced by quinols (Garcia-Horsman et
al., 1994a).
Membership in the heme-copper oxidase superfamily is determined by the
presence of a homolog to subunit I of the mammalian cytochrome c oxidase. Subunit I is
the largest subunit of the cytochrome c oxidase, and it contains the unique bimetallic or
binuclear center where O2 binds and is reduced to water (Garcia-Horsman et al., 1994a).
This bimetallic center consists of a heme and a copper ion, CuB. Subunit I has a second
heme group in addition to the one at the binuclear center. This secondary heme facilitates
electron transfer to the binuclear center. The mitochondrial heme-copper oxidase is
comprised of 13 subunits, of which 3 (subunits I, II, and III) are encoded by the
18
mitochondrial genome, whereas most bacterial heme-copper respiratory oxidases have 3
to 4 subunits. Three of these bacterial subunits are homologues of subunit I, subunit II,
and subunit III of mitochondria, and the fourth bacterial subunit, when present, is
unrelated to any mitochondrial subunit. Figure 2 shows a scheme with the main catalytic
subunits of the different types of heme-copper oxidases from the heme-copper oxidase
superfamily. The variation observed in the heme-copper oxidases derives from the fact
that individual oxidases use different combinations of the hemes a, o, and b in association
with subunit I. Hemes a, o, and b can reside at the binuclear center and either heme b or
heme a is found at the low-spin site. The proton-conducting channel is also associated
with subunit I (Figure 2).
Subunit II is responsible for shuttling electrons from the redox partner of the
oxidase to the binuclear center. The binding site for cytochrome c is located on subunit
II. Subunit II may also contain a second copper-containing redox center, denoted CuA
(Figure 2). This CuA is the initial electron acceptor from reduced cytochrome c in the
case of cytochrome c oxidase. Amino acid residues in subunit II that are conserved in all
cytochrome c oxidases have been implicated in either the binding of cytochrome c or in
liganding the CuA center (Saraste, 1990). These conserved amino acids are missing in
subunit II of quinol-type oxidases, which additionally do not have a CuA center
(Abramson et al., 2000; Fukaya et al., 1993; Lauraeus et al., 1991; Minghetti et al., 1992).
Under changing oxidative conditions, the cells must balance different needs in
order to optimize their respiratory electron transfer chains. Terminal oxidases may be
used to generate a maximal H+/e- gradient (Puustinen et al., 1991), to remove excess
19
Figure 2. Schematic illustration of the similarities and differences of four subclasses of
the heme-copper oxidase superfamily adapted from (Garcia-Horsman et al., 1994a).
Models indicate the major subunits (I and II) of the heme-copper oxidases as well as
electron donors to the enzyme. Predicted electron flow follows the path of the arrows.
Panels A, B, and C depict three types of cytochrome c oxidases. Panel D depicts the
generic model for a quinol oxidase.
20
Quinol oxidases
Fe CuB
e- e-
H+
H+
FeQH2
D
aa3, ba3, bb3, bo3
II III I
Fe CuB
aa3, ba3
B
A
Cyt c oxidases
Cyt c
Fe CuB
e-
e-
e-
H+
H+
Fe
CuA
e-
e-
e-
H+
Fe
CuA
Fe c
e-
Cyt c caa3, cao3
II I
H+
Cyt c
Fe CuB
e-
e-
H+
H+
Fe
FeFeFe
e-
C
cbb3
I
21
reducing equivalents, to consume oxygen to maintain anaerobicity or to lower the oxygen
concentration for the cell (Kelly et al., 1990). R. sphaeroides represents a model
organism that expresses a diverse number of terminal oxidases that may be used to
regulate its metabolism, since the organism can grow aerobically, anaerobically,
heterotrophically or photosynthetically (Garcia-Horsman et al., 1994a; Garcia-Horsman
et al., 1994b). R. sphaeroides has three distinct respiratory oxidases, two of which utilize
cytochrome c as an electron donor, whereas one terminal oxidase utilizes quinol as a
substrate. The predominant terminal oxidase is of the aa3 type when the cells are grown
aerobically with high O2 tension (Hosler et al., 1992). The alternate cytochrome c
oxidase is a cbb3 type oxidase and is present under microaerophilic conditions and when
the cells are grown under photosynthetic conditions (Garcia-Horsman et al., 1994b). The
quinol oxidase is able to support aerobic growth in R. sphaeroides strains that lack the bc1
complex (Yun et al., 1990).
The majority of studies performed thus far regarding the function of cytochrome
oxidases in cyanobacteria have been directed at examining cytochrome oxidase protein
biochemistry or the function of the enzyme during respiration (Alge and Peschek, 1993b;
Howitt and Vermaas, 1998; Obinger et al., 1990; Peschek et al., 1989; Sone et al., 1993;
Tano et al., 1991; Trnka and Peschek, 1986; Wastyn et al., 1987). Previous biochemical
studies examining P700+ reduction kinetics in cyanobacteria such as Synechococcus sp.
PCC 7002 (Yu et al., 1993) and Fremyella diplosiphon indicate that these organisms may
use cytochrome oxidases as a sink for removing excess electrons not accounted for by
PSI activity (Schubert et al., 1995).
22
The first cyanobacterial ctaCIDIEI operon encoding the primary aa3-type
cytochrome heme-copper oxidase was cloned from Synechocystis sp. PCC 6803 (Alge
and Peschek, 1993a; Alge and Peschek, 1993b) and a similar gene cluster has been
cloned from Synechococcus vulcanus (Sone et al., 1993). Two other respiratory oxidases
were identified and characterized in Synechocystis sp. PCC 6803 by (Howitt and
Vermaas, 1998). One is a quinol oxidase of the bd-type and the second oxidase appears
to be an oxidase of the bo-type. After characterization of mutants lacking all
combinations of the oxidases, it was concluded that ctaCIDIEI and cydAB encode
functional oxidases in Synechocystis sp. PCC 6803. However, these oxidases contributed
little to the normal growth characteristics of the organism under the conditions that were
studied (Howitt and Vermaas, 1998).
1.2 Purpose of the Present Work
The current project sought to identify similarities and differences between the
freshwater, unicellular cyanobacterium Synechocystis sp. PCC 6803 and the marine
cyanobacterium Synechococcus sp. PCC 7002 with regards to genes potentially involved
in either photosynthetic or respiratory electron transport. The purpose of this project was
to clone and sequence the genes for the type I NADH dehydrogenase, type II NADH
dehydrogenases, hydrogenase, mobile electron transport proteins and terminal oxidases
from the marine cyanobacterium Synechococcus sp. PCC 7002 for comparison of amino
acid sequences and gene organizations with other cyanobacteria.
23
Another purpose of this work was to initiate the characterization of some of these
genes and to examine the effects of mutations in specific genes that encode electron
transport proteins on the physiology of Synechococcus sp. PCC 7002. Results from
attempts to inactivate the ndhB and petJ1 genes are described. The initial
characterization of mutations in the following genes are also described: the hydrogenase
gene cluster, two putative mobile electron carriers, petJ2 and bcpA, and the ctaDI and
ctaDII genes, that purportedly encode the large subunits of heme-copper oxidase
complexes. The initial characterization of these genes is an important first step in
understanding the roles that these electron transport proteins play within cyanobacteria,
under different growth conditions.
Chapter 2
MATERIALS AND METHODS
2.1 Bacterial strains and culture conditions
2.1.1 Synechococcus sp. PCC 7002
Synechococcus sp. PCC 7002 is a unicellular or filamentous, naturally
transformable, marine cyanobacterium that was isolated by Van Baalen (Van Baalen,
1962). Synechococcus sp. strain PCC 7002 can be grown as a facultative
photoheterotroph in the presence of glycerol (Rippka, 1972). It has a well-defined
natural DNA uptake system that makes the organism attractive for genetic manipulation
(Stevens and Porter, 1980). The laboratory wild-type strain Synechococcus sp. PCC 7002
(formerly Agmenellum quadruplicatum strain PR6) was originally obtained from the
Pasteur Culture Collection, Unité de Physiologie Microbienne, Institut Pasteur, Paris,
France.
Synechococcus sp. PCC 7002 was grown in medium A (Stevens and van Baalen,
1973) supplemented with 1 g l-1 NaNO3 (referred to as A+ medium) in liquid culture and
on 1.5% (w/v) agar plates. Medium A consists of: 18 g l-1 NaCl, 5 g l-1 MgSO4•7H2O, 1
g l-1 Tris-HCl pH 8.2, 600 mg l-1 KCl, 270 mg l-1 CaCl2, 50 mg l-1 KH2PO4, 30 mg l-1
25
tetrasodium EDTA, 34.3 mg l-1 H3BO3, 4.32 mg l-1 MnCl2 •4H2O, 3.89 mg l-1 FeCl3
•6H2O, 315 mg l-1 ZnCl2, 3 mg l-1 CuSO4•5H2O, 30 mg l-1 MoO3, 12.2 mg l-1 CoCl2
•6H2O, and 4 mg l-1 vitamin B12. Antibiotic concentrations used to select or maintain
mutant strains were 100 µg ml-1 kanamycin, 100 µg ml-1 streptomycin, 100 µg ml-1
spectinomycin, 50 µg ml-1 chloramphenicol, and 100 µg ml-1 erythromycin.
Stock cultures were maintained on 1.5% (w/v) agar in A+ media, containing the
appropriate antibiotic(s) for mutant strains of Synechococcus sp. PCC 7002, as necessary,
in Petri dishes at approximately 28˚C under continuous illumination at an approximate
light intensity of 60 µE m-2 s-1. Individual colonies for each strain were re-streaked on
fresh plates once every three weeks.
Small-scale liquid cultures (25 ml) were grown in 22 × 175 mm culture tubes at
38˚C in aquarium water baths to maintain constant temperature. For large-scale cultures,
cells were grown in sterile flasks at room temperature and were constantly bubbled with
CO2 in air with continuous stirring. Constant illumination was provided with
F72T12/CW fluorescent bulbs. The standard illumination conditions for liquid cultures
were approximately 200-300 µE m-2 s-1. For lower light intensities, paper was used to
shade the aquariums from light until the appropriate light level was achieved. For higher
light intensities, 150 W halogen bulbs were added at appropriate distances to obtain the
surface light intensity desired. Light intensities were measured using a model QSL-100
quantum scalar irradiance meter (Biospherical Instruments, Inc., San Diego, CA). For
growth curve measurements, all strains were grown under standard conditions (250 µE
m-2s-1 at 38°C with 1-5% (v/v) CO2/air constantly bubbling) into exponential phase and
26
were diluted to an OD550nm = 0.05 prior to a shift to a different light intensity. Once the
cells were shifted to a new light intensity, the cell growth was monitored at that new light
intensity over a 24-hour period.
Growth was determined by monitoring the turbidity of cells at 550 nm in a
Bausch and Lomb (now Milton Roy, Rochester, NY) Spectronic 20 spectrophotometer.
To determine the doubling time, the absorbance of cells in various growth phases was
measured at various time points, minimally in triplicate, and the results were plotted on a
semi-log scale.
2.1.2 Synechocystis sp. PCC 6803
Synechocystis sp. PCC 6803 is a unicellular, naturally transformable freshwater
cyanobacterium (Grigorieva and Shestakov, 1982). Synechocystis sp. PCC 6803 cells
were grown at 32˚C in liquid medium BG-11 (Stanier et al., 1971) buffered with 5 mM
HEPES in 22 x 175 mm culture tubes bubbled with 1.5% CO2 in air and on 1.5% (w/v)
agar plates in B-HEPES medium. B-HEPES medium consists of: 1.5g l-1 NaNO3 , 50 mg
l-1 KH2PO4, 75 mg l-1 MgSO4•7H2O, 272 mg l-1 CaCl2, 6.56 mg l-1 citric acid•H2O, 12
mg l-1 ferric ammonium citrate, 2 mg l-1 tetrasodium EDTA, 2.86 mg l-1 H3BO3, 20 mg
l-1 NaCO3, 1.81 mg l-1 MnCl2 •4H2O, 3.89 mg l-1 FeCl3 •6H2O, 222 mg l-1 ZnCl2, 390
mg l-1 NaMoO4•2H2O 79 mg l-1 CuSO4•5H2O, 49.4 mg l-1 Co(NO3)2•6H2O, 1.1 g l-1
HEPES pH 8.0 (titrated with 2 M KOH). Cells in liquid culture were bubbled constantly
27
with 1.5% CO2. Light intensities were 100-150 µE m-2 s-1 for growth of liquid cultures
and 60 µE m-2 s-1 for stock cultures on plates.
2.1.3 Escherichia coli
E. coli DH5α (genotype: F-, endA, hsdR17, supE44, recA1, gyrA96, relA1, argF)
(Bethesda Research Laboratories, Gaithersburg, MD) was used for all recombinant DNA
manipulations. This strain is a recombination-deficient, phage-suppressing strain that is
capable of α-complementation with the amino-terminal, α−fragment of beta-
galactosidase that is encoded by pUC and pBluescript vectors. The cells were grown in
liquid cultures or on 1.5% (w/v) agar plates of LB (Luria-Bertani) medium at 37˚C. LB
medium contains 1% (w/v) bacto-tryptone, 0.5% (w/v) yeast extract, and 1% (w/v) NaCl,
pH 7.5 (adjusted with NaOH). Antibiotics (ampicillin 100 µg ml-1, kanamycin 40 µg ml-
1; chloramphenicol 25 µg ml-1; and erythromycin 50 µg ml-1) were added when required.
E. coli BL21(DE3) (genotype: F-, ompT, rB-, mB- (DE3)) and E. coli
BL21(DE3)(pLysS) (Novagen, Madison, WI) were used for overproduction of proteins.
The cells were grown in NZCYM medium (pH 7.0) containing 1% (w/v) bacto-tryptone,
0.5% (w/v) yeast extract, 0.5% (w/v) NaCl, 0.1% (w/v) casamino acids, and 0.2 % (w/v)
MgCl2•7H2O.
28
2.2 Standard laboratory methods
Standard recombinant DNA procedures were performed according to the
recommendations of the manufacturer of the respective kit or enzyme and by consulting
current laboratory manuals (Ausubel et al., 1987; Sambrook et al., 1989). Restriction
endonucleases and other DNA modifying enzymes were purchased from New England
BioLabs (Beverly, MA), Promega Corporation (Madison, WI), Bethesda Research
Laboratories (Gaithersburg, MD), and Boehringer Mannheim Biochemicals
(Indianapolis, IN). Radiolabelled [α-32P] dATP and [α-35S] dATP were purchased from
New England Nuclear Products, Dupont de Nemours & Co. (Boston, MA).
DNA fragments generated by restriction digests were separated by electrophoresis
on agarose gels and purified using GENECLEAN® DNA isolation kit (Bio101, La Jolla,
CA), or isolated from the excised gel slice with Sigma (formerly Supelco)-spin columns
(Sigma, Bellefonte, PA). DNA fragments were labeled with [α−32P] dATP, using a
Random Primed DNA Labeling kit (Boehringer Mannheim Biochemicals, Indianapolis,
IN).
Polymerase chain reaction (PCR), using Taq polymerase from Boehringer
Mannheim Biochemicals (Indianapolis, IN), was performed with one of the following
thermocyclers: a Barnstead/Thermolyne Corporation Temp•Tronic® Series 669
thermocycler (Dubuque, IA), a Techne Progene thermocycler (PGC Scientific, Frederick,
MD), or an Eppendorf® Mastercycler Gradient Thermal Cycler (Fisher Scientific,
29
Pittsburgh, PA). Annealing temperatures were varied ± 5˚C from the calculated Tm of
the synthetic primers (as calculated in the MacVector version 6.5 software package,
Oxford Molecular Group). Reactions of 30-35 cycles were generally carried out with a
denaturation temperature of 94˚C and an extension reaction temperature of 72˚C.
Template quantities per reaction for plasmid DNA and chromosomal DNA were
generally 1-10 ng and 100-500 ng, respectively. 50 pmoles of each primer was added to
the reactions.
Radioactive signals from Southern blots, Northern blots, colony filters, and
sequencing gels were exposed to Kodak X-OMAT AR, BIOMAX MR, or BIOMAX MS
X-ray film using intensifying screens (Lightening Plus, DuPont, Wilmington, DE) at -
80˚C or to a Molecular Dynamics PhosporImager screen when appropriate.
2.3 Isolation and manipulation of DNA
2.3.1 Plasmid and cosmid isolation from Escherichia coli
An alkaline lysis method was used for small-scale plasmid and cosmid DNA
preparations from E. coli. The cells were resuspended in 200 µl of solution 1 (0.05 M
Tris-HCl (pH 7.5), 0.01 M EDTA (pH 8.0), 0.1 mg ml-1 RNaseA). An equal amount of
solution 2 was added (0.2 M NaOH, 1% SDS) and as soon as the suspension cleared, 200
µl of solution 3 (3M NaOAc at pH 5.0) was added. Cell debris was pelleted by
centrifugation and the cleared supernatant was transferred to a new tube. The plasmid
30
DNA was precipitated by the addition of an equal volume of isopropanol to the collected
aqueous solution.
An alkaline lysis method was also used for large-scale plasmid isolations
(Birnboim and Doly, 1979). Plasmid DNA was separated from chromosomal DNA by
CsCl-ethidium bromide, equilibrium density-gradient ultracentrifugation (Sambrook et
al., 1989). The super-coiled plasmid DNA fraction was collected from the gradient,
extracted with NaCl-saturated isopropanol (to remove ethidium bromide), and dialyzed
against TE buffer (10mM Tris-HCl pH 8.0, 1 mM EDTA).
2.3.2 Total DNA isolation from Synechococcus sp. PCC 7002 and
Synechocystis sp. PCC 6803
For large-scale preparations of total DNA from Synechococcus sp. PCC 7002 or
Synechocystis sp. PCC 6803, cells were isolated by first collecting 1 L of exponentially
growing cells by centrifugation at 5000×g. The resulting cell pellet was resuspended in 4
ml of 5 mM Tris-HCl (pH 7.5), 5 mM EDTA, 50 mM NaCl with 3 mg ml-1 lysozyme.
The cells were then incubated for one hour with agitation to keep the cells in suspension
at 37˚C. The cells were subsequently lysed by adding 400 µl of 10% (w/v) sarkosyl and
an equal amount of Tris-buffered phenol. This mixture was agitated by vortexing for 15
min. The organic and aqueous phases were separated by centrifugation at 8000×g for 10
min. The aqueous layer was collected and 1 ml of 5.0 M NaCl and 1 ml of 10% (w/v)
cetyltrimethylamine bromide (CTAB)-700 mM NaCl were added to remove
31
polysaccharides. An equal volume of chloroform was added to the mixture and the
suspension was agitated by vortexing or shaking and subsequently centrifuged at 8000Xg
for 5 min to separate the organic and aqueous phases. The aqueous phase was collected
and the DNA was precipitated by the addition of an equal amount of isopropanol to the
aqueous phase.
Small-scale chromosomal DNA extractions were performed by harvesting
exponentially growing cyanobacterial cells (25 ml) by centrifugation at 5000×g for 5
min, resuspending the cell pellet in 500 µl of lysis buffer (10% (w/v) sucrose, 50 mM
Tris-HCl pH 8.0, 10 mM EDTA), and subjecting the suspension to a freeze/thaw cycle.
Lysozyme was added (5 mg) and the mixture was incubated at 37˚C for 30 min. SDS (to
1% w/v) was added and the resulting solution incubated at 37˚C for one hour before
phenol:chloroform extractions were performed. The aqueous phase was removed and
combined with NaCl (to 1 M), one-fifth volume CTAB/NaCl solution (10% (w/v) CTAB,
700 mM NaCl), and an equal volume of chloroform to separate polysaccharides from the
DNA. DNA was precipitated with an equal volume of isopropanol.
2.4 Transformation procedures
2.4.1 E. coli transformation procedures
E. coli cells were prepared for transformation using the following methods. To
prepare cells for transformation by electroporation, E. coli of the desired strain (BL21R,
32
BL21(DE3), DH5-a, or DH10-B) were streaked onto an LB plate with no antibiotics from
an axenic freezer stock and allowed to grow overnight at 37°C. A single colony was
selected from the plate and transferred to 5 ml of LB liquid media and grown overnight at
37°C with vigorous shaking. This 5 ml starter culture was transferred to 1L of LB and
grown for 2-3 hours at 37°C to an OD500 of 0.5-1.0. Cells were harvested by
centrifugation in sterile tubes by at 4000×g at 4°C for 15 min. The cell pellet was
resuspended in 1L of sterile, cold water and collected by centrifugation (4000×g, 4°C) for
10 min. The cells were washed three more times with 500 ml of sterile cold water, and
were resuspended in 3 ml of filter-sterilized cold 10% (v/v) glycerol, and aliquots (40 µl)
of this solution was transferred into Eppendorf tubes in. Plasmid DNA (10-50 ng) was
added to E. coli cells that had been prepared for transformation. E. coli cells were
transformed by electroporation (Transporator Plus, BXT, San Diego, CA, USA) at 1.5V
using ice-cold transformation cuvettes with a 1.5 mm gap space. The cells were
incubated in 1 ml of SOC or LB for 30-60 min at 37°C. Aliquots of the cells incubated in
SOC were transferred to LB plates containing the appropriate antibiotic(s).
Transformants were usually selected by blue-white screening for initial clones on LB
ampicillin (100 µg ml-1) and X-gal (40 µg ml -1) agar plates.
2.4.2 Cyanobacterial transformation procedures
Synechococcus sp. PCC 7002 cells are naturally transformable (Stevens and
Porter, 1980) and take up either circular or linear DNA. Insertional inactivation of genes
33
was performed by interposon mutagenesis. A DNA fragment containing a gene
conferring resistance to an antibiotic was ligated into a restriction site within the coding
region of the gene selected for mutation. Replacement of the wild-type gene in the
Synechococcus sp. PCC 7002 genome occurred via double crossover homologous
recombination by transforming cells with linear DNA fragments containing an
insertionally inactivated gene. Generally, at least 200 bp of homologous sequence were
flanking each side of the drug resistance cartridge. Approximately 1-2 µg of linear DNA
was added to 0.9 ml of cells that were grown to a transmittance of 20% at 550 nm. The
culture was exposed to full light and bubbled with 1.5% CO2 in air for 1.5 doubling
times. Aliquots of this transformation mixture were spread on A+ plates and incubated at
low light intensity for 2-3 days. Plates were then overlaid with the appropriate antibiotic
in 0.8% top agar and incubated under standard light intensity. Transformants were visible
4-10 days after being challenged with the appropriate antibiotic. Single colonies were
inoculated into liquid culture (10-20 ml), and the resulting cultures were, again, diluted 4-
6 times with media containing the appropriate antibiotic in order to allow segregation of
mutant and wild-type alleles. Southern blot hybridization analyses and PCR
amplifications of chromosomal DNA from the transformants were performed to verify
the gene interruption. Amplification of the DNA from whole cells was performed as
described (Howitt et al., 1996). Synechococcus sp. PCC 7002 cells were resuspended in
sterile water at an optical density at 550 nm of 15 ml-1. One µl of this suspension was
used in a standard PCR reaction.
34
2.5 Southern blot transfer and hybridization
Agarose gels containing restricted DNA fragments that had been separated by
electrophoresis were photographed alongside a fluorescent ruler under UV light using the
Biophotonics (Ann Arbor, MI) Gel Print 2000i video imaging system. The DNA was
denatured by soaking the gel in denaturation solution (0.5 M NaOH, 1.5 M NaCl) for 1
hour and subsequently neutralized by soaking in neutralizing solution (0.5 M Tris pH 8.0,
1.5 M NaCl) for 1 hour. The DNA fragments were transferred to nitrocellulose
membranes or nylon membranes (Schleicher & Schuell, Keene, NH) by capillary action
in 10X SSC (0.15 M sodium citrate pH 7.0, 1.5 M NaCl) overnight. Nitrocellulose and
nylon membranes were baked for 1 hour under vacuum at 80˚C before use in
hybridization experiments. Pre-hybridization of the nitrocellulose filters or nylon filters
was performed in a modified hybridization buffer described by Church and Gilbert
(1984). The modified version of the hybridization buffer consists of 5% (w/v) SDS,
0.5M NaPO4, and 1% (w/v) Bovine Serum Albumin (fraction V from Sigma, St. Louis).
The temperature of incubation determined the stringency of the hybridization and was
determined empirically (Bryant and Tandeau de Marsac, 1988). Typically, low-
stringency hybridizations were carried out between 50-55˚C, whereas high-stringency
conditions were 60˚C - 70˚C. Following hybridization for at least 12 hours, blots were
washed with wash buffer (Church and Gilbert, 1984) at the hybridization temperature six
times or until there were no radioactive counts in the wash eluent. The wash buffer
consisted of 1% (w/v) SDS, 40 mM NaPO4, pH 7.2.
35
2.6 Cloning and sequencing
2.6.1 General cloning strategy
The following procedures were adapted for gene cloning from Ausubel et al. (1987)
and Maniatis et al. (1982). First, a Southern blot hybridization experiment was carried
out with chromosomal DNA, which had been digested with various combinations of
restriction enzymes. The respective hybridization probe was chosen to maximize the
signal intensity of the targeted gene to be cloned. In some cases several hybridization
experiments were necessary to determine this. After analysis of the results of the
hybridization experiment(s), a convenient restriction fragment was chosen for cloning
based on size and restriction enzyme cutting efficiency. In the case of Synechococcus sp.
PCC 7002 a partial cosmid library was available, and if genes from this organism were to
be cloned, this library was first screened to determine if the gene of interest was present.
Otherwise, size-fractionated libraries of fragments were created by restriction digestion
of chromosomal DNA, by gel-purifying the desired range of fragments, and by ligating
the fragments into a plasmid vector. The usual vector of choice was the high-copy-
number vector pUC19 (Yanisch-Perron et al., 1985) or pBluescript SK+ (Stratagene, La
Jolla, CA). The fragment library was transformed into competent E. coli DH5α cells,
and colonies were screened to be able to identify the targeted gene. Approximately 1000
colonies were generally analyzed by colony hybridization with the same probe that was
36
used for the initial Southern hybridization. Positive clones were selected and confirmed
by further hybridization experiments. Final confirmation was obtained by DNA sequence
analysis.
2.6.2 Construction of a Synechococcus sp. PCC 7002 genomic library
A total of 701 E. coli DH5-α clones made up the genomic library. Each clone
harbors a cosmid containing a Synechococcus sp. PCC 7002 DNA fragment insert
ranging in size from 20-kb to 45-kb. 576 clones originated from genomic DNA partially
digested with EcoRI and ligated into cosmid vector pHC79. The remaining 125 clones (a
gift from Dr. William R. Widger, University of Houston, USA) were made by partially
digesting genomic DNA with Sau3A and with subsequent insertion of the resulting
fragments into the cosmid vector SuperCos (Stratagene, La Jolla, CA, USA). The library
was maintained on eight 96-well master plates; each clone was grown in 100 µL LB
medium containing ampicillin, and stored at -80˚C after the addition of sterile glycerol to
a final concentration of 10 % (v/v).
2.6.3 Probes
Genes putatively involved in electron transport were identified by sequence
homology from the fully sequenced genome of the cyanobacterium Synechocystis sp.
PCC 6803 (Kaneko et al., 1996) and primers were made to amplify these genes (Table 2)
37
Table 2. Synechocystis sp. PCC 6803 electron transfer protein gene PCR primers
Primer name Nucleotide sequence (5’-3’) ORF number gene nameslr1379R TTA CTC CTT AAC AAC AGC slr1379 cydAslr1379F ATG CAG GAT TTT TTG AGT AAC slr1379 cydAslr1185R TCA AAC CCA CCA GGG slr1185 petCslr1185F ATG GAT AAC ACA CAG GCG slr1185 petCslr1281F GTG GCT GAG GAA GTG AAC slr1281 ndhJslr1281R CTA ATA GGC ATC CTG GAG slr1281 ndhJslr1280R TCA GCC ACG GTT TAA TTG slr1280 ndhKslr1280F ATG AGT CCC AAC CCT GCT slr1280 ndhKsll0823F ATG GAA ATT GTT TGC AAA ATA sll0823 sdhBsll0823R TTA AAG GGG ATC CAT CAA GTC sll0823 sdhBsll0629F ATG ACG GCG ATC GCC AGG sll0629 psaKsll0629R CTA GAG AAT GCC ACT ACT AGC sll0629 psaKslr2009F ATG AAT TTT CCC ACG GCG slr2009 ndhFslr2009R TCA TAT TAA TAC CGA GCT slr2009 ndhFslr0261F ATG ACC AAG ATT GAA ACC slr0261 ndhHslr0261R CTA GCG GTC CAC CGA TCC slr0261 ndhHslr2007F ATG ACC CTA TTT GCA GTT slr2007 ndhDslr2007R TCA TAC TAA AAT CCA GGC slr2007 ndhDsll0027F ATG CTC AGT GCT CTC ATC sll0027 ndhDsll0027R CTA CAC TTC CCC CAA TTC sll0027 ndhDslr0851R CTA GGG GCC CAC TAG TTC slr0851 ndbAslr0851F ATG AAT TCC CCC ACC TC slr0851 ndbAslr1743F ATG ACG GAC GCT CGA CC slr1743 ndbBslr1743R GGA AGG TTC ATT TTT GAG slr1743 ndbBslr1484R CTA CCG ATT CAT ACC GG slr1484 ndbCslr1484F GTG GGG TTC CAG TTT CCA slr1484 ndbC5'1226 ATG TCT AAA ACC ATT GTT ATC sll1226 hoxH3' 1226 TTA ATC CCG CTG GAT GGA sll1226 hoxH5'ndhB ATG GAC TTT TCT AGT AAC sll0223 ndhB3'ndhB CTA GGG TAA ATC ATG GGA sll0223 ndhBslr1743.1 ATG ACG GAC GCT CGA CCA slr1743 ndbBslr1743.2 GAG CCA ATC CCC CAG GGG slr1743 ndbBslr1279 GTG TTT GTT TTA ACC GGT TAC slr1279 ndhCslr1279R CTA GGA CCA CTC CAG AGC slr1279 ndhCsll0026 250F CGG TGG AGA TTT CCC CGG sll0026 ndhFslr1380R CTA GTC GGT GAC AAT TTT GCC slr1380 cydBslr1380F ATG GAA CCG TTA GAA CCG slr1380 cydB
38
by PCR. Amplified PCR products were electrophoresed on standard agarose gels and
purified as described previously (see 2.2). The purified Synechocystis sp. strain PCC
6803 PCR products were labeled with (α32P)-dATP by use of a Random Primed DNA
Labeling Kit (Boehringer Mannheim, Germany) and used as probes to screen cosmid
libraries, partial genomic libraries and genomic DNA blots. Hybridization probes made
from Synechococcus sp. PCC 7002 DNA that had been isolated from clones or PCR
products, were made in the same way as those for Synechocystis sp. PCC 6803.
2.6.4 Colony hybridization
Colony hybridizations were carried out as described by Sambrook et al. (1989).
Usually 100 individual transformants were transferred to a fresh LB agar plate containing
the appropriate antibiotic(s) with a sterile loop. The cells were scraped into a grid
template and the plate was marked on a single side as a frame of orientation for the
colonies. The plate was replicated as described in Sambrook et al. (1989), and one plate
served as the reference plate. The colonies were lifted from one plate onto circular
nitrocellulose membranes (Schleicher & Schuell, Keene, NH), or in the case of the
rectangular plates containing the cosmid library onto rectangular-cut Nytran-Plus
membranes (Schleicher & Schuell, Keene, NH). After the colony transfer, the
membranes were placed on filter paper soaked with denaturation solution to lyse the cells
and denature the DNA. The membranes were then transferred to a filter paper soaked in
neutralization solution to neutralize the denaturation solution. The membranes were air-
39
dried between the two steps. These are the same solutions as described above for
Southern transfers. The following steps are also similar to the steps leading to
hybridization described in the section on Southern transfers. To lower the background
hybridization of colony filters, cell debris was removed by gently rubbing the pre-
hybridized filters, fresh pre-hybridization solution was added, and the filters were pre-
hybridized for at least another hour prior to probe addition.
2.6.5 DNA sequencing and analysis
The sequences of all cloned genes were completely determined on both strands.
Manual sequencing was carried out by the dideoxy chain-termination method (Hattori
and Sakaki, 1986; Tabor and Richardson, 1987a) following the protocol supplied with the
SEQUENASE® Version 2.0 DNA sequencing kit (U. S. Biochemical Company, Inc.,
Cleveland, OH). The templates were base-denatured (Hattori and Sakaki, 1986) prior to
sequencing. Sequencing reactions were electrophoresed on 5% (w/v) acrylamide
(acrylamide:bis-acrylamide, 19:1 (w/w)) gels (40 cm x 33 cm x 0.4 mm) containing 8 M
urea and TBE buffer (89 mM Tris pH 8.3, 89 mM boric acid, 2.75 mM EDTA) at a
constant power of 52-60 W. Gels were soaked in 10% (v/v) acetic acid, 10% (v/v)
methanol for 15-30 min before being transferred to Whatman paper (3MM) and dried
under heat and vacuum. Dried gels were exposed to Kodak X-Omat AR or Biomax AR
film. Other sequencing reactions were performed at the Nucleic Acid Facility at The
Pennsylvania State University with an ABI PRISM 377 DNA sequencer (Perkin-Elmer
40
ABI, Foster City, CA, USA) by using the method of 3' BigDye-labeled
dideoxynucleotide triphosphates. Sequences were assembled in MacDNASIS Pro V2.5
(Hitachi Software Engineering Co., Ltd., Japan). Other sequence manipulations were
performed with MacVector 6.5 (Oxford Molecular Co., Ltd., UK) software. Amino-acid
sequences obtained in this work were compared with sequences obtained from the
National Center for Biotechnology Information (NCBI) databases and aligned with the
ClustalW multiple sequence alignment program (Thompson et al., 1994a) by using
default parameters within the MacVector 6.5 program.
2.7 RNA isolation, Northern-blot hybridization, and RT-PCR analysis
2.7.1 RNA isolation
All glassware was baked at 400˚C for 4 hours, aqueous solutions were treated with
0.1% (v/v) DEPC (diethylpyrocarbonate) for 12 hours, and gloves were worn throughout
the entire procedure in order to prevent contamination by RNases. Total RNA from
Synechococcus sp. PCC 7002 was isolated using a modified mechanical breakage
protocol (Golden et al., 1987). Cultures (150-200 ml) were harvested by centrifugation
and the cells were resuspended in 5 ml 50/100 TE (50 mM Tris-HCl pH 8.0, 100 mM
EDTA) and transferred to glass centrifuge tubes. Chloroform was added and incubated
with the cell mixture for 5 min on ice with occasional shaking. After centrifugation at
5000×g for 10 min at 4˚C, the cell pellet was resuspended in 5 ml of breakage buffer (30
41
ml 50/100 TE, 0.15 ml Triton X-100, 0.6 ml 25% (w/v) sarkosyl, 0.6 ml 20% (w/v)
SDS). Acid-washed, baked 0.45-mm glass beads (4.0 g) and a 1:1 mixture of
phenol/chloroform (5 ml) were added. This suspension was vortexed in three bursts of 3
min each, with incubation on ice for 2 min between each burst. After centrifugation, the
aqueous phase was transferred to a new tube and extracted twice with phenol/chloroform
and twice with chloroform. The RNA and DNA was precipitated with ethanol, pelleted
by centrifugation, dried at room temperature, and resuspended in 50/100 TE. The RNA
was separated from DNA by precipitation with 2.5 M LiCl (Sakamoto and Bryant, 1997).
RNA concentrations were quantified by a fluorometric assay using the dye SYBR
Green II (Molecular Probes, Inc., Eugene, OR) as described by Schmidt and Ernst (1995)
and/or by inspection of a UV spectrum for the RNA sample. A standard curve was made
by measuring the fluorescence of known quantities of RNA, which had been determined
by measuring the absorbance of a stock solution at 260 nm with a Cary 14
spectrophotometer (Olis, Bogart, GA); An absorbance of A260nm = 1.0 was assumed to
contain 40 µg ml-1 RNA.
2.7.2 Northern-blot hybridization analyses
RNAs were size-fractionated by electrophoresis through 1.5 % (w/v) agarose gels
made with running buffer (50 mM HEPES pH 7.8, 1 mM EDTA including 16% (v/v)
formaldehyde). An equal volume of loading buffer (50% (v/v) formamide, 16% (v/v)
formaldehyde, HEPES/EDTA buffer, 10% (w/v) glycerol, 0.025% (w/v) xylene cyanol,
42
0.025% (w/v) bromphenol blue, 0.025% (w/v) ethidium bromide) was added to each
RNA sample. The samples were then incubated in a 68˚C water bath for 5 min prior to
loading on the gel. Gels were typically run at 25 volts for 16 hours in the dark at room
temperature. The ribosomal RNA bands were visualized under UV light and
photographed with a Biophotonics (Ann Arbor, MI) Gel Print 2000i video imaging
system. The formaldehyde-agarose gels containing RNA were soaked in distilled water
five times for 30 min. This was followed by a 30-minute soak in 20×SSC. The filter
papers were also soaked in 20×SSC. The membrane was soaked in 10×SSC prior to
transfer of the RNA by capillary action to Nytran Plus nylon membranes (Schleicher &
Schuell, Keene, NH) using 20×SSC (1×SSC = 0.015 M Na citrate pH 7.0, 0.15 M NaCl)
for 16-24 hours. After the transfer, the RNA was fixed to the nylon membrane for two
min by UV crosslinking. A slightly modified set of hybridization and washing conditions
were used as described in Church and Gilbert (1984). The membranes were incubated in
hybridization buffer (5% (w/v) SDS, 0.5 M NaPO4, pH 7.2, 1% (w/v) Bovine Serum
Albumin, Fraction V) for a minimum of 2 hours. After this time, the hybridization buffer
was replaced and a denatured gene-specific probe was added to the membranes. The
membranes and probe were incubated overnight at 55°C. The membranes were washed
with wash buffer (1% (w/v) SDS, 40 mM NaPO4, pH 7.2) six times for 15 min each until
no counts were detectable with a pancake radiation monitor.
43
2.7.3 Reverse transcriptase-polymerase chain reaction (RT-PCR)
The following primers were used for the RT-PCR reactions with the
Synechococcus sp. PCC 7002 total RNA or cDNA templates. For ctaCII, primer ctaCII.1
(5'-CAC TTT CGG CGA TCG CCC TAC TTT TGG GGG-3') and primer ctaCII.2 (5'-
GGG ATG ATA ATT CAC CAC-3') were used. For ctaDII, primer ctaDII.1 (5'-CCA
TGA CCC AAG CTC CC-3') and primer ctaDII.2 (5'-CGG GAA CCA GTG GTA CAC
CGC-3') were used. For ctaEII, primer ctaEII.1 (5'-GAC TGC CAT CAA TGA AAC C-
3') and primer ctaEII.2 (5'-ATT GCC AGA GAT AAA TCA GCC-3') were used. 1 µg of
total RNA isolated from Synechococcus sp. PCC 7002 was diluted to a final volume of 20
µl with 20 µM of primer (either primer 683, primer 1407, or primer 1288) and incubated
at 70˚C for 5 min to denature secondary structure. Each mixture was briefly incubated on
ice. 5 µl of 5×M-MLV Reaction buffer (5 µl of 5×M-MLV Reaction buffer = 250 mM
Tris-HCl, pH 8.3, 375 mM KCl, 15 mM MgCl2, 50 mM DTT), 1µl of 25 mM dNTPs,
0.625 µl RNAsin, and 1.5 µl of M-MLV reverse transcriptase was added to the mixture,
and the mixture was incubated at 42˚C for 1 hour. Individual reaction mixtures were
incubated at 70˚C for 5 min to stop the extension reaction and treated with 5U of RNAse
H and 5U of RNAse A for 20 min at 37˚C. Each reaction mixture was brought up to a
volume of 200 µl with TE and concentrated using Nanosep 100 spin columns. The three
different cDNAs were collected from the membrane by washing with 20 µl of TE.
Aliquots (4 µl) of this mixture were used for second strand synthesis by PCR.
44
2.8 Overproduction of the rBcpA protein in E. coli
2.8.1 Generation of E. coli expression plasmids
Plasmids for the overproduction of proteins in E. coli were constructed using the pET
plasmid system (Studier et al., 1990). Appropriate restriction sites were engineered by
PCR into genes destined for overexpression. An NcoI or NdeI site was introduced at the
start codon of the gene via the design of the 5' oligonucleotide primer, and a BlpI site was
introduced downstream of the stop codon in the 3' oligonucleotide primer. The modified
and amplified PCR product was digested with appropriate restriction enzymes and ligated
into the equivalent sites of the chosen pET vectors, pET30C+. or pET16 The pET vectors
were purchased from Novagen (Madison, WI), and the conditions for overexpression
were according to those recommended by the manufacturer. Briefly, E. coli DH5-α cells
were transformed with the pETBcpA plasmids as described in 2.4.1 and kanamycin
resistant colonies were selected. Selected individual colonies picked were transferred to
5 ml of LB-kanamycin or LB-ampicillin liquid medium and grown overnight. These
isolated pETBcpA plasmids were used to transform either BL21R or BL21(DE3) E. coli
strains for recombinant protein overproduction.
45
2.8.2 Protein overproduction in E. coli and isolation of inclusion bodies
A frozen stock of pET30C+/BCP or pET16BCP was used to inoculate an LB-
kanamycin or LB-ampicillin plate to grow individual colonies harboring the
pET30C+/BCP or pET16BCP plasmid. The plates were allowed to grow overnight at
37˚C. An individual colony was picked from the plate and grown in either 5 ml of LB-
ampicillin or 5 ml of LB-kanamycin media (depending on the plasmid) at 37˚C to an
OD600 of 0.5. These 5 ml starter cultures were used to inoculate 500 ml or 1 L of LB-
kanamycin or LB-ampicillin liquid media and the cultures were incubated with rigorous
shaking at 37˚C until the OD600 was between 0.4 and 1.0. This usually took between 2-3
hours, after which, IPTG from 100 mM stock was added to a final concentration of 0.4
mM and the incubation was continued for 2-3 hours. The flasks were placed on ice for 5
min and the cells were harvested by centrifugation at 5000×g for 5 min at 4˚C. The cell
pellet was resuspended in 0.25× culture volume (125 ml) of TS (50 mM Tris, pH 8.0, 100
mM NaCl) and centrifuged again to wash the cells. The pellet was resuspended in 30 ml
of TS, and the cells were broken by using a French Pressure cell operated at 20,000 psi
and maintained at 4˚C. The broken cell mixture was centrifuged at 10,000×g in a
Beckman JA14 rotor to separate the inclusion bodies/membrane fraction from soluble
proteins for 5 min at 4˚C. The inclusion body pellet was resuspended in 3 ml of TS
buffer and centrifuged to wash the pellet. The pellet was washed twice with 0.1% (v/v)
Triton X-100 and washed again with TS buffer to remove Triton X-100. Next, the
46
inclusion body pellet was resuspended in 10 M urea (TS, 4 mM 2-mercaptoethanol) using
a minimal amount of buffer. This mixture was allowed to sit at room temperature for 1
hour and then centrifuged at 10,000×g in a Beckman JA14 rotor for 10 min to remove
insoluble material. The protein concentration was determined with the Pierce Coomassie
protein concentration assay. The solubilized pellet was then diluted with buffers as
described in the Results section to remove the chaotrope.
2.8.3 Purification of rBcpA from E. coli
The solubilized inclusion bodies were purified either by size exclusion
chromatography using a Sephadex G-75 resin (Amersham Pharmacia, Piscataway, NJ) or
a carboxymethyl (CM) sepharose resin (Amersham Pharmacia, Piscataway, NJ). For size
exclusion chromatography, the rBcpA protein was diluted in 0.8 M Urea TS buffer and
was loaded onto a pre-equilibrated G75 column. The running buffer was TS that had
been filter-sterilized and degassed. 2-mercaptoethanol was added to final concentration
of 4 mM to reduce disulfide bridges formed during the overproduction of the
recombinant protein in E. coli. The column was kept at 4˚C throughout the experiment.
Fractions were collected according to the 280 nm absorbance of the protein. The
fractions were dialyzed against TS after the column chromatography to remove the 2-
mercaptoethanol. For the purification of rBcpA using an ion exchange column, the
sample was diluted in 5PEM buffer (5 mM sodium phosphate, 1 mM EDTA, 2 mM 2-
mercaptoethanol) until the urea concentration was 0.8 M. The sample was loaded onto a
47
CM-sepharose column and the protein was eluted from the column at room temperature
by using a NaCl gradient (0-500 mM). The fractions collected were tested for protein
content with the Pierce BCA Protein Assay and were electrophoresed on SDS
polyacrylamide gels to determine the purity of the rBcpA protein.
2.9 Cytochrome c6 purification from Synechococcus sp. PCC 7002 and amino
terminal sequencing
20 L of Synechococcus sp. PCC 7002 cells were grown in A+ medium. Cells were
collected by centrifugation at 10,000×g, resuspended in 100 ml of Tris, pH 7.5 and
broken by passage through a French Press. The soluble fraction of the crude extract was
separated from the insoluble fraction by ultracentrifugation at 45000 × g for 1 hour. The
supernatant was collected and precipitated overnight at 4˚C with NH4SO4 (65%
saturation). The soluble fraction was separated by centrifugation at 8000 × g for 10 min
and the supernatant was collected. The soluble supernatant was fractionated again by an
overnight NH4SO4 (95% saturation) precipitation at 4˚C. The proteins were collected by
centrifugation at 10000 × g for 10 min. The pellet was resuspended in a minimal amount
of 20 mM Tris, pH 7.8 and deionized by dialysis in 20 mM Tris, pH 7.8 overnight at 4˚C.
Cytochrome c6 was separated from other proteins by FPLC using a linear gradient of 1M
NaCl in 50 mM Tris, pH 8.0 on a mono-Q column. Fractions from the mono-Q
chromatography were collected and cytochrome c6 was identified by its absorbance
properties. Fractions containing cytochrome c6 were pooled and precipitated by
48
trichloroacetic acid. The precipitated samples of cytochrome c6 were resuspended in
sodium dodecyl sulfate (SDS) sample buffer (60 mM Tris-HCl, pH 6.8, 2% (w/v) SDS,
10% (v/v) glycerol, 0.1 mg bromphenol blue per ml, 5% (v/v) 2-mercaptoethanol), and
loaded onto SDS polyacrylamide gradient gels (10-20%) and electrophoresed as
described below. Cytochrome c6 was identified by TMBZ staining (Thomas et al., 1976)
and Coomassie staining (see below). The protein band corresponding to cytochrome c6
was excised from the gel and electroeluted to PVDF membrane for N-terminal
sequencing at the University of Nebraska Protein Core Facility (Department of
Biochemistry, The Beadle Center, P.O. Box 880664, Lincoln, NE 68588-0664).
2.10 SDS-polyacrylamide gel electrophoresis, TMBZ staining and immunoblot
analysis
2.10.1 SDS-PAGE
Polyacrylamide gel electrophoresis in the presence of SDS was performed as
described by (Laemmli, 1970). Gels used in immunoblot studies and in protein
expression studies were 10%, 15 %, or 10-20% (w/v) linear gradient polyacrylamide slab
gels (30:0.8 (w/w) acrylamide:bisacrylamide). Stacking gels were typically 4% (w/v)
acrylamide. Protein samples were heated to 65˚C in SDS-sample buffer (60 mM Tris-
HCl, pH 6.8, 2% (w/v) SDS, 10% (v/v) glycerol, 0.1 mg bromphenol blue ml-1, 5% (v/v)
2-mercaptoethanol) for 5 min prior to loading. Typically the gels were electrophoresed
49
for 12-16 hours under constant amperage in a buffer containing 25 mM Tris, 0.2 M
glycine, 0.1% (w/v) SDS for 16 cm gels. Proteins separated by electrophoresis were
stained with Coomassie blue stain (0.125% (w/v) Coomassie Brilliant Blue, 50% (v/v)
methanol, 10% (v/v) acetic acid) and destained to visualize the protein by washing the gel
with destaining solution (50% (w/v) methanol, 10% (v/v) acetic acid) until the protein
bands were clearly visible.
2.10.2 3, 3', 5, 5'-tetramethylbenzidine (TMBZ) staining
Proteins with c-heme groups (such as cytochrome c6) can be specifically
visualized by 3, 3', 5, 5'-tetramethylbenzidine (TMBZ)-H2O2 staining (Thomas et al.,
1976). A 6.3 mM TMBZ (Sigma-Aldrich Chemical Co., St. Louis, MO) solution was
freshly prepared in methanol. Immediately before use, 3 parts of the TMBZ solution
were mixed with 7 parts of 0.25 M sodium acetate, pH 5.0. The gels were immersed in
this mixture at room temperature in the dark. After 1-2 hours with occasional mixing
(every 10-15 min), H2O2 was added to a final concentration of 30 mM. The staining was
visible within 3-30 min. The gels were washed with a (3:7) isopropanol:0.25 M sodium
acetate, pH 5.0 solution that was replaced once or twice to remove any precipitated
TMBZ.
50
2.10.3 Immunoblot analysis
The nitrocellulose membrane, filter papers, and SDS-polyacrylamide gel were
pre-soaked in transfer buffer (25mM Tris, 192 mM glycine (20% (v/v) methanol), pH
8.3) for 30 min. Proteins separated by SDS-PAGE were electrophoretically transferred
onto the nitrocellulose membranes (Schleicher & Schuell, Keene, NH) for 10-45 min at
10-15V using a BioRad Trans-Blot SD Semi-dry transfer cell (Richmond, CA). The
membrane was stained with 0.5% (w/v) Ponceau-S in 1.0% (v/v) acetic acid for 30
seconds to determine the transfer efficiency. The Ponceau-S stain was removed by
washing the membrane with several changes of distilled water. The membrane was then
placed in 5% (w/v) nonfat milk in TBS buffer (10 mM Tris-HCl, pH 8.0, 150 mM NaCl)
for 1 hour on a shaking platform at room temperature. Next, the membrane was
incubated in a 3% nonfat milk/TBS solution with the primary antibody at a dilution of
1:5000 to 1:10000 for 1-4 hours at room temperature. After incubation with the primary
antibody, the membrane was washed with TBST (10 mM Tris-HCl, pH 8.0, 150 mM
NaCl, 0.05% (v/v) Tween-20) buffer for 15 min, followed by two more washes with
TBST buffer of 5 min each. The membrane was then incubated with the secondary
antibody (goat, anti-rabbit IgG-alkaline phosphatase conjugate, Sigma, A-8025, St.
Louis, MO) diluted 1:3000 in 1% (w/v) nonfat milk/TBS for 2-3 hours. The membrane
was rinsed briefly with TBS buffer and then washed with TBST buffer once for 15 min,
followed by 4 washes in TBST buffer of 5 min each. The membrane was finally washed
51
with TBS buffer to remove the Tween-20. The color of the blot was developed in 50 ml
of alkaline phosphatase buffer (100 mM Tris-HCl, pH 9.5, 100 mM NaCl, 5 mM MgCl2)
containing 0.33 ml of nitroblue tetrazolium (50 mg ml-1 in 70% (v/v) dimethyl
formamide) and 0.165 ml of bromochlorindolyl phosphate (50 mg ml-1 in ddH2O). The
membrane was incubated in this solution at room temperature with agitation until bands
were suitably dark. The blot was rinsed with ddH2O to stop the reaction and allowed to
air dry. After color development was complete, the blots were rinsed in distilled water
and stored in the dark.
2.11 Determination of chlorophyll a and carotenoid contents
Chlorophyll a and total carotenoid concentrations were determined as described
by Sakamoto and Bryant (1998). Cyanobacterial cells (1.0 ml) in exponential growth
phase were harvested by centrifugation in 15 ml COREX tubes at 10,000×g at room
temperature for 5 min. The supernatant was removed and chlorophyll a and carotenoids
were extracted with dimethylformamide for 15 min at room temperature. The
concentrations of chlorophyll a and carotenoids in the dimethylformamide extract were
calculated from the following equations (A750nm was subtracted to correct for light
scattering):
[Chlorophyll a (µg ml-1)] = (A664nm-A750nm) × 11.92
[Carotenoids (µg ml-1)] = (A461nm-A750nm) - 0.046 × (A664nm-A750nm) × 4
52
2.12 Oxygen evolution and consumption rates
Oxygen evolution and consumption rates were determined using a Clark-type
electrode from Hansatech, Inc. (UK). The cyanobacterial cells were harvested by
centrifugation, washed with fresh A+ media, and centrifuged again. The cyanobacterial
cell pellets were resuspended to a final chlorophyll concentration of 3-5 µg chlorophyll a
per ml for oxygen evolution measurements. The cyanobacterial cell pellets were
resuspended to a final concentration of 50-100 µg of chlorophyll a per ml for oxygen
consumption measurements. The cells were kept in the dark 10-15 min and then agitated
to saturate the samples with air levels of oxygen prior to addition to the electrode
chamber. A total of 1 ml of cells was added to the electrode chamber for each individual
experiment. The electrode chamber temperature was maintained at 38˚C with a
circulating water bath, and the chamber contents were continuously stirred.
Measurements of oxygen evolution were determined after two min in the dark by
stimulation with saturating amounts of light (2.5 mE m-2 s-1). Measurements of oxygen
consumption were determined in complete darkness.
2.13 P700+ kinetic measurements
The photoinduced absorption change attributable to P700 can be monitored with a
single beam spectrophotometer (Maxwell and Biggins, 1976). Thus, the reduction
53
kinetics of P700+ in whole cells of Synechococcus sp. PCC 7002 were measured using a
detection system similar to the one described by (Yu et al., 1993) and originally by
(Maxwell and Biggins, 1976). Cyanobacterial cells were grown under standard growth
conditions (38˚C, 250 µE m-2 s-1, 1.5% (v/v) CO2/air) and harvested by centrifugation.
The cells were resuspended in 1 ml Ficoll saturated (200 mg ml-1) 25 mM Tris, pH 8.3
and adjusted to a chlorophyll a content of 40 µg ml -1. Modulated light was provided with
an oscillation frequency of 22 kHz, and the measuring beam was passed through a 5-mm-
wide sample cuvette with a 1-cm pathlength. A colored glass bandpass filter (λcuton > 800
nm) was inserted between the cuvette and the detector to minimize scattered actinic
illumination and fluorescence artifacts. The modulated near-IR measuring beam was
detected with a photodiode (RCA 30810), and the signal was demodulated with a lock-in
amplifier (model 5101; EG&G, Sunnyvale, CA). The instrument was operated with a
time constant of 1 ms. The sample was excited with white light generated from a 150-W
quartz-tungsten lamp that was focused on the cuvette (Yu et al., 1993). The cells were
kept in the dark prior to the experiment when they were subjected to a flash of saturating
light lasting 22 seconds to fully oxidize P700, followed by a dark incubation for 15
seconds to reduce P700+. The signal-to-noise ratio was improved by averaging 128 light-
dark cycles with a Nicolet 4095A digital oscilloscope (12-bit resolution, 500 µs point -1).
The first light dark cycle was recorded separately and compared with the average; no
changes in kinetics were found during the course of any measurement. The data were
transferred to and analyzed with IGORPRO v. 3.5 as described in (Yu et al., 1993). In all
cases, a single-exponential curve provided the best fit for the P700+ reduction kinetics.
54
2.14 77K fluorescence emission measurements
Exponentially growing cyanobacterial cells (10 ml) grown under various light
intensities (see Results) were harvested by centrifugation at room temperature at 8000×g
for 5 min and resuspended in 200 µl of 60% (v/v) glycerol, 50 mM HEPES, pH 7.0.
Samples were adjusted to a final OD730 nm of 1.0 in 1.0 ml of 60% glycerol, 50 mM
HEPES, pH 7.0. Measurement of 77K fluorescence was determined with an SLM-
Aminco 8000C spectrofluorometer (Spectronic Instruments Inc, Rochester, NY, USA);
the excitation wavelength was 440 nm for chlorophyll excitation. Absorption spectra
were obtained with a Cary-14R spectrophotometer modified for computerized operation,
data collection and data analysis by On-Line Instruments Systems (Bogart, GA, USA) as
described by (Sakamoto and Bryant, 1998).
2.15 Pulse Amplitude Modulated (PAM) fluorescence measurements
In PAM fluorescence measurements, a constant, weak modulated light source is
combined with a system that selectively detects the fluorescence emitted at the same
frequency and phase as the modulated source (Schreiber et al., 1986). Because the
modulated source is of constant irradiance, one can measure the modulated fluorescence
signal once the unmodulated background irradiance is high enough to close all of the
reaction centers (Geider and Osborne, 1992). Therefore, one can use PAM fluorescence
55
to determine the number of functional PSII complexes in whole cells. Exponentially
growing cyanobacterial cells (25 ml) grown under standard growth conditions (38°C, 250
µE m -2 s-1, 1.5% (v/v) CO2) were harvested by centrifugation at room temperature at
8000×g for 5 min. The cell pellet was resuspended and washed with 25 mM HEPES-
NaOH buffer, pH 7.0. The cells were collected again by centrifugation and resuspended
to a final OD550 nm = 1.0. Cells (1-3 ml) at a chlorophyll concentration of 5 µg ml -1 in
25mM HEPES-NaOH buffer, pH 7.0 were added to the cuvette. Sodium bicarbonate (1.0
mM final concentration) was added to the cuvette as well. The cells were incubated for 2
min in complete darkness before establishing F0, or the ground state. FM’, or the maximal
fluorescence, was obtained by exposing the cyanobacterial cells to a saturating light pulse
for 1 second intervals. The fluorometer settings were changed to 100 kHz from 1,600
kHz to obtain FV’, or variable fluorescence. After establishing Fm’, DCMU was added to
1 µM in order to obtain FM. by closing all PSII reaction centers completely. By
examining the ratio of FV’/FM’ (variable fluorescence to maximal fluorescence) one can
determine the total number of active PSII clusters compared to total chlorophyll content.
2.16 Superoxide dismutase (SOD) activity measurements
SOD inhibits the reduction of nitro-blue tetrazolium by superoxide by dismutation
of superoxide to hydrogen peroxide and oxygen (Winterbourn et al., 1975). This allows
one to measure the amount of SOD activity in soluble and membrane fractions of
Synechococcus sp. PCC 7002. Cyanobacterial cells (150 ml) were collected by
56
centrifugation and washed with 50 mM potassium phosphate buffer, pH 7.8. The
recollected cells were resuspended in 20 ml of 50 mM potassium phosphate buffer and
broken by passage through an SLM-AMINCO French Press. The broken cells were
centrifuged at 4°C, 5000×g for 5 min to collect unbroken cells. The supernatant was
collected and subjected to a high-speed centrifugation (45,000×g for 1 h) at 4°C to
separate the soluble and membrane fractions. The soluble fraction was collected and
immediately used in SOD assays. The membrane fraction was collected and resuspended
in a minimal amount of 50 mM potassium phosphate buffer, pH 7.8 and used for SOD
assays. The protein concentration for each fraction was determined by Coomassie assay.
SOD activity assays were performed according to the protocol of (Winterbourn et al.,
1975).
2.17 Detection of hydroperoxides
A modified ferrous oxidation/xylenol orange assay was used to determine the
levels of hydroperoxides in cyanobacterial cells (Sakamoto and Bryant, 1998).
Exponentially growing cyanobacterial cells were first diluted to an OD550nm of 0.2 and
were incubated under standard growth conditions (250 µE m-2s-1 at 38°C) or in darkness
for 4 hours in the presence or absence of 50 µM methyl viologen. The cells were
harvested by centrifugation at 5000×g for 10 min. The cell pellets were resuspended in
0.8 ml of methanol/0.01% (w/v) BHT, 0.1 ml of Reagent A (2.5 mM ammonium iron (II)
sulfate/0.25M sulfuric acid) and 0.1 ml of Reagent B (40 mM BHT/1.25 mM xylenol
57
orange in methanol) to an OD550nm of 1.0 per ml. The mixture was incubated for 30 min
at room temperature and then centrifuged at 10,000×g to remove any cell debris. The
absorbance at 560 nm was determined for the samples and the concentration of
hydroperoxides was determined by using the extinction coefficient (E560nm = 4.3X104 M -1
cm-1) (Sakamoto and Bryant, 1998).
2.18 Catalase activity
To measure catalase activity in Synechococcus sp. PCC 7002, the method of
(Beers and Sizer, 1952) was followed using whole cells. Exponentially growing
cyanobacterial cells (25 ml) grown under standard growth conditions (38°C, 250 µE m -2
s-1, 1.5% (v/v) CO2) were harvested by centrifugation at room temperature at 8000×g for
5 min. The cell pellet was resuspended and washed with 50 mM potassium phosphate
buffer, pH 7.0. The washed cells were again collected by centrifugation and resuspended
to give a final OD550nm = 1.0 ml-1 in a 3 ml sample. Catalase activity was measured by
monitoring the rate of H2O2 decomposition at 240 nm by using the extinction coefficient
for H2O2 of 43.6 M-1cm-1 (Beers and Sizer, 1952).
2.19 Peroxidase activity
Peroxidase activity can be measured by the following reaction: H2O2 + peroxidase
+ oxygen acceptor (colorless)àH2O + oxidized acceptor (colored) (Trinder, 1966). In
58
this case the oxygen acceptor is 4-aminoantipyrine in the presence of phenol.
Exponentially growing cyanobacterial cells (25 ml) grown under standard growth
conditions (38°C, 250 µE m -2 s-1, 1.5% CO2) were harvested by centrifugation at room
temperature at 8000× g for 5 min. The cell pellet was resuspended and washed with 20
mM potassium phosphate buffer, pH 7.0. The washed cells were again collected by
centrifugation and resuspended to give a final OD550nm of 2.0 per ml in a 3.0 ml sample.
Peroxidase activity was measured by monitoring the change in color of phenol in the
presence of 4-aminoantipyrene and H2O2 at 510 nm (Trinder, 1966).
2.20 Cell viability
Exponentially growing cells (OD550nm = 0.25) from the wild-type, ctaDI-, ctaDII -,
and ctaDI- ctaDII- strains of Synechococcus sp. strain PCC 7002 were divided into four
culture tubes. Two tubes of the cultures of each strain were incubated under standard
growth conditions with or without the addition of 50 µM MV. The other two tubes of the
cultures of each strain were incubated in the dark at 37°C with or without the addition of
50 µM MV for 4 hours. The cells were harvested by centrifugation and washed with
fresh A+ liquid media to remove the methyl viologen. The cells were resuspended in A+
to give a final OD550nm of 0.1 per ml. This culture was diluted 1000-fold and 10 µl of
each dilute, resuspended cell culture was spread on an appropriate A+ plate. The cells
were grown on plates for seven days prior to counting of colony forming units by visual
inspection. Under normal growth conditions at 38°C, 1 ml of Synechococcus sp. PCC
59
7002 culture with an OD550nm = 1.0, is equivalent to 4.7 + 0.6 × 107 CFU (Sakamoto and
Bryant, 1998).
60
Chapter 3
RESULTS
Cloning of electron transport protein genes
In order to determine the number and type of electron transport protein encoding
genes in Synechococcus sp. PCC 7002, forty open reading frames, putatively encoding
proteins involved in electron transport based on sequence similarities to known electron
transport proteins, were cloned and sequenced from the marine cyanobacterium
Synechococcus sp. PCC 7002. The ndh and ndb genes were cloned by Søren Persson, a
lab technician who worked under my direct supervision. All of the clones were used to
compare the number of electron transport genes and organization of electron transport
genes to those of other cyanobacteria. The genes identified in this study can be divided
into five major groups according to their presumed products: (1) the type I NADH
dehydrogenase, (2) the type II NADH dehydrogenases, (3) the hydrogenase and its
assembly proteins, (4) mobile electron transport proteins, and (5) cytochrome oxidases.
Table 3 shows the genes and some predicted properties of the proteins encoded by those
genes identified or used in this study. The percent similarity and identity of the
Synechococcus sp. PCC 7002 electron transport proteins to their respective homologs
from other bacteria or chloroplasts are shown in Table 4. Restriction maps of all of the
61
Synechococcus sp. PCC 7002 electron transport genes identified in this study are
shown in APPENDIX A . ClustalW protein alignments of the electron transfer proteins
from this study with their respective homologs from other bacteria and chloroplasts not
shown in the RESULTS are located in APPENDIX B.
62
Table 3. Genes that encode electron transport proteins from Synechococcus sp. PCC
7002 and some predicted properties of those proteins.
63
Gene / gene clusterSynechococcus sp. PCC7002
Synechocystissp. PCC6803
homolog
Source Aminoacids
MW (kDa)
PredictedpI
Type I NADH dehydrogenasendhB sll0223 this study 527 56.1 5.28ndhC
ndhK/psbGndhJ
slr1279slr1280slr1281
this studythis studythis study
121242174
13.526.919.9
6.545.714.22
ndhD1 slr0331 this study 527 57.6 6.83ndhD2 slr1291 this study 532 58.0 9.12ndhF3 sll1732 Klughammer et al., 1999 617 67.2 6.35ndhD3 sll1733 Klughammer et al., 1999 499 53.9 8.16ndhF4 sll0026 this study 633 69.3 6.25ndhD4 sll0027 this study 494 53.5 7.43ndhF1 slr0844 Schluchter et al., 1993 665 72.9 5.27ndhF5 slr2007 this study 479 52.2 9.25ndhF2 slr2009 this study 479 52.1 9.24ndhH slr0261 this study 395 45.6 5.66ndhAndhIndhGndhE
sll0519sll0520sll0521sll0522
this studythis studythis studythis study
372203206104
40.523.421.911.4
4.825.654.924.95
ndhL ssr1386 this study 77 9.2 9.42Type II NADH dehydrogenase
ndbA slr0851 this study 460 50.6 5.82ndbB slr1743 this study 391 43.3 5.61
Cytochrome oxidasectaCIctaDIctaEI
slr1136slr1137slr1138
this studythis studythis study
362590197
39.660.922.0
4.555.625.96
ctaCIIctaDIIctaEII
sll0813sll2082sll2083
this studythis studythis study
298551198
33.861.022.4
6.506.374.89
HydrogenasehypEhoxEhoxFhoxUhoxYhyp3hoxHhoxWhypAhypBhypFhypChypD
sll1464sll1220sll1221sll1223sll1224
NAsll1226slr1876slr1675sll1079sll0322ssl3580slr1498
this studythis studythis studythis studythis studythis studythis studythis studythis studythis studythis studythis studythis study
34817153623818920947614311427476680362
36.918.457.926.120.923.352.916.212.030.486.08.2
39.3
4.806.344.785.734.665.696.624.594.406.217.184.046.71
Mobile electron transport proteinsbcpA NA this study 106 10.6 6.06petJ1 sll1796 Nomura and Bryant 1997 112 9.4 4.89petJ2 NA this study 88 9.4 9.29cytM sll1245 this study 98 10.5 8.53
NA: not available, since no homolog occurs in Synechocystis sp. PCC 6803
64
Table 4. Percent identity and similarity of Synechococcus sp. PCC 7002 electron
transport proteins with homologs from other bacteria and the spinach chloroplast.
X/Y in table represents the % identity / % similarity of the predicted protein from
Synechococcus sp. PCC 7002 compared to Synechocystis sp. PCC 6803, Anabaena
sp. PCC 7120, E. coli, Spinacia oleracea, and R. eutropha.
ND: not determined.
(*) compared to sll1796 (PetJ) from Synechocystis sp. PCC 6803.
N-317 refers to the 317 amino acids at the amino terminus of the E. coli NuoF
protein.
C-338 refers to the 338 amino acids at the carboxy-terminus of the R. eutropha HypF
protein.
65
Predicted ProteinSynechococcus sp.
PCC7002
Synechocystissp. PCC6803
Anabaena sp.PCC7120
E. coli S. oleracea R. eutropha
Type I NADH dehydrogenase
NdhB 73/85 72/83 27/45 46/64 NDNdhC
NdhK/PsbGNdhJ
91/9780/8674/84
87/9573/8575/84
27/4634/5223/43
65/8150/6448/61
NDNDND
NdhD1 80/87 80/88 33/51 45/62 NDNdhD2 57/70 55/69 31/48 42/58 NDNdhD3 65/78 59/76 30/51 27/38 NDNdhD4 64/78 33/52 33/53 29/49 NDNdhF1 76/85 73/83 33/50 44/68 NDNdhF2 49/68 33/52 14/30 15/28 NDNdhF3 64/78 57/76 24/43 23/39 NDNdhF4 64/81 58/74 33/53 17/31 NDNdhF5 54/70 62/78 15/31 16/33 NDNdhH 92/95 85/92 37/55 71/83 NDNdhANdhINdhGNdhE
84/8481/8866/8180/92
75/8781/8762/7481/88
33/5421/3522/4229/56
58/7854/6436/5358/80
NDNDNDND
NdhL 58/71 51/65 ND ND NDType II NADH
dehydrogenaseNdbA 61/74 57/69 19/32 ND NDNdbB 53//74 50/68 20/31 ND ND
Cytochrome oxidaseCtaCICtaDICtaEICtaCIICtaDIICtaEII
61/7480/9067/7840/5764/7656/69
47/6563/7855/7451/7072/8661/71
NDNDNDNDNDND
NDNDNDNDNDND
NDNDNDNDNDND
HydrogenaseHypEHoxEHoxFHoxUHoxYHyp3HoxHHoxWHypAHypBHypFHypCHypD
57/7367/8269/8375/8660/75ND
66/8936/5150/7051/6448/6352/7261/75
ND56/7365/7968/8548/6446/6564/8030/4630/5136/57NDND
49/68
42/5624/4430/47
24/36(N-317)13/23ND
18/33ND
29/5237/5739/5632/4745/65
NDNDNDNDNDNDNDNDNDNDNDNDND
43/61ND
28/4532/4930/50ND
39/5713/26ND
30/4330/44 (C-388)
34/5043/62
Mobile electron transport proteins
BcpA ND 62/75 ND ND NDPetJ1 46/64 46/56 ND ND NDPetJ2 37/51* 47/59 ND ND NDCytM 49/65 ND ND ND ND
66
3.1 Type I NADH Dehydrogenase
A full complement of genes that are predicted to encode subunits of the type I
NADH dehydrogenase and that exhibit strong sequence similarity to the genes found in
the freshwater cyanobacterium Synechocystis sp. PCC 6803 were isolated from the
marine cyanobacterium Synechococcus sp. PCC 7002. These include the ndhA, ndhI,
ndhG, ndhE, ndhB, ndhC, ndhK, ndhJ, ndhD1, ndhD2, ndhD3, ndhD4, ndhF1, ndhF2,
ndhF3, ndhF4, ndhF5, ndhH, and ndhL genes. The NADH dehydrogenase genes were
cloned and the predicted protein sequences were used in comparisons with other
cyanobacterial subunits of the NADH dehydrogenase as well as with their respective
counterparts found in the chloroplast of Spinacia oleracea and in the gram-negative
bacterium E. coli. As in chloroplasts and other cyanobacteria, Synechococcus sp. PCC
7002 is missing the homologs to the nuoE, nuoF and nuoG genes of E. coli and R.
sphaeroides. Restriction maps for all Synechococcus sp. PCC 7002 ndh genes are shown
in APPENDIX A.
67
3.1.1 ndhAIGE
The ndhAIGE gene cluster was isolated on a 3.8-kb EcoRI fragment from the
Synechococcus sp. PCC 7002 EcoRI cosmid library. Figure 3 reveals that the ndhAIGE
gene cluster arrangement is nearly identical to that of the freshwater cyanobacterium,
Synechocystis sp. PCC 6803 and the gene cluster arrangement of the filamentous,
nitrogen-fixing cyanobacterium, Anabaena sp. PCC 7120. This ndhAIGE gene
organization may be conserved throughout all cyanobacteria. However, the genes
flanking the ndhAIGE gene cluster are very different from one another. In
Synechococcus sp. PCC 7002, the ndhAIGE gene cluster is flanked by a homolog to the
fraH gene, which would encode a putative zinc-finger protein related to the slr0298 open
reading frame of Synechocystis sp. PCC 6803 at the 5’ end of the cluster. Examination of
the 3’ end of the cluster revealed that it is homologous to the slr1536 open reading frame
from Synechocystis sp. PCC 6803. Both of these reading frames are in the same
transcriptional orientation as the ndhAIGE gene cluster. In Synechocystis sp. PCC 6803
the genes flanking the ndhAIGE gene cluster are transcribed divergently and consist of a
valyl-tRNA and a gene encoding a putative dihydroorotate dehydrogenase (Kaneko et al.,
1996). The Anabaena sp. PCC 7120 ndhAIGE gene cluster is flanked by a predicted
citrate synthase gene as well as a homolog to the hypothetical protein slr1415 (Kazusa
DNA Research Institute, 2000; Kaneko et al., 1996). The similarities of the ndhAIGE
gene clusters and differences in the genes flanking the ndhAIGE gene cluster illustrate
68
Figure 3. Gene organization of the ndhAIGE gene cluster in cyanobacteria. Arrow
directions indicate the direction of transcription. Scale is as indicated in bp for each
individual map. (A) The gene organization of the ndhAIGE gene cluster of
Synechococcus sp. PCC 7002. (B) The gene organization of the ndhAIGE gene cluster of
Synechocystis sp. PCC 6803. (C) The gene organization of the ndhAIGE gene cluster of
Anabaena sp. PCC 7120.
Synechococcus sp. PCC7002
Synechocystis sp. PCC6803
Anabaena sp. PCC7120
1K 2K 3K 4K 5K
ndhA sll1415citrate synthase ndhGndhI ndhE
1K 2K 3K 4K 5K 6K 7K
Valyl-T-RNA synthetaseDihydroorotate dehydrogenase (slr1418) ndhA ndhGndhIslr1419 ndhE
1K 2K 3K
ndhA ndhGndhI ndhEfraH (slr0298) slr1536
ndhAIGE gene organization in cyanobacteria
69
both the conservation of gene organization as well as the diversity in gene
organization among these three cyanobacterial species.
The predicted proteins encoded by the Synechococcus sp. PCC 7002 ndhAIGE
gene cluster are most similar to their homologs from the freshwater cyanobacterium
Synechocystis sp. PCC 6803 (Table 4), and have the least amount of similarity with the
homologs from E. coli.
Sequence analysis indicates that the ndhA gene is predicted to encode a protein of
372 amino acids in Synechococcus sp. PCC 7002 with a predicted pI of 4.82. In E. coli,
the NdhA homolog NuoH contains the ubiquinone binding site (Yagi et al., 1998). Based
on the amino acid sequence similarity of the Synechococcus sp. PCC 7002 NdhA protein
to other NdhA and ND1 proteins and the similarity of the NdhA and ND1 hydrophobicity
profiles, the NdhA subunit may also contain a quinone-binding site (Ellersiek and
Steinmüller, 1992; Friedrich et al., 1990; Matsubayashi et al., 1987).
The ndhI gene is predicted to encode a protein of 203 amino acids with a
predicted pI of 5.65. NdhI is the only protein in the predicted membrane arm of the
NDH-1 complex that would have iron-sulfur clusters. These clusters are bound to the
NdhI protein in a similar manner to the 4Fe-4S clusters of bacterial ferredoxins. The
consensus-binding motif CysXXCysXXCysXXXCysPro, which provides ligands for
tetranuclear iron-sulfur centers in ferredoxins, occurs twice in the predicted NdhI
sequence of Synechococcus sp. PCC 7002. The first occurrence of the motif begins at
Cys-92 for the amino acid sequence CIACEVCVRVCP and the second begins at Cys-137
70
for the amino acid sequence CIFCGNCVEYCP. The NdhI protein is homologous to
the E. coli NuoI protein that most likely contains two 4Fe-4S clusters, as well.
The ndhG gene putatively encodes a protein of 206 amino acids and a pI of 4.92
in Synechococcus sp. PCC 7002. The NdhG protein is homologous to the ND6 protein
from the mitochondrial complex (Walker et al., 1992) and the NuoJ protein of E. coli
(Weidner et al., 1993). Analyses of NdhG predict a protein with 5 transmembrane
helices and suggest that this gene encodes part of the membrane bound arm of the NDH-1
complex.
The ndhE gene of Synechococcus sp. PCC 7002 is the last gene found in this gene
cluster. Analysis predicts a protein of 104 amino acids with a pI of 4.95. The NdhE
protein is homologous to the NuoK protein from E. coli and the ND4L protein from the
mitochondrial NDH-1 complex (Friedrich et al., 1995).
3.1.2 ndhB
A portion of the ndhB gene from Synechococcus sp. PCC 7002 was initially
isolated on a 0.9-kb BamHI-EcoRI fragment, and the gene was completely sequenced by
adding sequence from a cosmid 4A-B3 isolated from the EcoRI cosmid library. The
ndhB gene of Synechococcus sp. PCC 7002 is predicted to encode a protein of 527 amino
acids with a pI of 5.28. The NdhB protein is homologous to the NuoN protein in E. coli
and the ND2 protein from the mitochondrial NDH-1 complex.
71
The gene organization of the Synechococcus sp. PCC 7002 ndhB gene is
similar to that of Synechocystis sp. PCC 6803 since the gene in both organisms is not
associated with any other ndh gene. Figure 4 shows that the ndhB gene from
Synechococcus sp. PCC 7002 is adjacent to an open reading frame predicted to encode a
2Fe-2S ferredoxin-like protein similar to one found in Clostridium pasteurianum
(Fujinaga and Meyer, 1993). These two genes are upstream of an open reading frame
predicted to encode an adenine phosphoribosyltransferase related to the sll1430 open
reading frame from Synechocystis sp. PCC 6803 (Kaneko et al., 1996).
72
Figure 4. Gene organization around the ndhB gene in cyanobacteria. Gene maps of
three different cyanobacteria: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. Arrows represent transcriptional direction. Maps are
individually scaled in bp.
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300
ndhB (sll0223)phoA (sll0222) ssl0410 psaK (ssr0390)
300 600 900 1200 1500 1800 2100 2400 2700
ndhB Ferredoxin sll1430
300 600 900 1200 1500 1800 2100 2400 2700 3000
ndhB PAB1763hstK
7120 ndhB
6803 ndhB
7002 ndhB
73
3.1.2.1 Attempted mutagenesis of the Synechococcus sp. PCC 7002 ndhB gene by
interposon mutagenesis
The inactivation of the ndhB gene in Synechococcus sp. PCC 7002 was attempted
for two reasons: (1) The ndhB genes in Synechocystis sp. PCC 6803 (Ogawa, 1991) and
Synechococcus sp. PCC 7942 (Marco et al., 1993) were previously insertionally
inactivated. Analysis of these ndhB mutants revealed strains that required much higher
concentrations of CO2 for growth. However, in tobacco, mutagenesis of the ndhB gene
revealed impairment of cyclic electron flow around PSI (Shikanai et al., 1998). In order
to determine whether the ndhB gene product from Synechococcus sp. PCC 7002 was
involved in carbon uptake, cyclic electron flow or both, inactivation of the ndhB gene
was attempted. (2) It had been previously demonstrated by Schluchter et al. (1993) that
inactivation of ndhF1 of Synechococcus sp. PCC 7002 had an effect on cyclic electron
flow. It had also been demonstrated by Klughammer et al. (1998) that inactivation of the
ndhF3 and ndhD3 genes in Synechococcus sp. PCC 7002 lead to phenotypes affecting
carbon assimilation. Both of these studies were done with genes (ndhF1, ndhF3 and
ndhD3) that are present in multiple copies in the genome of Synechococcus sp. PCC 7002
(Table 3). On the other hand, ndhB is present as a single copy gene in Synechococcus sp.
PCC 7002 and its inactivation may lead to phenotypes different than those observed for
genes present in multiple copies. Therefore, to examine the effects of inactivation of a
74
single copy ndh gene in Synechococcus sp. PCC 7002, mutagenesis of the ndhB gene
was attempted.
The ndhB gene was interrupted by the insertion of a Klenow-filled-in HincII-
EcoRV 1.3-kb fragment containing the chloramphenicol acetyltransferase (cat) gene
isolated from the plasmid pRL425 (Elhai and Wolk, 1988) into a unique BglII site within
the ndhB coding sequence which was made blunt by Klenow treatment (see Figures 4
and 5). Several hundred independently isolated chloramphenicol/ampicillin E. coli
colonies were screened to isolate the mutant plasmid. Two plasmids were isolated for
both the parallel and anti-parallel orientation of the antibiotic resistance cartridge relative
to the reading frame orientation of the cloned ndhB sequence. The resultant plasmids,
pNDHB1 (parallel transcriptional orientation of the cat insert relative to ndhB) and
pNDHB2 (antiparallel transcriptional orientation of the cat insert relative to ndhB) were
used to transform the wild-type strain of Synechococcus sp. PCC 7002 (Figure 5A).
Transformants were selected for as described in Materials and Methods.
Chloramphenicol-resistant transformants were selected and grown in liquid culture. The
segregation of the ndhB::cat locus in the chloramphenicol resistant transformants was
checked by Southern Blot analysis. All of the transformants were merodiploid for the
ndhB::cat and the wild-type copy of the ndhB gene (Figure 5B). The merodiploid
ndhB::cat/wild-type ndhB strains of Synechococcus sp. PCC 7002 were grown on
selective medium A+ plates under high CO2 concentrations (5% CO2, 95% air) to try and
force full segregation of the alleles, since ndhB- strains of Synechocystis sp. PCC 6803
(Ogawa, 1991) and Synechococcus sp. PCC 7942 (Marco et al., 1993) require higher
75
levels of CO2 for viability. This treatment failed to cause segregation of the alleles.
All subsequent attempts to segregate the mutant (increasing the antibiotic concentration
or increasing of CO2 levels) also failed to segregate the ndhB loci. These results imply
that the ndhB gene is essential to Synechococcus sp. PCC 7002 under all of the conditions
that were tested for segregation. The inability to segregate the ndhB mutant and wild-
type alleles may be indicative of the central role of the type I NADH dehydrogenase in
Synechococcus sp. PCC 7002 compared to the enzymes function in other cyanobacterial
species.
76
Figure 5. The ndhB gene of Synechococcus sp. PCC 7002 is required for cell viability.
(A) The ndhB gene was interrupted by insertion of the chloramphenicol acetyltransferase
(cat) gene into a unique BglII site within the coding sequence. The cat gene was digested
with HincII and EcoRV. The ndhB plasmid was digested with BglII. This site was made
blunt via a Klenow fill-in reaction and the cat gene was ligated into the now blunt BglII
site. Transformants containing both orientations of the cat gene were isolated, and the
new plasmids pNDHB(1.1), corresponding to the parallel orientation of insert and
pNDHB(2.2), corresponding to the antiparallel orientation of insert. These plasmids were
used to transform the wild-type strain of Synechococcus sp. PCC 7002. (B) Southern blot
hybridization analysis of the ndhB loci. Genomic DNA was digested with EcoRI,
transferred to a nylon membrane, and probed with a 32P-labeled PCR product for ndhB.
The sizes of the hybridizing bands are indicated on the left. Lane 1 represents genomic
DNA isolated from the merodiploid wild-type ndhB/ndhB::cat strain grown under normal
growth conditions. Lane 2 represents genomic DNA isolated from the merodiploid wild-
type ndhB/ndhB::cat strain grown under 5% CO2/95% air. Lane 3 represents genomic
DNA isolated from the merodiploid wild-type ndhB/ndhB::cat strain grown in the
presence of 10 mM glycerol under normal growth conditions. Lane 4 represents genomic
DNA isolated from the merodiploid wild-type ndhB/ndhB::cat strain grown under 5%
CO2/95% air and 10 mM glycerol. Lane 5 represents genomic DNA isolated from the
wild-type strain of Synechococcus sp. PCC 7002.
77
1 2 3 4 5
2.1 kb
0.89 kb
B.
A.
ndhB
CATCAT
ndhB
Bam
HI
Eco
RI
Bam
HI
Eco
RI
BglII
Eco
RI
Eco
RI
Eco
RV
Eco
RV
Hin
dIII
Hin
dIII
Hin
cII
Hin
cII
BglII
78
3.1.3 ndhD1, ndhD2, ndhD3, ndhD4
Synechococcus sp. PCC 7002 has 4 genes that putatively encode proteins with
strong sequence similarity to NdhD. There are an equivalent number of ndhD genes
found in the genome of Synechocystis sp. PCC 6803 (Kaneko et al., 1996). NdhD is
homologous to the NuoM protein of E. coli and to the ND4 protein from the
mitochondrial NDH-1 complex (Friedrich et al., 1995). Non-cyanobacteria have only
one gene copy of the ndhD-like genes; thus they lack the diversity displayed by the
multiple copies of the ndhD-type genes seen in cyanobacteria (Kaneko et al., 1996). All
of the ndhD genes are predicted to encode highly hydrophobic subunits that are most
likely membrane bound. Restriction maps of all Synechococcus sp. PCC 7002 ndhD
genes are shown in APPENDIX A.
The ndhD1 gene (slr0331 homolog) was cloned on 2.2-kb BamHI-HincII
fragment and sequence analysis is predicted to encode a protein of 521 amino acids with
a pI of 6.83. NdhD1 from Synechococcus sp. PCC 7002 is most similar to NdhD1 gene
from Synechocystis sp. PCC 6803.
The ndhD2 gene (slr1291 homolog) was isolated on a 3.5-kb-XbaI fragment. The
analysis of the ndhD2 nucleotide sequence is predicted to encode a protein of 532 amino
acids with a pI of 9.12. NdhD2 from Synechococcus sp. PCC 7002 is most similar to
NdhD2 gene from Synechocystis sp. PCC 6803.
79
The ndhD3 gene (sll1773 homolog, accession number U97516) was cloned by
Klughammer et al. (Klughammer et al., 1999), and the gene is predicted to encode a
protein of 499 amino acids with a pI of 8.16. NdhD3 from Synechococcus sp. PCC 7002
is most similar to NdhD3 gene from Synechocystis sp. PCC 6803.
The ndhD4 gene (sll0027 homolog) from Synechococcus sp. PCC 7002 was
cloned and sequenced from two independently isolated clones. The first plasmid consists
of a 2.6-kb EcoRV-EcoRI fragment, which contains the 3' end of the ndhF4 gene and the
5' end of the ndhD4 gene. A second clone consisting of a 2.5-kb HincII-HindIII fragment
contains the 3' end of the ndhD4 gene. The ndhD4 gene is predicted to encode a protein
of 494 amino acids with a pI of 7.43.
Figure 6 shows the ClustalW amino acid alignment of the four Synechococcus sp.
PCC 7002 NdhD proteins. Table 5 shows percent identity and similarity of the four
Synechococcus sp. PCC 7002 NdhD proteins to each other. It is clear from these results
that, although the NdhD proteins from Synechococcus sp. PCC 7002 are related to one
another, they are very diverse in size, homology, and pI. The differences observed in the
NdhD subunits may be responsible for the diversity in the number and types of NDH-1
complexes in cyanobacteria.
80
Figure 6. ClustalW alignment of the four NdhD amino acid sequences from
Synechococcus sp. PCC 7002. The Synechococcus sp. PCC 7002 NdhD amino acid
sequences were aligned using the ClustalW alignment program from MacVector v. 6.5.
The consensus amino acid sequence is underneath the alignments. Gray high-lighted
regions indicate identical amino acids and lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids were determined with the
ClustalW program (Thompson et al., 1994b).
81
7002 NdhD ClustalW Formatted Alignments
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
10 20 30 40 50
M N F A N F P WL S T I I L F P I I A A L F L P L I P D K D G K T V R WY A L T I G L I D F V I I VM D S L Q I P WL T T A I A F P L L A A L V I P L I P D K E G K T I R WY T L WR C P H R F C L L V
M L S F L L F L P L V G I G A I A L F P R - - - - P L T R I V A T V F T V V T L A I SM L S A L I WL P L A G A L L V A I L P Q G E K N Q F S R T M A L G A A A L V F V WT
L S . I P L . . A L . . L . P . . . . . . .
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
60 70 80 90 100
T A F Y T G Y D F G N P N L Q L V E S Y T WV E A I D L R WS V G A D G L S M P L I L L T G F I T TT A F WQ N Y D F G R T E F Q L T K N F A WI P Q L G L N WS L G V D G L S M P L I I L A T L I T TS G L L I N L N L Q D A G M Q Y T E F H N WL S I L G L N Y N L G V D G L S L P L I V L N S L L T LA WL G F H Y D V A I A G L Q F V E H Y L WI E WL G L N Y D L G V D G L S L P L L A L N A L L T L. . Y D . Q E . W. L G L N . L G V D G L S P L I . L L . T
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
110 120 130 140 150
L A I L A A WP V S F K P K L F Y F L M L L M Y G G Q I A V F A V Q D M L L F F F T WE L E L V P VL A T L A A WN V T K K P K L F A G L I L V M L S A Q I G V F A V Q D L L L F F I M WE L E L V P VV A I Y S I G E S N H R P K L Y Y S L I L L I N S G I T G A L I A N N L L L F F L F Y E I E L I P FV A L WI S P K D L H R P R F Y Y A L F L L L Q A S V N G A F L A Q D V L L F F L F Y E I E I I P L. A . . . . . P K L . Y L L L . G . F . . Q D . L L F F . . E . E L . P .
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
160 170 180 190 200
Y L I L S I WG G K K R L Y A A T K F I L Y T A G G S L F I L I A A L T M A F Y G D T V T F D M T AY L L I S I WG G K K R L Y A A T K F I L Y T A L G S V F I L A F T L A L A F Y G G D V T F D M Q AY L L I A I WG G E K K G Y A S T K F L I Y T A I S G L C V L A A F L G I V WL S Q S S N F D F E NY F L I A I WG G K K R G Y A A I K F L L Y T A V S G I L I L A S F L G L A F L T E S N T F A Y S AY L L I I WG G K K R . Y A A T K F . L Y T A . . I L A L . . A F . T F D A
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
210 220 230 240 250
I A Q K D F G I N L Q L L L Y G G L L I A Y G V K L P I F P L H T WL P D A H G E A T A P A H M L LL G L K D Y P L A L E L L A Y A G F L I G F G V K L P I F P L H S WL P D A H S E A S A P V S M I LL T L E N L E F N T K V I L L T I L L I G F G I K I P L V P L H T WL P D A Y V E A N P A V T V L LL H S D L L P L T T Q L I L L G G I L V G F G I K I P F L P F H T WL P D A H V E A S T P V S V I LL . L . L . G . L I G F G . K . P . P L H T WL P D A H . E A . P V . . L
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
260 270 280 290 300
A G I L L K M G G Y A L L R M N A G M L P D A H A L F G P V L V I L G V V N I V Y A A L T S F A Q RA G V L L K M G G Y G L I R L N M E M L P D A H I R F A P L L I V L G I V N I V Y G A L T A F G Q TG G V F A K L G T Y G L V R F G L Q L F P D V WS T V S P A L A V I G T V S V M Y G S L A A I A Q RA G V L L K V G T Y G L L K F G I G L F P L A WA V V A P WL A I WA A I S A L Y G A S C A I A Q KA G V L L K G Y G L . R . P D A . . P . L . . . G . V . . Y G A L A A Q .
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
310 320 330 340 350
N L K R K I A Y S S I S H M G F V L I G M A S F T D L G T S G A M L Q M I S H G L I G A S L F F M VN L K R R L A S S S I F P H G L S S L G L L S F T D L G M N G A V L Q M L S H G F I A A A L F F L SD L K R M V A Y S S I G H M G Y I L V S T A A G T E L S L L G A V A Q M I S H S L I L A L L F H L VD M K K V V A Y S S I A H M A F I L L A A A A A T P L S L A A A E I Q M V S H G L I S G L L F L L V
L K R . A Y S S I H M G . . L . . A T . L G A . Q M . S H G L I . A . L F L V
82
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
360 370 380 390 400
G A T Y D R T H T L M L D E M G G V G - - - K K M K K I F A M WT T C S M A S L A L P G M S G F V AG V T Y E R T H T L M M D E M S G I A - - - R L M P K T F A M F T A A A M A S L A L P G M S G F V SG I I E R K V G T R D L D V L N G L M N P V R G L P L T S S L L I L A G M A S A G I P G L V G F V AG I V Y K K T G S R D V D Y L R G L L T P E R G L P L T G S L M I L G V M A S A G L P G M A G F I AG . Y . T T . D G . . R . P T . . . M A S . . L P G M G F V A
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
410 420 430 440 450
E L M V F V G F A T S D A Y S P T F R V I I V F L A A V G V I L T P I Y L L S M L R E I L Y G P E NE L T V F L G L S N S D A Y S Y G F K P I A I F L T A V G V I L T P I T C F Q C C G V F Y G - - K GE F L V F Q G - - - - - - - S F S R F P I P T L F C I I A S G L T A V Y F V I L L N R T C F G R L DE F L I F R G - - - - - - - S F P V Y P V A T L L C M V G T G L T A V Y F L L M I N K V F F G R L TE V F G S . P I . L . V G . L T . Y . . . . G
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
460 470 480 490 500
K E L V A H E K L I D A E P R E V F V I A C L L I P I I G I G L Y P K A V T Q I Y A S T T E N L T AS Q A P P R C G G E D A K P R E I F V A V C L L A P I I A I G L Y P K L A T T T Y D L K T V E V A SS H T A Y Y P K V F A S - - - E K I P A I A L T V I I L F L G L Q P A WL T R WI E P T T S Q F I AP E L I N M S P V N WA - - - D Q F P A V M L V I L L F V F G L Q P Q WL V R WS E I D T A A L V A
. . . A E F A . . L . . I . . . G L P . T . T . . A
NdhD1-7002(slr0331)NdhD2-7002(slr1291)NdhD3-7002(slr1773)NdhD4 -7002(sll0027)
510 520 530 540 550
I L R Q S V P S L Q Q T A Q A P - - - - - - - - - S L D V A V L R A P E I RK V R A A L P L Y A E Q L P Q N G D R Q A Q M G L S S Q M P A L I A P R FA I P T V Q T I A L T P A E L S - - - - - - - - - - - - - - - - K A PS P T A I E I S L K N
. . . . A P
83
Table 5. Percent identity and similarity (identity/similarity) of the four NdhD proteins
from Synechococcus sp. PCC 7002 compared to one another as determined by a ClustalW
alignment (Thompson et al., 1994a).
% identity/similarity to Synechococcus sp. PCC 7002protein
NdhD1 NdhD2 NdhD3 NdhD4
Synechococcus sp. PCC 7002NdhD1
100/100 56/70 30/48 34/55
Synechococcus sp. PCC 7002NdhD2
56/70 100/100 30/47 33/51
Synechococcus sp. PCC 7002NdhD3
30/48 30/47 100/100 49/69
Synechococcus sp. PCC 7002NdhD4
34/55 33/51 49/69 100/100
3.1.4 ndhF1, ndhF2, ndhF3, ndhF4, ndhF5
Cyanobacteria display great genetic diversity at the ndhF loci. Synechococcus sp.
PCC 7002 has five divergent copies of the ndhF gene. NdhF proteins are homologs to
the NuoL protein from E. coli and the ND5 protein from the mitochondrial complex I
(Friedrich et al., 1995). All of the ndhF genes from Synechococcus sp. PCC 7002 are
predicted to encode hydrophobic proteins, likely associated with the membrane.
84
The ndhF1 (slr0844 homolog, accession number AAA27311) gene from
Synechococcus sp. PCC 7002 was originally identified and characterized by Schluchter et
al. (1993) and analysis of the gene sequence is predicted to encode a protein of 665
amino acids with a pI of 5.27.
The ndhF2 (slr2009 homolog) gene was originally cloned on a 1.5-kb BamHI-
HindIII fragment adjacent to the ndhF5 (slr2007 homolog) gene. The remaining
sequence to obtain the entire sequence of both the ndhF2 and ndF5 genes was acquired
from a cosmid from the cosmid library. The ndhF2 (slr2009 homolog) gene is predicted
to encode a protein of 479 amino acids and a pI of 9.24.
The ndhF3 (sll1773 homolog) gene was originally identified by Klughammer et
al. (1999) and the gene sequence is predicted to encode a protein of 617 amino acids with
a pI of 6.35.
The ndhF4 (sll0026 homolog) gene was cloned with the ndhD4 gene on a 2.6-kb
EcoRV-EcoRI fragment. The 5' end of the ndhF4 gene was cloned on a 3.15-kb XbaI-
EcoRI fragment. The ndhF4 gene is predicted to encode a protein of 633 amino acids
and a pI of 6.25.
The ndhF5 (slr2007 homolog) was partially isolated on a 1.5-kb BamHI-HindIII
fragment. The remaining nucleotide sequence was obtained from cosmid AG-4. The
ndhF5 gene is predicted to encode a protein of 479 amino acids and a pI of 9.25.
Figure 7 shows the ClustalW alignment of the five Synechococcus sp. PCC 7002
NdhF proteins. Table 6 shows the percent identity and similarity of the five
Synechococcus sp. PCC 7002 NdhF proteins to each other. Again, the diversity observed
85
Figure 7. ClustalW alignment of the five NdhF amino acid sequences from
Synechococcus sp. PCC 7002. The Synechococcus sp. PCC 7002 NdhD amino acid
sequences were aligned using the ClustalW alignment program from MacVector v. 6.5.
The consensus amino acid sequence is underneath the alignments. Gray high-lighted
regions indicate identical amino acids and lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids were determined with the
ClustalW program (Thompson et al., 1994b).
86
7002 NdhF ClustlW Amino Acid Alignments
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
10 20 30 40 50
M E P L Y Q Y A WL I P V L P L L G A M V I G I G L I S L N K F T N K L R Q L Y A V F V L S L I GM N E L T I G WV I - - - - - - - - - - - F P F V V G F S I Y L L P K I D R Y L A I F V S I C S
M N T F F S Q S V WL V P C Y P L L G M G L S A L WM P S I T R K T G P R P A G Y V N M L L T F M AM S E F L L Q S V WL V P V Y G I T G A L L T L P WS L G L I R R T G P R P A A Y L N L I M T F L G
M I D D I T I I WI L - - - - - - - - - - - L P F V V G F S I Y L L P R WN R Y F A L A I A A L S. Q . WL . P . G . . P . . . G . . . . T . P R . Y . . . . . . . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
60 70 80 90 100
T S M A L S - F G L L WS Q I Q G H E A F T Y T L E WA A A G D F H L Q M G Y T V D H L S A L M S VL I F G - - - F V Q I F Q P E P - - - - - - Y S L K L L G M Y G V D L L V D - D Q S G Y F I L T N AL V H S C L A F I E R WE Q P A L - - - - K P S L T WL Q A A D L T L S I D L D I S S I T I G A L IL L H G S F A F A S L WN M P P Q - - - - Q L S L E WL Q V A D L N L S L V I E I S P V N L G A M EV V Y S - - - I G L L WS L E P - - - - - - F T L E L L D S F G V T L M F D - E L S G Y F I L M N GL . . F . L W P . S L E WL . . D . L . D . . S . I L .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
110 120 130 140 150
I V T T V A L L V M I Y T D G Y M A H D P G Y V R F Y A Y L S I F S S S M L G L V F S P N L V Q V YA V A I A - - - - - - - V T V Y C WK S A K S A F F F T Q L V V L Q G A L N A V F V C A D L I S L YL I A G I N L L A Q L Y A V A Y L E M D WG WA R F F A T M S L F E A G M C A L V L C N S L F F S YL V T G I C F M A Q L Y G L G Y L E K D WS I A R F Y G L M G F F E A A L S G L A I S D S L L L S YL V T G A - - - - - - - V L L Y C F D K Q K S P F F Y T Q L V I L H G A V N A T F C C A D L I S L YL V T G . . . Y . . . Y . . D A R F Y . L . . F . A . A L . . C L . . Y
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
160 170 180 190 200
I F WE L V G M C S Y L L I G F WY D R K A A A D A C Q K A F V T N R V G D F G L L L G M L G L Y WV A L E A I S I A A F L L M T Y Q R T D R S I WI G L R Y L F L S N - T A M L F Y L I G A V L V Y QV V L E I L T L G T Y L L I G Y WF N Q S L V V T G A R D A F L T K R V G D L F L L M G V V A L L PG L L E V L T L S T Y L L V G F WY A Q P L V V T A A R D A F L T K R V G D I L L L M G I V A L S SV A L E C I G I A A F L L I T Y S R S D R S L WV G L R Y L F I S N - T A M L F Y L I G A V L V Y QV . L E . . . . . . Y L L I G Y W. . . . . G . R A F L T N R V G D L F L L . G . V . L Y
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
210 220 230 240 250
A T G S F E F D L M G D R L M D L V S T G Q I S S L L A I V F A V L V F L G P V A K S A Q F P L H VA T K S F A F V G L - - - - - - - - A E A P S D A I A L I F L G L L T K G G - - - - - - V F V S G LL A G T WN F D G L A E - - - - WA A T A E L D P T L A T L L C L A L I A G P L G K C A Q F P L H LY G T G L T F S E L E T - - - - WA A N P P L P P WE A S L V G L A L I S G P I G K C A Q F P L N LA S N S F A F S G L - - - - - - - - A V A P K E A I A L I F L G L L T K G G - - - - - - I F V S G LA . S F F G L . A A P . . . . A I . L G L L . . G P . . K A Q F P L L
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
260 270 280 290 300
WL P D A M - E G P T P I S A L I H A A T M V A A G V F L V A R M Y P V F E P I P E A M N V I A WTWL P L T H S E A E T P V S A M L - S G V V V K A G I F P L L R C G - - - I L V P D L D L WL R L FWL D E A M - E S P V P A T V V R - N S L V V G T G A WV L I K L Q P I F A L S D F A S T F M I A IWL D E A M - E G P N P A G I I R - N S V V V S A G A Y V L L K M E P V F T I T P I T S D A L I I IWL P L T H G E S E T P V S A L L - S G V V V K A G V F P L A R C A - - - L L V P E L D P V V R L FWL P . A M E . P T P . S A . . . V V V A G . F . L . R . P . F . L . P . . . . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
310 320 330 340 350
G A T T A F L G A T I A L T Q N D I K K G L A Y S T M S Q L G Y M V M A M G I G G Y T A G L F H L MG L A T A L L G I I F A I L E T D A K R L L A F S T I S K L G L L L S A P - - - - - A V A G L A A LG A T T A L G A A M V A I A Q I D I K R S L S Y S V S A Y M G M V F M A V G S Q Q D Q T T L V L L LG T V T T V G A S L V A L A Q I D I K R A L S H S T S A Y L G L V F I A V G L N Q V D I A L L L L LG V G T A L L G V G Y A V F E K D T K R M L A F H T V S Q L G F V L A A P - - - - - A V G G F Y A LG . . T A L L G . . . A . . Q D I K R . L A . S T S L G V . A G . . L . . L L
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
360 370 380 390 400
T H A Y F K A M L F L G S G S V I H G M E E V V G H N A V L A Q D M R L M G G L R K Y M P I T A T TS H G L V K S S L F L M A G - - - - - - - - - - - - - - - - - - - - - Q L P - - - T R - - - - - - -T Y G V A M A I L V M A I G G V V L - - - - - - - - - V N I S Q D L T Q Y G G L WS R R P I T G I CT H A I A K A L L F M S I G A V I L - - - - - - - - - N T H G Q N I T E M G G L WS R M P A T T S AT H G L V K G A L F L T A G - - - - - - - - - - - - - - - - - - - - - Q L S - - - S R - - - - - - -T H G . . K A . L F L . G V . Q Q G G L S R P . T
87
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
410 420 430 440 450
F L I G T L A I C G I P P F A G F WS K D E - - I L G L A F E A N P V L WF I G WA T A G M T A F Y- - - - - - - - - - - - - - - N F Q E L R Q T K I A S S L WL P L A I A C L S M V G M P L L V G F SY L V G A A S L V A L P P F G G F WS L A Q - - L T T N F WK T S P I L A V I L I T V N A L T S F SF V V G S A G L V C L F P L G T F WT M R R - - WV D G F WD T P P WL V L L L V G V N F C S S F N- - - - - - - - - - - - - - - N F K V L R E Q S I P R A Y WWV L V L A C A S I S G L P F L A G Y S. . . G . . . P . F W. L R I . . . W P . L . . . . G . . L . . F S
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
460 470 480 490 500
M F R M Y F L T F E G E F R G T D Q Q L Q E K L L T A A G Q A P E E G H H G S K P H E S P L T M T FS K A L L L K N I A - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - P - - - - - - - - -I M R E F G L I F G G K - - - - - - - - - - - - - - - - - - - - - - - - - - - - P K Q M T V R S P EL T R V F R S V F L G A - - - - - - - - - - - - - - - - - - - - - - - - - - - - P K P K T R R S P ES K I L T M K N I L - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - P - - - - - - - - -
R . . F . G P .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
510 520 530 540 550
P L M A L A V P S V L I G L L G V P WG N R F E A F V F S P N E - - - - A A E A A E H G F E L T E F- - WQ A M G L N I A A V G T A L A - - - - F A K F L F I P - - - - - - - - - - - - - - H D A A T KG L WA L V L P M V I L A G F A L H S P F I L A K L N F L P - - - - - - - - D WH Q L N L P L A A VV V WQ M A V P M V S L I L M T L M V P F F L H Q WQ L L F N P S L P T L V E R P L I V T L A I P A- - WQ S F A L N V A A V G T A I S - - - - F A K F I F L P - - - - - - - - - - - - - - K K T D P S
. WQ . . . P V . . . G A L F A K F . F L P . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
560 570 580 590 600
L I M G G N S V G I A L I G I T I A S L M Y L Q Q R I D P A R L A E K F P V L Y Q L S L N K WY F DF T A K S T T F WG A I A F L F S G V I L G N - - - - - - - - - - - - - - - - - - - - - - G F Y L EL I I S T M V G G G T A M Y L Y L N E K I S K P I H I F S D P V R E F F - - - - - - - A K D L Y T AL M I T G G L G L V A G L T I T L N P S L S R P R Q L Y L R F L Q D L L - - - - - - - A Y D F Y I DL K I L P N - F Y G A I V L L L G G L F V T N - - - - - - - - - - - - - - - - - - - - - - S F Y L EL I . . G A . . L . . . . . . . . F Y . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
610 620 630 640 650
D I Y N N V F V M G T R R L A R Q I L E V D Y R V V D G A V N L T G I A T L L S G E G L K Y I E N GA Y Q L D N I P K A L I K I A - - I G WA L Y WL I M K R I E F K - L P R I F - - - - - - - - - - -E L Y K N T V I F A V A L I S K I I D WL D R Y F V D G V I N F L G L A T L F G G Q S L K Y N N S GR I Y N V T V V WL V T T L S K L A A WF D R Y V V D G F V N L T G L A T L F S G S A L R Y N V S GA Y Q P S N I L K A L V T I G - - I G WL A Y G L I F Q R I T V K - L P R V V - - - - - - - - - - -
. Y . . A . . I . . I . W. D Y . . V D G I N . G L A T L F G L . Y G
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
660 670 680 690 700
R V Q F Y A L I V F G A V L G - - - - F V I F F S V A- E A F E Q L I G A M S V V L - - - - - T G L F WM V T L NQ S Q S Y A L S I V A G I L L - - - - F I A A L S Y P L L K H WQ FQ S Q F Y V L T I V L G M I L G L V WF M A T G Q WT M I T D F WS N Q L A- E Q F E H L V G V M S L V L - - - - - T G L F WL V L A N
Q F Y . L . . V . . . . L F . . F . .
88
Table 6. Percent identity and similarity (identity/similarity) of the four NdhD proteins
from Synechococcus sp. PCC 7002 compared to one another as determined by a ClustalW
alignment (Thompson et al., 1994a).
% identity/similarity to Synechococcus sp. PCC 7002protein
NdhF1 NdhF2 NdhF3 NdhF4 NdhF5
Synechococcus sp. PCC7002 NdhF1
100/100 13/28 25/46 25/25 15/29
Synechococcus sp. PCC7002 NdhF2
13/28 100/100 14/28 14/29 60/76
Synechococcus sp. PCC7002 NdhF3
25/46 14/28 100/100 45/64 62/75
Synechococcus sp. PCC7002 NdhF4
25/25 14/29 22/40 100/100 13/27
Synechococcus sp. PCC7002 NdhF5
15/29 60/76 24/44 13/27 100/100
89
among NdhF subunits may be responsible for multiple forms of the NDH-1 complex in
cyanobacteria.
3.1.5 Gene organization and phylogenetic analysis of the ndhD and ndhF
genes in cyanobacteria
The gene organization of several of the ndhD-ndhF gene clusters of cyanobacteria
is shown in Figure 8. The ndhF1 and ndhD1 genes from Synechococcus sp. PCC 7002
and Synechocystis sp. PCC 6803 are not found in close proximity to other ndh genes.
However, in Anabaena sp. PCC 7120, the ndhF1 gene and the ndhD1 gene are arranged
in a cluster with thioredoxin and an open reading frame homologous to the slr1621 gene
from Synechocystis sp. PCC 6803. The ndhF3 and ndhD3 genes are oriented in the same
transcriptional direction and arranged as clusters in all three cyanobacterial species. The
ndhF4 and ndhD4 genes are also arranged in similar clusters for all three cyanobacterial
species. The ndhF5 and ndhF2 genes are found nearly adjacent to one another in both
Synechococcus sp. PCC 7002 and Synechocystis sp. PCC 6803. However, Anabaena sp.
PCC 7120 appears to have only one homolog of the ndhF5 gene. In all three
cyanobacterial species, the ndhD2 gene is not found to be associated with other ndh
genes.
Sequence analysis shows that the NdhF and NdhD proteins are derived from a
common ancestor. Phylogenetic analysis reveals that the original nomenclature
Figure 8. Gene organization of the ndhF and ndhD genes in cyanobacteria. The ndhD and ndhF genes and gene clusters in
Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803 and Anabaena sp. PCC 7120 are shown. NADH dehydrogenase genes
shown in dark gray, genes not related to the NADH dehydrogenase shown in light gray. HP: hypothetical protein. h: homolog.
90
91
ndhD10.5 kbP1174 (1492<-1512)
ndhF3 ndhD3
HP, ORF427, sll1734 h.
rbcR HP, sll1735 h.
1 kb
ndhF4 ndhD4
HP, slr1302 h.
1 kb
ndhF5 ndhF2Na+/H+ antiporter
HP, slr2010 h.HP, slr2006 h.
HP, ssr3410 h.
1 kb
ndhD2SigG HP, slr1546 h.
1 kb
ndhF1 HP, sll0175 h.
HP, ssl3451 h.
PetH1kb
ndhF1ycf34 , ssr1425 HP, ssl1533
0.5 kb
ndhD1 HP, slr0333PsaC
0.5 kb
ndhF3 ndhD3 HP, sll1734HP, sll1730
1 kb
ndhF4 ndhD4HP, sll0518 NrtD, slr0044
1 kb
ndhF2ndhF5
HP, slr2010
HP, slr2008HP, slr2006 HP, ssr3409
1 kb
ndhD2 HP, sll1200HP, sll1201
1 kb
ndhF3 ndhD3 HP, sll1734 h.HP, slr1429 h.
1 kb
ndhF4 ndhD4 HP, slr1302 h./sll1734 h.ccmk-2, sll…
1 kb
ndhF5HP, sll0528 h. HP, slr2011 h.HP, slr2010 h.HP, slr2006 h.
1 kb
ndhD2 HP, slr1083 h.ExbB, sll1404 h.
1 kb
ndhF1 ndhD1 HP, slr1612 h.thioredoxin, …
1 kb
Synechococcus sp. PCC7002 Synechocystis sp. PCC6803 Anabaena sp. PCC7120
and classification of the cyanobacterial NdhF and NdhD proteins was incorrect. Figure 9
shows a representative phylogenetic tree with the evolutionary relationships among the
predicted NdhF and NdhD proteins from Synechococcus sp. PCC 7002, Synechocystis sp.
PCC 6803, Anabaena sp. PCC 7120, as well as the Ndh4 protein (accession number
NP_039348) and Ndh5 protein (accession number NP_039343) from the liverwort
Marchantia polymorpha (Ohyama et al., 1986), the NuoM and NuoL proteins from E.
coli (Weidner et al., 1993), and the NuoM protein (accession number AAC25004) and
NuoL (accession number AAC25003) proteins from R. capsulatus (Dupuis et al., 1995).
The Ndh protein sequences were aligned with the ClustalW program (Thompson et al.,
1994a) in MacVector v. 7.0 using MD (Dayhoff and National Biomedical Research
Foundation., 1965), PAM (Dayhoff and National Biomedical Research Foundation.,
1965), and BLOSUM (Henikoff and Henikoff, 1992) matrices to obtain the best
alignments of the protein sequences. The phylogenetic trees were made using the
neighbor-joining program in MacVector v. 7.0. Trees calculated from each of the
matrices were subjected to bootstrapping over 1000 cycles to assure that the relationships
displayed in the tree were those most likely to occur. The cyanobacterial NdhF proteins
cluster with the NuoL proteins E. coli and R. capsulatus as well as the liverwort Ndh5
protein from M. polymorpha. The cyanobacterial NdhD proteins group with the NuoM
proteins from E. coli and R. capsulatus as well as the liverwort Ndh4 protein from M.
polymorpha. The cyanobacterial Ndh proteins are most closely related to their identified
paralog in the other cyanobacteria except in the case of the NdhF5 and NdhF2 proteins.
NdhF5 was originally classified as an NdhD protein (Kaneko et al., 1996). However,
93
Figure 9. Phylogenetic analysis of NdhF and NdhD proteins. NdhF and NdhD
proteins Synechococcus sp. PCC7002, Synechocystis sp. PCC6803 and Anabaena sp.
PCC7120 and homologs from E. coli, R. capsulatus and M. polymorpha were subjected
to phylogenetic analysis with MacVector 7.0. A BLOSUM series matrix was used to
calculate the alignment between individual sequences with an Open/Extended gap
penalty of 10/0.05, Delay divergent: 60%. This is a representative tree after
bootstrapping for 1000 iterations. Numbers on branches represent percentages of times
that the proteins group together after analysis.
94
Method: Neighbor Joining; Bootstrap (1000 reps); tie breaking = SystematicDistance: Uncorrected (“p”)
Gaps distributed proportionally
NdhD1 7120
NdhD1 7002
NdhD1 6803
NdhD2 7002
NdhD2 7120
NdhD2 6803
NdhF2 7002
NdhF5 7120
NdhF5 7002
NdhF5 6803
NdhF2 6803
NdhF4 7120
NdhF4 7002
NdhF4 6803
NdhF3 7120
NdhF3 7002
NdhF3 6803
NdhF1 7120
NdhF1 7002
NdhF1 6803
NdhD3 7120
NdhD3 7002
NdhD3 6803
NdhD4 7120
NdhD4 7002
NdhD4 6803
NuoM R. capsulatus
NuoM E. coli
Ndh4 M. polymorpha
NdhD1 7120
NdhD1 7002
NdhD1 6803
NdhD2 7002
NdhD2 7120
NdhD2 6803
NdhF2 7002
NdhF5 7120
NdhF5 7002
NdhF5 6803
NdhF2 6803
NdhF4 7120
NdhF4 7002
NdhF4 6803
NdhF3 7120
NdhF3 7002
NdhF3 6803
NuoL E. coli
NuoL R. capsulatus
Ndh5 M. polymorpha
NdhF1 7120
NdhF1 7002
NdhF1 6803
NdhD3 7120
NdhD3 7002
NdhD3 6803
NdhD4 7120
NdhD4 7002
NdhD4 6803
100
100
100
100
100
99
100
100
100
100
99
99
100
100
95
100
99
98
94
72
100
100
95
phylogenetic analysis reveals that NdhF5 (slr2007 homolog) should be classified as an
NdhF protein and is actually most closely related to the NdhF2 protein from the same
organism. The ndhF5 and ndhF2 genes lie adjacent to each other in both the
Synechococcus sp. PCC 7002 and Synechocystis sp. PCC 6803 genomes; the high degree
of sequence similarity between the NdhF5 and NdhF2 proteins may be indicative of a
relatively recent gene duplication of either ndhF5 or ndhF2.
3.1.6 ndhCKJ
The Synechococcus sp. PCC 7002 ndhCKJ gene cluster was cloned on a 3.5-kb
BamHI fragment. The organization of the ndhCKJ gene cluster organization is shown
and compared to the other cyanobacterial ndhCKJ gene clusters in Figure 10. This
figure reveals that the Synechococcus sp. PCC 7002 ndhCKJ gene cluster is organized in
a manner very similar to the ndhCKJ gene clusters from both Synechocystis sp. PCC
6803 and Anabaena sp. PCC 7120, although the genes flanking the ndhCKJ cluster are,
as in the case of the cyanobacterial ndhAIGE gene cluster, very different from species to
species.
The ndhC gene from Synechococcus sp. PCC 7002 is predicted to encode a
protein of 121 amino acids and a pI of 6.54. Based on sequence similarity, NdhC is a
homolog to the NuoA protein from E. coli as well as the ND3 protein from the
mitochondrial complex I.
96
Figure 10. Gene organization of ndhCKJ gene clusters. Arrows indicate the direction
of transcription and size of the gene. (A) Synechococcus sp. PCC 7002 ndhCKJ gene
cluster. (B) Synechocystis sp. PCC 6803 ndhCKJ gene cluster. (C) Anabaena sp. PCC
7120 ndhCKJ gene cluster. Gene names or predicted protein names are indicated above
each gene. Individual scales for each map are in bp.
1K 2K 3K 4K 5K
kinesin light chainndhK ndhJndhCrubredoxin
transposase (slr1283)
sll0997
1K 2K 3K
ndhK ndhJsll0606 ndhChypothetical protein
Anabaena sp. PCC7120
1K 2K 3K 4K
transposase (slr1282)Threonine synthase ndhK ndhJndhC
ndhCKJ-6803Synechosystis sp. PCC6803
Synechococcus sp PCC7002
97
Sequence analysis predicts that the ndhK gene encodes a protein of 242 amino
acids and a pI of 5.71. NdhK is a homolog to the NuoB protein from E. coli and the
PSST protein from the mitochondrial complex I of B. taurus (Friedrich et al., 1995). The
Synechococcus sp. PCC 7002 NdhK protein contains the putative binding motifs (C57,
C58, C122, and C153) for a 4Fe-4S cluster. These cysteines are conserved in all NdhK
proteins from cyanobacteria (Kazusa DNA Research Institute, 2000; Kaneko et al.,
1996), the NdhK from spinach chloroplast (Schmitz-Linneweber et al., 2000), and the
homologous NuoB protein from E. coli (Weidner et al., 1993).
The ndhJ gene is predicted to encode a protein of 174 amino acids and a pI of
4.22. Based on sequence similarity, NdhJ is a homolog to the NuoC protein from E. coli
and the 30(IP) protein from B. taurus (Friedrich et al., 1995).
3.1.7 ndhH
The ndhH gene from Synechococcus sp. PCC 7002 was isolated on a 2.4-kb
BamHI fragment and is predicted to encode a protein of 395 amino acids and a pI of 5.66.
NdhH is a homolog of the NuoD protein from E. coli. It is also related to the 49(IP)
protein of mitochondrial complex I from B. taurus (Friedrich et al., 1995). The ndhH
gene is not associated with other ndh-genes, similar to the gene organization of the
Synechocystis sp. PCC 6803 ndhH gene (Figure 11). The genes downstream of the ndhH
gene are different in each species of cyanobacteria. In Synechococcus sp. PCC 7002,
there is a putative serine esterase gene downstream of the ndhH gene, followed by a
98
Figure 11. Gene organization around the ndhH genes of Synechococcus sp. PCC 7002
and Synechocystis sp. PCC 6803. Arrows indicate the direction of transcription and size
of the gene. (A) Gene organization around the Synechococcus sp. PCC 7002 ndhH gene.
(B) Gene organization around the Synechocystis sp. PCC 6803 ndhH gene. Maps are
individually scaled and numbers below each represent bp. Gene names and predicted
product names are indicated above genes (arrows). Restriction sites are as indicated on
maps.
99
300 600 900 1200 1500 1800 2100
ndhH sl r0262 s l r0263NdeI
HincII
StyI
HindIII
PvuI
SspI
PvuIPstI
SspI
SphI StyI
StyI
HincII
SspI
NcoI
StyI
EcoRI NcoI
StyI
B. Synechocystis sp. PCC 6803 ndhH
300 600 900 1200 1500 1800 2100
ndhH serine esterase
sl l0355BamHI BamHI
StyI
PvuI
EcoRV
PvuI
EcoRI
PvuIStyI
SspI
PvuI
HincII
A. Synechococcus sp. PCC 7002 ndhH
100
homolog of the Synechocystis sp. PCC 6803 hypothetical protein encoding sll0355.
In Synechocystis sp. PCC 6803 the gene organization around the ndhH gene is different.
It has two ORFs 3’ to ndhH (slr0262 and slr0263) which encode genes of unknown
function. These results reinforce that there are organizational differences for the genes
immediately surrounding the ndh genes amongst cyanobacteria.
3.1.8 ndhL
The ndhL gene from Synechococcus sp. PCC 7002 was isolated on a 1.75-kb
HindIII fragment from a partial genomic library. Sequence analysis of the
Synechococcus sp. PCC 7002 ndhL gene is predicted to encode a protein of 77 amino
acids and a pI of 9.42. A comparison of the gene organization around the ndhL genes of
Synechococcus sp. PCC 7002 and Synechocystis sp. PCC 6803 is shown in Figure 12.
The genes immediately downstream from the ndhL genes in both Synechococcus sp. PCC
7002 and Synechocystis sp. PCC 6803 are the same (slr0815). Based on proximity and
gene organization, these ORFs may be involved with the assembly or the function of the
type I NADH dehydrogenase. The ndhL gene was originally isolated as a mutant
deficient in its ability to uptake inorganic carbon and was called ictA (for inorganic
carbon transport A) (Ogawa, 1990) and was later renamed when found to be associated
with the type I NADH dehydrogenase complex (Ogawa, 1991). The predicted NdhL
proteins from cyanobacteria are very similar, and inactivation of the ndhL gene in
101
Figure 12. Gene organization around the ndhL genes of Synechococcus sp. PCC
7002 and Synechocystis sp. PCC 6803. Arrows indicate the direction of transcription and
size of the gene. (A) Synechococcus sp. PCC 7002 ndhL gene. (B) Synechocystis sp.
PCC 6803 ndhL gene. Maps are individually scaled and numbers below each represent
bp. Gene names and predicted product names are indicated above genes (arrows).
Restriction sites are as indicated on maps.
102
300 600 900 1200 1500 1800
sl r0816s l r0815sl l0812 ndhL
PvuI
StyI
StyI
StyI
PstI
NcoI
StyI
PvuI
NcoI
StyI
SspI
StuI
300 600 900 1200 1500
ndhL
TrpA (slr0966)slr0815 HindIII HindIII
NcoI
StyI
PvuI
StyI
StyI
PvuI
BamHI
EcoRV StyI
PvuI
PstI NcoI
StyI
PvuI
A. Synechococcus sp. PCC 7002 ndhL
B. Synechocystis sp. PCC 6803 ndhL
103
Synechococcus sp. PCC 7002 may lead to a phenotype similar to that observed by
Ogawa (Ogawa, 1990).
3.2 Type II NADH dehydrogenases
The ndb genes encode proteins of the type II NADH dehydrogenase family.
There are three ndb genes in Synechocystis sp. PCC 6803 (Howitt et al., 1999), but only
two of the three ndb genes could be found in Synechococcus sp. PCC 7002: ndbA and
ndbB. The Synechococcus sp. PCC 7002 ndbA open reading frame was isolated on a 2.1-
kb HindIII fragment from a partial genomic library. The ndbA gene is predicted to
encode a protein of 460 amino acids and a pI of 5.82. A comparison of the gene
organization around the ndbA genes from Synechococcus sp. PCC 7002 and
Synechocystis sp. PCC 6803 is shown in Figure 13. The genes flanking the ndbA genes
are very different in the two cyanobacteria.
The Synechococcus sp. PCC 7002 ndbB gene was isolated on a 2.0-kb BamHI-
HindIII fragment from a partial genomic library. Sequence analysis of the ndbB gene
from Synechococcus sp. PCC 7002 is predicted to encode a protein of 391 amino acids
with a pI of 5.61. A comparison of the gene organization around the ndbB genes from
Synechococcus sp. PCC 7002 and Synechocystis sp. PCC 6803 is shown in Figure 14.
As in Synechocystis sp. PCC 6803, the ndbA and ndbB proteins from Synechococcus sp.
PCC 7002 are predicted to be hydrophilic proteins.
104
Figure 13. Gene organization around the ndbA genes of Synechococcus sp. PCC
7002 and Synechocystis sp. PCC 6803. Restriction sites are as indicated. Arrows
indicate the direction of transcription and size of the gene. (A) Synechococcus sp. PCC
7002 ndbA gene. (B) Synechocystis sp. PCC 6803 ndbA gene. Maps are individually
scaled and numbers below each represent bp. Gene names and predicted product names
are indicated above genes (arrows). Restriction sites are as indicated on maps.
300 600 900 1200 1500 1800 2100 2400
ndbA ribosomal-protein-alanine N-acetyltransferase
s l r0852
SacI
DraI
NcoI
StyI
EcoRV
StyI
SspI NcoI
StyIEcoRI
EcoRI
PvuISspI
XbaI
HindIII
DraI
HincII
PvuI
AvaIStyI
HincII
EcoRIPvuI
HincII StyI
HincIIStyI
500 1000 1500 2000 2500 3000 3500 4000 4500 5000 5500 6000 6500 7000 7500 8000 8500
hypothetical protein ndbA (slr0851) s l r1938s l r0722hypothetical protein
HincII
DraI
HindIII
PvuI
DraI
StyI
SspI
PvuI
EcoRV
SspI
PvuI
PvuI
SspI
PvuI
PvuI
StyI
StyI
SspI
PvuI NcoI
StyI
StuI
PvuI
NcoI
StyI
EcoRI
BamHI
StyI
SspI
DraI
DraI
SspI
PvuI
HindIII
StyI
PvuI
SspI
HincII
DraI
HindIII
PvuIBamHI
BglII
PvuI
EcoRV
EcoRI
SspI
NcoI
StyI
PvuI
HincII
BglII HincII
SspI
PvuI
BglII
StyI
EcoRI
PstI
StuI
StyI
EcoRV
HindIII
HincII
A. Synechococcus sp. PCC 7002 ndbA
B. Synechocystis sp. PCC 6803 ndbA
105
Figure 14. Gene organization around the ndbB genes of Synechococcus sp. PCC
7002 and Synechocystis sp. PCC 6803. Arrows indicate the direction of transcription and
size of the gene. (A) Synechococcus sp. PCC 7002 ndbB gene. (B) Synechocystis sp.
PCC 6803 ndbB gene. Maps are individually scaled and numbers below each represent
bp. Gene names and predicted product names are indicated above genes (arrows).
Restriction sites are as indicated on maps.
106
300 600 900 1200 1500 1800 2100 2400 2700 3000
ndbBsl r1742 N-acetylmuramoyl-L-alanine amidase
SacI
HincII
PvuI
EcoRV
BamHI
SmaI
AvaI
NcoI
StyI
HincII
NcoI
StyI
HincII
PvuI
NcoI
StyI
HincII
StyI
AvaI
PvuI
HincII
HincII
StyI
StyI
StyI
StyI
PvuI
StyI
HincII
StyI
B. Synechocystis sp. PCC 6803 ndbB
300 600 900 1200 1500 1800 2100
ndbB hypothetical Zn-finger protein (CAB07242)KpnI
SacI
StyI NcoI
StyI
HincII
PvuI
XbaI
DraI
SphI
StyI
PvuI PvuI
SspI
BamHI
PvuI
StyI
BglII
StyI
StyIDraI
A. Synechococcus sp. PCC 7002 ndbB
107
The results presented here suggest that there are only two Ndb-type II NADH
dehydrogenases in Synechococcus sp. PCC 7002 that are most similar to the NdbA and
NdbB proteins from Synechocystis sp. PCC 6803.
3.3 Bi-directional Hydrogenase
The bi-directional hydrogenase apparent operon spans a 15-kb region of DNA in
Synechococcus sp. PCC 7002. Initially, the Synechococcus sp. PCC 7002 hoxH gene was
identified by Southern blot hybridization with a Synechocystis sp. PCC 6803 hoxH-
specific probe on a 4.5-kb EcoRV fragment. A Synechocystis sp. PCC 6803 hoxF-
specific probe was similarly used to identify cosmid 3C8 containing the Synechococcus
sp. PCC 7002 hoxF gene. The hydrogenase gene cluster was identified on two other
overlapping cosmids, cosmid 1C6 and cosmid 1E6, by using the hoxH and hoxF clones
isolated from Synechococcus sp. PCC 7002 as probes to screen the cosmid library. The
entire hydrogenase gene cluster was also identified on cosmid 5D12. The genes encoding
the bi-directional hydrogenase as well as the genes encoding the hydrogenase assembly
factors are tightly clustered in Synechococcus sp. PCC 7002 compared to other
cyanobacterial species (Figure 15). There are a total of 13 genes that have been
identified in this 15-kb nucleotide sequence that may encode subunits of the hydrogenase
enzyme or proteins involved in the assembly and maturation of the functional
hydrogenase. The 15-kb of nucleotide sequence has the following open reading frames:
hypE, hoxE, hoxF, hoxU, hoxY, hyp3, hoxH, hoxW, hypA, hypB, hypF, hypC, and hypD.
108
Figure 15. Hydrogenase gene cluster organization in cyanobacteria. Arrows indicate
the direction of transcription and size of the gene. (A) The hydrogenase gene cluster of
Synechococcus sp. PCC 7002. (B) The hydrogenase gene cluster of Synechocystis sp.
PCC 6803. (C) The hydrogenase gene cluster of Anabaena sp. PCC 7120. Maps are
individually scaled and numbers below each represent bp. Gene names and predicted
product names are indicated above genes (arrows).
1K 2K 3K 4K 5K 6K 7K 8K 9K 10K 11K 12K 13K 14K
hypFUreC hoxF hoxH hypDhypEhypB
hoxU hyp3hoxYhoxEhoxW
hypA hypC
1K 2K 3K 4K 5K 6K 7K 8K 9K 10K 11K 12K 13K 14K 15K 16K 17K 18K 19K 20K 21K 22K 23K 24K 25K
acetyl CoA synthasesll0163 hoxHslr0143 slr0090hoxF hoxU ORFhyp8
hoxYsll1388
hoxE hoxWsll0525
sll0163
1K 2K 3K 4K 5K 6K 7K 8K 9K 10K 11K
hoxF slr1334hoxHslr1233 hoxUsll1222
hoxY hoxEsll1225 transposasessl2420
Synechococcus sp. PCC7002
Synechocystis sp. PCC6803
Anabaena sp. PCC7120
Cyanobacterial hydrogenase gene organization
109
3.3.1 hypE
The hypE gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 348 amino acids with a pI of 4.8. This protein is presumably involved in assembly of
the hydrogenase enzyme in other bacteria (Colbeau et al., 1993; Drapal and Bock, 1998;
Jacobi et al., 1992), and is predicted to serve a similar purpose in Synechococcus sp. PCC
7002.
3.3.2 hoxE
The hoxE gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 171 amino acids with a pI of 6.34. There is a putative 2Fe-2S binding motif
(CX4CX35CXGXC) beginning at Cys-86 and ending at Cys-131 of the predicted
Synechococcus sp. PCC 7002 HoxE protein; this motif is similar to the that found in the
Synechococcus sp. PCC 6301 HoxE protein (Boison et al., 1998). The HoxE protein also
has some sequence similarity to the NuoE subunit of the E. coli (Blattner et al., 1997;
Weidner et al., 1993).
3.3.3 hoxF
The hoxF gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 536 amino acids and a pI of 4.78. This large subunit of the predicted diaphorase
110
heterodimer has sequence similarity to the NuoF subunit of the E. coli Type I NADH
dehydrogenase and is predicted to contain a flavin mononucleotide binding site for
transfer of electrons to and from NAD(P)H/NAD(P)+ (Appel and Schulz, 1996).
3.3.4 hoxU
The hoxU gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 238 amino acids and a pI of 5.73. The HoxU protein from Synechococcus sp. PCC
7002 is predicted to have 15 cysteines and has a conserved 2Fe-2S cluster binding motif
(PXLCX10-15CRXCXVX11-16C) near the amino terminus, starting at Pro-33 and ending at
Cys-64. This motif is found in all other HoxU proteins to date (Appel and Schulz, 1996).
Another conserved region of amino acids found in the Synechococcus sp. PCC 7002
HoxU protein which begins at Asp-95, is the sequence (EGNHXC). This sequence
overlaps the conserved sequence HXCX2CX5C and corresponds either to a 4Fe-4S cluster
similar to the 4Fe-4S cluster found in the small, crystallized subunit from D. gigas
(Volbeda et al., 1995) or 3Fe-4S cluster binding site (Appel and Schulz, 1996). There is
also a ferredoxin-like binding motif (CX2CX2CX3CXnCX2CX2CX3CP) located near the
carboxy terminus of the Synechococcus sp. PCC 7002 HoxU protein that begins at Cys-
148 and ends at Pro-203. These putative Fe-S clusters may be involved in distributing
electrons to either NAD+ or other electron acceptors from hydrogen to transfer electrons
through the respiratory electron transport chains.
111
3.3.5 hoxY
The hoxY gene of Synechococcus sp. PCC 7002 is predicted to encode a protein of
189 amino acids with five cysteines and a pI of 4.66. This proteins also has the Ni-Fe
hydrogenase consensus sequence CXGCXnGXCX3GXmGCPP, which is predicted to
ligand the proximal 4Fe-4S cluster in the hydrogenase moiety of the enzyme (Appel and
Schulz, 1996).
3.3.6 hyp3
The hyp3 gene of Synechococcus sp. PCC 7002 is predicted to encode a protein of
209 amino acids and a pI of 5.6. Synechocystis sp. PCC 6803 has no homolog of this
protein, but there are copies of the gene in Anabaena variabilis (Boison et al., 1998) and
Anabaena sp. PCC 7120. The function of this protein is unknown.
3.3.7 hoxH
The hoxH gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 476 amino acids and a pI of 6.62. This large subunit of the hydrogenase moiety is
predicted to have binding ligands for the Ni cofactor. There is a putative protease-
cleavage site located at His-448 (His-475 in the numbering scheme in Figure 40 because
of insertions to maximize the sequence similarity) in the Synechococcus sp. PCC 7002
112
HoxH protein. The 26 amino acids of the carboxy terminus may have to be cleaved
by HoxW in order for the maturation of the protein similar to the HoxH proteins of R.
eutropha (Massanz et al., 1997).
3.3.8 hoxW
The hoxW gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 143 amino acids and a pI of 4.59. The predicted HoxW protein has the consensus
hydrogenase-protease specific motif GXGNX4RDD/EGXG (Boison et al., 1998) and is
responsible for cleavage of HoxH for the maturation of the hydrogenase enzyme in other
bacteria (Massanz et al., 1997). It is likely that the HoxW protein serves the same
purpose in Synechococcus sp. PCC 7002.
3.3.9 hypA
The hypA gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 114 amino acids and a pI of 4.40. HypA is a predicted metal-binding assembly protein
containing five cysteines near the carboxy terminus of the protein. The HypA protein
from Synechococcus sp. PCC 7002 contains the CX2CX12-13CX2C motif that is found in
all HypA proteins and is characteristic of non-heme iron proteins like rubredoxins (Berg
and Holm, 1982) and certain zinc-binding regulatory domains (South and Summers,
1990), which suggests that HypA is a metalloprotein.
113
3.3.10 hypB
The hypB gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 274 amino acids and a pI of 6.21. This protein has 20 histidines in the amino terminus
that may be involved in Ni binding (Fu and Maier, 1994; Olson et al., 1997). Based on
the similarity that the Synechococcus sp. PCC 7002 HypB protein has with the HypB
proteins from other bacterial species, it also contains four regions that may be involved in
GTP binding (Fu and Maier, 1994; Olson et al., 1997).
3.3.11 hypF
The hypF gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 766 amino acids and a pI of 7.18. This protein has a Zn-finger binding motif
CX2X18CX24CX2CX18CX2C (Rey et al., 1993) beginning at Cys-110 and ending at Cys-
184. It has been postulated that HypF is required for the assembly of the mature
hydrogenase as well as hydrogen inducibility of the hydrogenase operon in R. capsulatus
(Colbeau et al., 1998). It has been postulated recently that the HypF proteins also have
an acylphosphatase (ACP) motif near the N-termini (Wolf et al., 1998). The biological
function of ACPs is unknown in prokaryotes. In eukaryotes, it is known that ACPs are
small (100 amino acids) proteins that catalyze the hydrolysis of the carboxyl-phosphate
bond in acylphosphates (Wolf et al., 1998).
114
Alignment of eukaryotic and prokaryotic ACPs with the Synechococcus sp. PCC
7002 HypF reveals that this regional similarity does exist in the HypF protein (Figure
16). Of note is the conserved Arg residue that corresponds to R23 in the well-defined
ACP from bovine testis. This Arg residue is conserved in all HypF proteins and ACPs to
date (Wolf et al., 1998). This residue is presumed to bind the phosphate moiety at the
active-site which abstracts a proton from a nucleophilic water molecule liganded to a
conserved Asn (N41). This residue is also conserved in all HypF sequences reported to
date. These observations point to the existence of an acylphosphatase domain in the
Synechococcus sp. PCC 7002 HypF protein.
Figure 16. ClustalW alignment of HypF proteins and acylphosphatases. The N-termini of several HypF proteins have been
aligned to the acylphosphatase proteins from three major acylphosphatase groups (Wolf et al., 1998). Groups are represented
as I, II, or III. Group I are the organ-common acylphosphatases, group II are the muscular acylphosphatases, and group III are
the prokaryotic homologs of acylphosphatases.
HypF and ACP ClustalW Formatted Alignments
chicken I PO7032pig I P24540human I P07311chicken II P07031Pig II P00819human II P14621E. coli III P75877A. fulgidus III O29440B. subtilis III O350317002 HypF N-term 1506803 HypF N-term 150R. eutropha HypF2 N-term 150E. coli HypF N-term 150R. capsulatus HypF N-term 150
5 10 15 20 25 30 35 40 45 50 55 60 65 70 75 80 85 90 95 100 105 110A G S E G L M S V D Y E V S G R V Q G V F F R K Y T Q S E A K R L G L V G WV R N T S H G T V Q G Q A Q G - P A A R V R E L Q E WL R K I G S P - Q S R I S R A E F T N E K E I A A L E H T D F Q I R K
S M A E G D T L I S V D Y E V F G K V Q G V F F R K Y T Q A E G K K L G L V G WV Q N T D Q G T V Q G Q L Q G - P T S K V R H M Q E WL E T R G S P - K S H I D R A S F N N E K V I S K L D Y S D F Q I V KM A E G N T L I S V D Y E I F G K V Q G V F F R K H T Q A E G K K L G L V G WV Q N T D R G T V Q G Q L Q G - P I S K V R H M Q E WL E T R G S P - K S H I D K A N F N N E K V I L K L D Y S D F Q I V K
S A L T K A S G S L K S V D Y E V F G R V Q G V C F R M Y T E E E A R K L G V V G WV K N T S Q G T V T G Q V Q G - P E D K V N A M K S WL S K V G S P - S S R I D R T K F S N E K E I S K L D F S G F S T R YS T A R P L K S V D Y E V F G R V Q G V C F R M Y T E D E A R K I G V V G WV K N T S K G T V T G Q V Q G - P E E K V N S M K S WL S K I G S P - S S R I D R T N F S N E K T I S K L E Y S N F S I R Y
M S T A Q S L K S V D Y E V F G R V Q G V C F R M Y T E D E A R K I G V V G WV K N T S K G T V T G Q V Q G - P E D K V N S M K S WL S K V G S P - S S R I D R T N F S N E K T I S K L E Y S N F S I R YM S K V C I I A WV Y G R V Q G V G F R Y T T Q Y E A K R L G L T G Y A K N L D D G S V E V V A C G - E E G Q V E K L M Q WL K S - G G P R S A R V E R V L S - - E P H H P S G E L T D F R I R
M I A L E I Y V S G N V Q G V G F R Y F T R R V A R E L G I K G Y V K N L P D G R V Y I Y A V G - E E L T L D K F L S A V K S - G P P - - - - L A T V R G V E V K K A E I E N Y E S F E V A YM L Q Y R I I V D G R V Q G V G F R Y F V Q M E A D K R K L A G WV K N R D D G R V E I L A E G - P E N A L Q S F V E A V K N - G S P - F S K V T D I S V T E S R S L E G - - H H R F S I V Y
L L V K R R L R L E I Q G T V Q G V G F R P F V Y Q L A T A L N L F G WV N N S T A G - V T I E V E G - G R S P L N L F L E K L Q A E L P P - - - - N A K I D A L K Y Q Y L E L I G Y N N F E I H AM L K T V A I Q V Q G R V Q G V G F R P F V Y T L A Q E M G L N G WV N N S T Q G A T V V I T A D - - E K A I A D F T E R L T K T L P P - - - - P G L I E Q L A V E Q L P L E S F T N F T I R P
M L M P R R P R N P R T V R I R I R V R G V V Q G V G F R P F V Y R L A R E L G L A G WV R N D G A G - V D I E A Q G S A A A L V D S R L R R L R R D A P P - L A R V D E I G E - - E R C A A Q V D A D G F A I L EM A K N T S C G V Q L R I R G K V Q G V G F R P F V WQ L A Q Q L N L H G D V C N D G D G - V E V R L R E - - - - D P E T F L V Q L Y Q H C P P - L A R I D S V E R - - E P F I WS Q L P T E F T I R Q
M Q A WR I R V R G Q V Q G V G F R P F V WQ L A R A R G L R G V V L N D A E G - V L I R V A G - - - - D L G D F A A A L R D Q A P P - L A R V D A V E V T - - A A V C D D L P E G F Q I A AV . V G . V Q G V G F R . T E A . . L G L . G WV . N . G V . . G V L G P R . D . . E . . . . . . F I .
115
3.3.12 hypC
The HypC protein is required for hydrogenase maturation in R. eutropha (Dernedde et al.,
1993), E. coli (Lutz et al., 1991), B. japonicum (Olson and Maier, 1997), and may also be
required for the maturation of the enzyme in cyanobacteria. The hypC gene of
Synechococcus sp. PCC 7002 is predicted to encode a protein of 80 amino acids and a pI
of 4.04 and has a high degree of similarity to other bacterial HypC proteins.
3.3.13 hypD
The hypD gene of Synechococcus sp. PCC 7002 is predicted to encode a protein
of 362 amino acids and a pI of 6.71. The predicted protein is very similar to other
bacterial HypD proteins. HypD has been shown to play a role in hydrogenase maturation
in E. coli (Jacobi et al., 1992) and in R. eutropha (Dernedde et al., 1993) and may play a
similar role in Synechococcus sp. PCC 7002.
116
117
3.3.14 Interposon mutagenesis of the hoxH and hoxF genes in
Synechococcus sp. PCC 7002
The physiological role of the hydrogenase in cyanobacteria is unknown. In order
to investigate its possible role in electron transport, the hoxH gene was chosen for
interposon mutagenesis. The hoxH gene is predicted to encode the large Ni-containing
subunit of the hydrogenase. It was hoped that the absence of the large subunit would lead
to an inactivation of the entire hydrogenase. The hoxH gene was interrupted by the
insertion of a 1.3-kb BamHI DNA fragment containing the aphII gene and conferring
kanamycin resistance into a unique BglII site in the coding sequence of hoxH (Figure
17). This plasmid, pHOXH, was used to transform the wild-type strain of Synechococcus
sp. PCC 7002. Segregation of alleles in the transformants hoxH::aphII of Synechococcus
sp. PCC 7002 was verified by PCR analysis using primer 311 (5'-G CCC CAT CAT
CAG AAC G-3') and primer 312 (5'-GAT TTA GCA CGG GGT GG-3') (Figure 17).
Results of the PCR analysis reveal that the hoxH::aphII locus successfully replaced all
wild-type copies of the hoxH gene. Therefore, the hoxH gene is not essential for cell
viability, under normal photoautotrophic growth conditions.
As mentioned in 1.1.2, although cyanobacteria have ndh genes, which putatively
encode a type I NADH dehydrogenase, genes encoding subunits for the diaphorase
activity of type I NADH dehydrogenase are missing. It may be possible that some of the
diaphorase active subunits from the hydrogenase of Synechococcus sp. PCC 7002 are
able to substitute for these missing type I NADH dehydrogenase subunits. In order to
118
Figure 17. Interposon mutagenesis of hoxH of Synechococcus sp. PCC 7002. (A)
Physical map of the Synechococcus sp. PCC 7002 hoxH gene. Arrows indicate the
direction of transcription and size of the gene. A 1.3-kb BamHI fragment containing the
aphII gene conferring kanamycin resistance was inserted into a unique BglII site within
the coding sequence of the hoxH gene. The resultant plasmid, pHOXH was used to
transform the wild-type strain of Synechococcus sp. PCC 7002. (B) PCR analysis of the
hoxH locus. Total DNA was isolated from the wild-type and kanamycin-resistant strains
of Synechococcus sp. PCC 7002. This DNA was used as a template for PCR reactions.
119
hoxH hoxW hypA
aphII
hypB
Eco
RV
Eco
RV
HIn
cII
HIn
cII
HIn
dIII
HIn
dIII
Bgl
II
Sph
I
∆hoxH::a
phII
wild ty
pe
HE ladder
A
B
1.1 kb
2.4 kb
120
assess the possible role of the bi-directional hydrogenase as a component of the type I
NADH dehydrogenase in Synechococcus sp. PCC 7002, interposon mutagenesis was
performed on the hoxF gene. The hoxF gene was interrupted by inserting a 2-kb BamHI
DNA fragment containing the aadA gene from the Ω fragment (Prentki and Krisch, 1984)
conferring spectinomycin and streptomycin resistance into a unique BglII site within the
predicted coding sequence of hoxF (Figure 18). The resulting plasmid, pHOXF, was
used to transform both the wild-type and hoxH::aphII strains of Synechococcus sp. PCC
7002. The segregation of the hoxF::Ω allele was verified by Southern blot analysis
(Figure 18). Results indicate that the all wild-type copies of the hoxF gene were
successfully displaced by the hoxF::Ω allele in three independently isolated
spectinomycin transformants and a kanamycin/spectinomycin resistant transformant.
These results indicate that the hoxF gene product is not essential for cell viability under
normal growth conditions. The absence of a clear phenotype similar to those associated
with either the NdhF1-deficient (Schluchter et al., 1993), or NdhF3-deficient and NdhD3-
deficient (Klughammer et al., 1999) strains of Synechococcus sp. PCC 7002 suggests that
the HoxF gene product is not important in complexes involving these Ndh proteins.
121
Figure 18. Interposon mutagenesis of hoxF of Synechococcus sp. PCC 7002. (A)
Physical map of the Synechococcus sp. PCC 7002 hoxF gene. Arrows indicate the
direction of transcription and size of the gene. A BamHI fragment containing the Ω
fragment conferring streptomycin and spectinomycin resistance was inserted into a
unique BglII site within the coding sequence of the hoxF gene. (B) Southern blot
analysis of the hoxF locus. Total DNA was isolated from the wild-type and
spectinomycin resistant strains of Synechococcus sp. PCC 7002. This DNA was digested
with HindIII and BamHI and transferred to a nylon membrane. A 32 P-labeled hoxF
specific probe was hybridized to the blot. Lanes 1 and 6, wild-type Synechococcus sp.
PCC 7002; lane 2, hoxF/hoxF::Ω merodiploid; lane 3, hoxF::Ω ; lane 4 and 5,
hoxH::aphII hoxF::Ω.
122
1 2 3 4 5 6
hoxE hoxF
Bam
HI
HIn
cII
Bgl
II
Eco
RI
Ω
Sph
IB
amH
I
Bam
HI
Hin
dIII
A
B
1.2 kb
3.2 kb
123
3.3.15 Chlorophyll and carotenoid contents of the hoxH- and hoxH- hoxF-
strains
Although no immediate growth phenotypes were obvious, inactivation of the hox
genes in Synechococcus sp. PCC 7002 may have had other effects on the cells and their
contents. Physiological parameters that can be readily measured are the chlorophyll and
carotenoid contents. The chlorophyll and carotenoid contents of the hoxH- and hoxH-
hoxF- mutant strains were compared to those of the wild-type strain grown under standard
conditions. The wild-type strain had a chlorophyll concentration of 3.6 ± 0.1 µg ml-1,
compared to 3.3 ± 0.1 µg ml-1 for the hoxH- strain and 3.5 ± 0.1 µg ml-1 for the hoxH-
hoxF- double mutant strain. The carotenoid contents were 1.0 ± 0.1 µg ml-1 for the wild-
type strain, 1.0 ± 0.1 µg ml-1 for the hoxH- strain, and 1.1 ± 0.1 µg ml-1 for the hoxH-
hoxF- double mutant strain. These results reveal that under standard growth conditions,
there is no significant difference in chlorophyll and carotenoid contents between the
hoxH- and hoxH- hoxF- mutant strains compared to the wild-type strain of Synechococcus
sp. PCC 7002.
124
3.3.16 Growth analysis of hoxH- and hoxH- hoxF- double mutants
The doubling times of both the hoxH- mutant and hoxH- hoxF- double mutant are
nearly identical to those of the wild-type strain (3.8 ± 0.1 hours) under standard growth
conditions. The doubling time of the hoxH- strain is 3.9 ± 0.3 hours and the doubling
time of the hoxH- hoxF- strain is 3.8 ± 0.2 hours. These results indicate that the doubling
times of the hoxH- and hoxH- hoxF- double mutant are identical to the wild-type under
normal growth conditions (38°C, 250 µE m-2s-1, 1.5% CO2/98.5% air). Temperature
shifts from normal growth temperatures to lower temperatures have been shown to lead
to photoinhibition in other cyanobacterial strains with lesions in the electron transport
chain (Clarke and Campbell, 1996). Therefore, a temperature shift experiment was
conducted to determine whether or not there would be an effect on the growth rates from
the inactivation of the hydrogenase genes. The effects of a temperature-shift from 38°C
to 22°C on the growth rates of hoxH- and wild-type strains of Synechococcus sp. PCC
7002 are shown in Figure 19. These results indicate that the hoxH gene is not involved
in low temperature adaptation in Synechococcus sp. PCC 7002. Moreover, the wild-type
and mutant strains do not grow differently under conditions of nickel limitation at 22°C
when nitrate is the nitrogen source (T. Sakamoto, personal communication). These
results prove the fact that neither the hoxH nor the hoxF gene products are necessary for
the cell under these conditions.
125
Figure 19. Temperature-shift effects on wild-type and hoxH::aphII. Exponentially
growing cells were either maintained at 38°C or transferred to a 22°C water bath. (A)
Temperature-shift effects on the hoxH::aphII strain of Synechococcus sp. PCC 7002.
The hoxH::aphII cells were grown at 38°C () and 22°C (). (B) Temperature-shift
effects on the wild-type strain of Synechococcus sp. PCC 7002. Wild-type cells grown at
38°C () and 22°C (). The data shown are from a single experiment. The experiment
was repeated three times, and the results were consistent and reproducible.
126
OD
550
0.01
0.1
1
10
0 10 20 30 40 50
Time (hours)
∆hoxH::aphII (22˚C)
∆hoxH::aphII (38˚C)
OD
550
0.01
0.1
1
10
0 10 20 30 40 50
Time (hours)
WT (22˚C)WT(38˚C)
A
B
127
3.4 Mobile Electron Carriers
Four potential mobile electron carriers were identified in this study. Three genes
(petJ1, petJ2, cytM) are predicted to encode cytochrome c proteins, and the fourth gene,
bcpA, is predicted to encode a putative type I blue-copper protein. Plastocyanin is
encoded by the petE gene in higher plants (Scawen et al., 1975), algae (Merchant et al.,
1990), and other cyanobacteria (Briggs et al., 1990; Clarke and Campbell, 1996).
Although Synechococcus sp. PCC 7002 genomic DNA blots were probed with
heterologous petE specific probes, no cross-hybridizing signals could be found for the
petE gene. Likewise, no plastocyanin-like sequences have yet been identified in the
Synechococcus sp. PCC 7002 genome sequencing project.
3.4.1 Cloning and sequence analysis of the Synechococcus sp. strain
PCC 7002 cytochrome c6 gene
Initial Southern blot hybridization screens using heterologous PCR product
probes (cytochrome c6 genes from Synechocystis sp. PCC 6803 or Synechococcus sp.
PCC 7942) at low stringency did not produce positive hybridization signals. Thus, the
Synechococcus sp. strain PCC 7002 protein was purified as described in Materials and
Methods. Cytochrome c6 was purified to apparent electrophoretic homogeneity based
upon its characteristic absorbance maxima at 416 nm and 553 nm. Purity was assessed
by electrophoresis on a polyacrylamide gel in the presence of SDS and was followed by
128
Coomassie staining of the proteins and TMBZ detection of heme groups (Figure 20).
The N-terminal sequence from the purified cytochrome c6 was determined to be
ADAAAGAQVFAANCA. Based upon this sequence, the degenerate oligonucleotide: 5'-
GCT GA(T/C) GCT GCT GCT GGT GCT CA(A/G) GTT TT(T/C) GCT GCT AA(T/C)
TG(T/C) GC-3' was synthesized for use as a forward primer for PCR. This primer was
used with the degenerate oligonucleotide: 5'-CCA CGC TTT TTC TGC TTG GTC GAG
TAC GTA GCG TGC NAC (A/G)TC-3' derived from the conserved cytochrome c6
sequence (DVAAYVLDQAEKGW) of other cytochrome c6 proteins as the reverse
primer to amplify by PCR an internal region of the Synechococcus sp. strain PCC 7002
cytochrome c6 gene sequence from total Synechococcus sp. strain PCC 7002 genomic
DNA for use as a probe. An internal degenerate PCR primer: 5'-AA(T/C) TG(T/C)
GC(T/C) GC(T/C) TG(C/T) C-3' was also designed from the conserved heme attachment
site amino acid sequence (NCAACH) as determined by cytochrome c6 sequence
alignments using the BLAST search tool (Altschul et al., 1990). The resultant PCR
products were used as probes for Southern blot analysis of Synechococcus sp. strain PCC
7002 chromosomal DNA digested with multiple restriction enzymes. A hybridizing
HindIII-XbaI fragment of approximately 1.7-kb was chosen to clone into pUC19 (Figure
21). The nucleotide sequence of this clone has been deposited into GenBank (accession
number: AF020306). The petJ coding region contains a single complete open reading
frame of 351 bp that can encode to a preprotein of 117 amino acids, this ORF was
positioned between the sequences of two other partial open reading frames (Figure 22).
Sequence analysis of the partially sequenced open reading frame 5'
129
Figure 20. SDS-PAGE analysis of cytochrome c6 from Synechococcus sp. PCC
7002. Soluble protein fractions from Synechococcus sp. PCC 7002 isolated by
ammonium sulfate precipitation and mono-Q ion-exchange chromatography were
electrophoresed on two SDS polyacrylamide gels. (A) TMBZ stained polyacrylamide
gel. Lane 1, fraction 1 from mono-Q column chromatography of purified cytochrome c6
from Synechococcus sp. PCC 7002. Lane 2, fraction 2, mono-Q column
chromatography, Lane 3, fraction 3, mono-Q column chromatography, Lane 4, 95%
ammonium sulfate cut supernatant, Lane 5, E. coli protein extract, Lane 6, horse heart
cytochrome c. (B) Coomassie stained polyacrylamide gel. Lanes are loaded identically
as for Panel A. A total of 5 µg of protein was loaded per lane.
131
Figure 21. Probe design and Southern blot hybridization of petJ from Synechococcus
sp. strain PCC 7002. (A) Degenerate oligonucleotide sequences for PCR primers.
Primer 1 was a degenerate oligonucleotide derived from the N-terminal sequence of the
Synechococcus sp. strain PCC 7002 purified cytochrome c6. Primer 2 was a degenerate
oligonucleotide derived from the conserved amino acid sequence of the heme attachment
motif (XCXXCH) of several cytochrome c6 proteins. Primer 3 was a degenerate
oligonucleotide derived from a semi-conserved amino acid sequence at the carboxy-
terminus of several cytochrome c6 proteins. The predicted amino acid sequences for the
degenerate oligonucleotides are shown above the nucleotide sequences. Two different
probes were made using these degenerate oligonucleotides, probe 1 and probe 2. The
predicted PCR probe product sizes are beneath the primer sequences. (B) Autoradiogram
of genomic digests of Synechococcus sp. strain PCC 7002 DNA probed with probe 2 (32P-
radiolabelled PCR product of primer 1 and primer 3 of (A)). Genomic DNA from 7002
was digested with various restriction enzymes and hybridized at 55°C. Lane 1: BglII,
Lane 2: BamHI, Lane 3: EcoRI, Lane 4: HindIII, Lane 5: XbaI, Lane 6: BglII/BamHI,
Lane 7: BglII/EcoRI, Lane 8: BglII/HindIII, Lane 9: BamHI /EcoRI, Lane 10:
BamHI/XbaI, Lane 11: BamHI/HindIII, Lane 12: EcoRI/HindIII, Lane 13: EcoRI/XbaI,
Lane 14: HindIII/XbaI. Only one copy of the petJ gene is detectable at a hybridization
temperature of 55˚C. The smallest hybridizing DNA fragment (1.74-kb HindIII-XbaI)
was chosen to clone into pUC19.
132
A.
A D A A A G A Q V F A A N C A5' GCT GAT/C GCT GCT GCT GGT GCT CAA/G GTT TTT/C GCT GCT AAT/C TGT/C GC 3'
Primer 1
N C A A C H5' AAT/C TGT/C GCT/C GCT/C TGC/T C 3'
Primer 2
D V A A Y V L D Q A E K G W3' CTG/A CAN CGT GCG ATG CAT GAG CTG GTT CGT CTT TTT CGC ACC 5'
Primer 3
Probe 1
Probe 2
238 bp
275 bp
B.
kb
4.27
21.7
5.00
3.48
1.981.901.601.37
0.94
1 2 3 4 5 6 7 8 9 10 11 12 13 14
133
Figure 22. Gene organization around petJ1 in cyanobacteria. Arrows indicate
direction of transcription. (A) Gene organization around the petJ1 gene of Synechococcus
sp. PCC 7002. (B) Gene organization around the petJ gene of Synechocystis sp. PCC
6803. Maps are individually scaled in bp. Gene names and predicted product names are
indicated above genes (arrows). Restriction sites are as indicated on maps.
134
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300
sll0185 homolog sll0415 ABC transporterpetJ1AvaI
PvuI
PvuI
PvuISacI
HindIIIEcoRV
BamHIStyI
KpnISspI
StyIPvuI
PvuI
HindIII
KpnI StyI
SspI
XbaI
PvuI
AvaI
HincII
A. Synechococcus sp. PCC 7002 petJ1
B. Synechocystis sp. PCC 6803 petJ
300 600 900 1200
sl r1896 petJ srrA (slr1897)
PvuI
NcoI
StyI
StyI
StyI
HincII
NcoI
StyI
HincII
PvuI
135
to the petJ gene indicates that it has high sequence homology to an ABC transporter
protein (sll0415 in the Synechocystis sp. PCC 6803) (Kaneko et al., 1996). The partial
open reading frame 3' to the petJ gene is predicted to encode a hypothetical protein of
unknown function (sll0815 in the Synechocystis sp. PCC 6803) (Kaneko et al., 1996).
Both of these hypothetical reading frames would be transcribed in the opposite direction
of Synechococcus sp. strain PCC 7002 petJ gene.
The petJ nucleotide sequence is predicted to encode a pre-protein transit peptide
sequence of 24 amino acids (Figure 23) that displays many characteristics (a basic N-
terminal region with two lysines (n-region), a central hydrophobic region (h-region) and a
C-terminal region that follows the '-3, -1 rule' where the amino acids at the -3 and -1
positions are alanines) common for prokaryotic signal peptides (Gierasch, 1989; von
Heijne, 1986). The mature protein would begin at amino acid 25 (Ala) and would thus
contain 93 amino acids. As noted above, this was confirmed by N-terminal amino acid
sequencing of the gel purified cytochrome c6 protein from Synechococcus sp. strain PCC
7002. PetJ has a molecular mass of 9.4 kDa and analysis of the amino acid sequence
predicts a pI of 4.89. There is a potential transcription terminator (-18 kcal mol-1)
located between nucleotide positions 970 and 990 about 150 bases downstream from the
stop codon of the petJ1 gene.
A comparison of mature cytochrome c6 proteins is shown in Figure 23. The
cytochrome c6 proteins were aligned using the ClustalW tool (Thompson et al., 1994b).
Cytochrome c6 from Synechococcus sp. strain PCC 7002 has 24 amino acids that are
absolutely conserved and overall is 46%-54% identical to other cytochrome c6 proteins.
136
Figure 23. ClustalW alignment of PetJ amino acid sequences. (A) Deduced cyt c6
amino acid sequences were aligned by the ClustalW multiple protein sequence alignment
function in the MacVector 6.5 program. Dark gray shading indicates identical or a
majority identical amino acid, the lighter gray shading indicates conserved amino acids,
and no shading indicates non-conserved amino acids. Dashes indicate gaps introduced to
maximize the alignment. The consensus cyt c6 sequence is indicated below. Organisms
are indicated as follows: 7002, Synechococcus sp. PCC 7002; S. maxima, Spirulina
maxima; C. reinhardtii, Chlamydomonas reinhardtii; 7937, Anabaena sp. PCC 7937;
7942, Synechococcus sp. PCC 7942; 7120, Anabaena sp. PCC 7120; 6803, Synechocystis
sp. PCC 6803. (B) Deduced cyanobacterial signal transit peptide sequences for cyt c6 and
plastocyanin from various organisms were aligned as described above. Dark gray
shading indicates identical or majority identical amino acids, while the lighter gray
shading indicates conserved amino acids, and no shading indicates non-conserved amino
acids. Hyphens indicate gaps introduced to maximize the alignment. Conserved or
identical signal sequence amino acids are indicated below. Organisms are as described
above. The signal peptide of cyt c6 from Synechococcus sp. PCC 7002 shares many of
the traits of typical prokaryotic signal peptide sequences (see text).
137
ClustalW Formatted Alignments
70026803S. maxima7942C. reinhardtii71207937
10 20 30 40 50
60 70 80 90
A D A A A G A Q V F A A N C A A C H A G G N N A V M P T K T L K A D A L K T Y L A G Y K D G S K S LA D L A H G K A I F A G N C A A C H N G G L N A I N P S K T L K M A D L E A N G K - - - - - - - N SG D V A A G A S V F S A N C A A C H M G G R N V I V A N K T L S K S D L A K Y L K G F D D - - - D AA D L A H G G Q V F S A N C A A C H L G G R N V V N P A K T L Q K A D L D Q Y G M - - - - - - - A SA D L A L G A Q V F N G N C A A C H M G G R N S V M P E K T L D K A A L E Q Y L D G G - - - - - F KA D S V N G A K I F S A N C A S C H A G G K N L V Q A Q K T L K K A D L E K Y G M - - - - - - - Y SA D V A N G A K I F S A N C A S C H A G G K N L V Q A Q K T L K K E D L E K F G M - - - - - - - Y SA D . A . G A . . F . A N C A A C H G G . N . . . P K T L K K A D L E . Y G S
70026803S. maxima7942C. reinhardtii71207937
10 20 30 40 50
60 70 80 90E E A V A Y Q V T N G Q G A M P A F G G R L S D A D I A N V A A Y I A D Q A E N N K WV A A I V A Q I T N G N G A M P G F K G R I S D S D M E D V A A Y V L D Q A E K G - WV A A V A Y Q V T N G K N A M P G F N G R L S P K Q I E D V A A Y V V D Q A E K G - WI E A I T T Q V T N G K G A M P A F G S K L S A D D I A D V A S Y V L D Q S E K G - W Q GV E S I I Y Q V E N G K G A M P A W A D R L S E E E I Q A V A E Y V F K Q A T D A A W K YA E A I I A Q V T N G K N A M P A F K G R L K P E Q I E D V A A Y V L G K A D A D - W KA E A I I A Q V T N G K N A M P A F K G R L K P D Q I E D V A A Y V L G Q A D K S - W K. E A I . . Q . T N G K G A M P A F G R . S . . . E D V A A Y V L D Q A E K . W .
A. ClustalW alignment of cytochrome c6 proteins
ClustalW Formatted Alignments
7002 C6 signal sequence7120 C6 signal sequence7937 C6 signal sequence7942 C6 signal sequence7942 PC signal sequence6803 PC signal sequence6803 C6 signal sequence
10 20 30L K K L L A I A L T - V L A T V F A F G - - - T P A F AM K K I F S L V L L G I A L F T F A F S - - - S P A L AM K K I F S L V L L G I A L F T F A F S - - - S P A L AM K R I L G T A I A - A L V V L L A F I - - - A P A Q A
M K V L A S F A R R L S F A V A - A V L C V G S F F L S A A P A S AM S K K F L T I L A G L L L V V S S F F L S V S P A A A A
M F K L F N Q A S R I F F G I A L P C L I F L G G I F S L G N T A L AM K K I F . . L . . L L . A F . P A A
B. ClustalW alignment of cyanobacterial signal sequences for cyt c6 and plastocyanin
138
It is interesting to note that there appears to be an insertion of 7 amino acids starting
at glycine 42 and ending at lysine 48. The most similar cytochrome c6 protein sequence
is that from Spirulina maxima, which also has extra inserted amino acids when compared
to cytochrome c6 proteins from other cyanobacteria.
3.4.2 Attempted deletion of the petJ1 gene by interposon mutagenesis.
The aphII gene, which confers kanamycin resistance, was inserted into a unique
BstXI site within the coding region of the Synechococcus sp. PCC 7002 petJ1 gene. This
plasmid was used to transform the wild-type strain of Synechococcus sp. PCC 7002 and
kanamycin resistant transformants from A+ plates were selected for growth in liquid
media. Unlike the previously described deletion of the petJ gene from Synechocystis sp.
strain PCC 6803 (Zhang et al., 1994) or the interposon interruption of the cytA gene from
Synechococcus sp. strain PCC 7942 (Laudenbach et al., 1990) where the functional
copies of the gene encoding cytochrome c6 were replaced by the mutant alleles, attempts
to segregate the petJ1::aphII and petJ1 alleles in Synechococcus sp. strain PCC 7002
failed. This indicates the importance of a wild-type copy of the gene for cell viability
(Figure 24). Northern blot analysis of the Synechococcus sp. strain PCC 7002 petJ1
transcript reveals that the petJ1 gene is accumulated as a monocistronic transcript
(Figure 25). The results of Northern blot analysis and the fact that the genes adjacent to
petJ1 would be transcribed in the opposite direction (Figure 22) reveal that a polar effect
in an essential gene is unlikely. Attempts to segregate the Synechococcus sp. strain PCC
7002 gene under different light intensities petJ were also unsuccessful. These results
support the idea that the petJ1 gene is essential to Synechococcus sp. PCC 7002.
139
Figure 24. Attempted interposon mutagenesis of the petJ1 gene. (A) Physical map
of the petJ1 gene and surrounding partial reading frames. The aphII gene, which confers
kanamycin resistance, was digested with the blunt cutting enzyme HincII. This HincII
fragment was inserted into a unique, blunt BstXI site within the coding region of petJ1.
The plasmid, pPETJ1 was used to transform the wild-type strain of Synechococcus sp.
PCC 7002. (B) DNA gel blot hybridization suggests that the gene encoding cytochrome
c6 may be an essential gene in Synechococcus sp. strain PCC 7002. Lanes 1 and 2
represent genomic DNA from two different Synechococcus sp. strain PCC 7002
transformants digested with Hind III. The two hybridizing bands represent the wild-type
petJ1 gene (2.0 kb) and the petJ1 gene interrupted with the aphII gene encoding
kanamycin resistance inserted at the BstXI site (petJ1::aphII, 3.3 kb) indicative of the
merodiploid status of these alleles. Lane 3 represents genomic DNA digested with
HindIII and wild-type petJ1 gene (2.0 kb).
140
Hin
d II
I
Xb
a I
bif A hyp I
Bst
X I
petJ
Bst
X I
∆∆∆∆ petJ::aphII
aphII
100 bp
Hin
c II
Hin
c II
Bam
HI
Xb
a I
Bam
HI
Bg
l II
Bcl
I
Pst
I
Sp
h I
Xb
a I
A1 2 3
21.7
5.004.27
1.981.90
1.601.37
0.940.83
B
2.0 kb
3.3 kb
141
Figure 25. Northern blot analysis of the petJ1 gene. The petJ1 gene is transcribed as
a monocistronic transcript. Total RNA was isolated from wild-type Synechococcus sp.
PCC 7002 under standard conditions. The RNA was denatured, electrophoresed, and
transferred to a nylon membrane. The blot was probed with a petJ1 specific probe
generated by the primers in Figure 21. The size of the petJ1 transcript (450 bp) was
estimated based on migration of the ribosomal RNA. Sizes of the ribosomal RNA
species are given on the left.
(knt)
2.31.5
0.5
2.8
petJ1 mRNA transcript
142
3.4.3 Attempted functional substitution of the Synechococcus sp.
strain PCC 7002 petJ1 gene with the petE and petJ genes from
Synechocystis sp. strain PCC 6803
Because attempts to segregate the petJ1::aphII allele in Synechococcus sp. PCC
7002 failed under the conditions tested, plasmids were made to determine whether the
functional analog of cytochrome c6, plastocyanin, from Synechocystis sp. strain PCC
6803, could replace the Synechococcus sp. strain PCC 7002 cytochrome c6, and thus
induce complete segregation of the petJ1::aphII allele. The petE gene, which encodes
plastocyanin, was amplified by PCR from Synechocystis sp. strain PCC 6803 genomic
DNA using the primers (5' petE): 5'-ATC GCC AAG AAA CAT GTC TAA AAA G-3'
and (3' petE): 5'-ATA GGC CTT GCC ATT GCG AGA AGC-3'. The PCR product was
isolated and digested with AflIII and StuI. This AflIII-StuI DNA fragment was inserted
into the pSE280 vector at the AflIII compatible NcoI site so that the gene would be in the
correct reading frame for expression and under the control of the E. coli trc (trp-lac)
promoter (Brosius, 1989). It has been demonstrated previously that introduced
heterologous genes can be expressed with the trc promoter in cyanobacteria (Geerts et al.,
1995). The 3' end of the PCR product was engineered to have a StuI site for insertion
into pSE280. This plasmid containing the Synechocystis sp. PCC 6803 petE gene was
inserted into the platform vector pLAT2. The pLAT2 vector contains a marker for
erythromycin resistance and a polylinker between Synechococcus sp. PCC 7002 argE3
gene flanking sequences. The argE3 gene has been determined to be a neutral site within
the Synechococcus sp. PCC 7002 genome (Muñiz, 1993). The pLAT2-petEtrc plasmid
143
was then transformed into the petJ1/petJ1::aphII merodiploid strain of
Synechococcus sp. PCC 7002 (Figure 26). Erythromycin/kanamycin resistant
transformants were selected and grown in liquid media with the appropriate antibiotics.
Chromosomal DNA was isolated from the cyanobacterial transformants as described in
the Materials and Methods and segregation of the petJ1::aphII allele was assayed by
Southern blot hybridization. Results indicate that the pLAT2-petEtrc plasmid had been
successfully inserted into the Synechococcus sp. PCC 7002 neutral site in all strains
tested (Figure 26). The inserted Synechocystis sp. PCC 6803 petE plasmids failed to
induce full segregation in the petJ1/petJ1::aphII merodiploid strain of Synechococcus sp.
PCC 7002. Even after transformation of this merodiploid strain with the pLAT2-petEtrc,
the wild-type copy of the petJ1 gene could not be segregated under either
photoautotrophic or photoheterotrophic growth conditions (Figure 27).
A similar strategy was used to incorporate the petJ gene from Synechocystis sp.
PCC 6803 into the Synechococcus sp. PCC 7002 petJ1/petJ1::aphII strain. The primer
6803 c6.1 (5'-CCT TGA AAG GAG AAC ACA TGT TTA AAT TAAT TC-3')
containing the NcoI-compatible AflIII site (underlined) and the primer 6803 c6.2 (5'-CTC
CCC AGC CCC AGA GGA TCC CTG ATC AAA-3') containing a BamHI site
(underlined) were used to amplify the petJ gene from Synechocystis sp. PCC 6803. The
resulting PCR product was digested and cloned into the pSE280 vector as described for
the Synechocystis sp. PCC 6803 petE/pSE280 plasmid. The Synechocystis sp. PCC 6803
petJ/pSE280 plasmid was digested with SspI and inserted into the platform vector
pLAT3. (Figure 28A).
144
Figure 26. (A) Schematic of the petJ1::aphII rescue strategy. This is an outline of the
strategy to inactivate the petJ1 gene from the Synechococcus sp. PCC 7002 merodiploid
strain by functionally replacing the Synechococcus sp. PCC 7002 PetJ1 (cyt c6) with the
Synechocystis sp. PCC 6803 PetE (plastocyanin) protein. The Synechocystis sp. PCC
6803 petE gene was cloned into the expression vector pSE280 so that petE expression
was regulated by the E. coli trc promoter. The plasmid pSE280-petEtrc was digested
with SspI and BamHI, the SspI-BamHI petEtrc-containing fragment was inserted into the
platform vector pLAT2. The pLAT2-petEtrc plasmid was used transform the
petJ1/petJ1::aphII merodiploid strain of Synechococcus sp. PCC 7002. (B) Southern blot
analysis of the petEtrc integration into the Synechococcus sp. PCC 7002 chromosome..
The blot was hybridized with a 32-P-labelled Synechocystis sp. PCC 6803 petE PCR
product. Lanes 1 and 2: the total DNA isolated from kanamycin/erythromycin resistant
strain transformed with the pLAT2-petEtrc plasmid digested with EcoRI. Lanes 3 and 4:
total DNA isolated from kanamycin/erythromycin resistant strain transformed with the
pLAT2 plasmid digested with EcoRI. Lane 5: pSE280-petEtrc digested with SspI. Lane
6: petE PCR product. Arrows adjacent to the autoradiogram indicate the Synechocystis
sp. PCC 6803 petE–specific cross-hybridizing DNA fragments.
145
Ssp IHind IIIHinc IISsp I
Ptrc petE
Ap r
7002 ARG
ORI
7002 ARG
Ptrc
pLAT2/petE trc9697 bp Eco RI
Not I
Not I
Apr
Em r
Eco RIpetE
Sst
I
Kpn
I
Bam
HI
Xba
I
Sal
I/A
cc I/
Hin
c II
Pst
I
Sph
I
Not I
Eco RI
Ap r
7002 ARG
ORI
7002 ARG pLAT28500 bp
Apr
Not I
Em rEco RI
Sst
I
Kpn
I
Bam
HI
Xba
I
Sal
I/A
cc I/
Hin
c II
Pst
I S
ph I
Hin
d III
digest pSE280 withSsp I
digest pLAT2with Hind III
Klenow fill-in
Ligate with T4 ligase
Eco RI
7002 ARG 7002 ARGPtrc
petE
Em r
Sst
I
Kpn
I
Bam
HI
Xba
I
Sal
I/A
cc I/
Hin
c II
Pst
I
Sph
I
7002 chromosome7002 chromosome
1 2 3 4 5 6
3.1 kb
1.0 kb
0.4 kb
146
Figure 27. Attempted displacement of the petJ1 gene in a Synechococcus sp. PCC
7002 petJ1/petJ1::aphII merodiploid strain with the pLAT2-petEtrc platform vector.
The petJ1::aphII/petJ1 merodiploid strain of Synechococcus sp. PCC 7002 was
transformed with the pLAT2-petEtrc plasmid. Kanamycin/erythromycin resistant
cyanobacterial colonies were selected and grown in liquid culture in the presence or
absence of 5µM Cu+ and in the presence or absence of 10 mM glycerol. Genomic DNA
was isolated from these cultures, digested with HindIII and transferred to a nylon
membrane and probed with a 32P-labelled Synechococcus sp. PCC 7002 petJ1-specific
probe. Genetic backgrounds of isolated DNA are as follows: Lanes 1 and 2:
petJ1/petJ1::aphII/pLAT2-petEtrc . Lanes 3 and 4: petJ1/petJ::aphII, not transformed
with pLAT2-petEtrc . Lane 5: wild-type. Arrows point out Synechococcus sp. PCC
7002 petJ1-specific hybridization with the radiolabelled probe.
148
Figure 28. Insertion of the Synechocystis sp. PCC 6803 petJ gene into the petJ1::aphII
merodiploid strain of Synechococcus sp. PCC 7002. (A) The pLAT3/Synechocystis sp.
PCC 6803 petJ plasmid. The petJ gene from Synechocystis sp. PCC 6803 was placed
into the multi-cloning site interrupting the amtA gene in the pLAT3 plasmid. This
plasmid was used to transform the petJ1/petJ1::aphII merodiploid strain of
Synechococcus sp. PCC 7002. Restriction sites in brackets ( ) represent sites that were
eliminated when the plasmid was made. (B) Southern blot analysis of the insertion of the
Synechocystis sp. PCC 6803 petJ gene into the petJ1/petJ1::aphII merodiploid strain of
Synechococcus sp. PCC 7002. Total DNA isolated from several independent
kanamycin/spectinomycin resistant transformants was digested with PstI and transferred
to a nylon membrane. The blot was probed with a 32P-labelled pLAT3/Synechocystis sp.
PCC 6803 petJ plasmid. Lanes 1-4, Genomic DNA isolated from the petJ1/petJ1::aphII
merodiploid strain of Synechococcus sp. PCC 7002 transformed with
pLAT3/Synechocystis sp. PCC 6803. Lane 5, genomic DNA isolated from the
Synechococcus sp. PCC 7002 wild-type strain. Lane 6, pLAT3 plasmid. Lane 7,
pLAT3/Synechocystis sp. PCC 6803 petJ plasmid.
149
Sph
I
Spe
I
Bam
H I
Bam
H I
Hin
d III
Hin
d III
Sma
I
Pst
I
Eco
RI
Hin
d III
Cla
II
(Hin
c II
)
amtA
Pst
I
Eco
RV/S
sp I
Hin
c II
Bam
HI
Eco
RV/S
sp I
petJ
Nco
I
amtA
1 2 3 4 5 6 7
1.7 kb
1.1 kb
WT∆amtA
6803 petJ
A.
B.
(Xba
I)
150
The pLAT3::petJtrc plasmid was used to transform the petJ1/petJ1::aphII
merodiploid strain of Synechococcus sp. PCC 7002 (Figure 28B). Figure 29 shows that,
even with the Synechocystis sp. PCC 6803 petJ gene, the Synechococcus sp. PCC 7002
petJ1/petJ1::aphII merodiploid strain did not segregate. Because there was no
segregation in either attempts at functional substitution of the petJ1 gene from
Synechococcus sp. PCC 7002 with the petE or petJ gene from Synechocystis sp. PCC
6803, the transcription of the Synechocystis sp. PCC 6803 petE and petJ genes was
assayed by RNA blot analysis (Figure 30 and Figure 31). The results indicated that
both the Synechocystis sp. PCC 6803 petE and petJ genes are transcribed in
Synechococcus sp. PCC 7002.
To insure that a protein was being translated from the petE mRNA transcript, the
production of the Synechocystis sp. PCC 6803 plastocyanin in the Synechococcus sp.
PCC 7002 petJ1/petJ1::aphII merodiploid strain was assayed by immunoblot analysis
with anti-Synechocystis sp. PCC 6803 plastocyanin antiserum (a gift from Dr. John
Whitmarsh, University of Chicago-Urbana). Figure 32 shows that the Synechocystis sp.
PCC 6803 plastocyanin is produced and processed to the correct mature size in
Synechococcus sp. PCC 7002. These results indicate that Synechococcus sp. PCC 7002 is
capable of producing and processing foreign proteins but that neither the Synechocystis
sp. PCC 6803 plastocyanin or cytochrome c6 can functionally substitute for the native
Synechococcus sp. PCC 7002 cytochrome c6 protein.
151
Figure 29. The petJ gene from Synechocystis sp. PCC 6803 cannot functionally
replace the petJ gene from Synechococcus sp. PCC 7002. The petJ1/petJ1::aphII
merodiploid/pLAT3 6803 petJ strain of Synechococcus sp. PCC 7002 was grown under
various light and photoheterotrophic conditions and total DNA was isolated from the
cells. DNA isolated from all strains was digested with HindIII. A Synechococcus sp.
PCC 7002 gene specific probe for petJ1 was used to check the segregation of the
petJ1::aphII and the wild-type petJ1 alleles. Lanes 1-3: genomic DNA isolated from the
petJ1/petJ1::aphII merodiploid/pLAT3 6803 petJ strain of Synechococcus sp. PCC 7002
grown under the different conditions listed. Lane 4: genomic DNA isolated from the
wild-type Synechococcus sp. PCC 7002 strain. Arrows point out Synechococcus sp. PCC
7002 petJ1-specific hybridization with the 32P-radiolabelled probe.
petJ1::aphII (3.4 kb)
petJ1 (2.0 kb)
10 mM glycerol250 µE m-2 s-1
1 2 3 4
++
-+
+-
+-
152
Figure 30. RNA blot hybridization analysis of the Synechocystis sp. PCC 6803 petE
gene in the petJ1/petJ1::aphII merodiploid strain of Synechococcus sp. PCC 7002. Total
RNA was isolated from merodiploid and wild-type strains. The RNA was
electrophoresed and transferred to a nylon membrane. (A) The blot was probed first with
a 32P-labelled Synechocystis sp. PCC 6803 petE gene specific probe. (B) As a control, the
membrane was stripped of radioactivity and then hybridized with a 32P-labelled
Synechococcus sp. PCC 7002 petJ1 gene specific probe. Sizes of the ribosomal RNA
bands are shown on the right.
153
2.82.31.5
0.5
wild-ty
pe
petJ
1::a
phII,
pet
J1, 6
803
petE
petJ
1::a
phII,
pet
J1, 6
803
petJ
wild-ty
pe
petJ
1::a
phII,
pet
J1, 6
803
petE
petJ
1::a
phII,
pet
J1, 6
803
petJ
A. B.
154
Figure 31. RNA blot hybridization analysis of the Synechocystis sp. PCC 6803 petJ
in the petJ1::aphII merodiploid strain of Synechococcus sp. PCC 7002. Total RNA was
isolated from mutant and wild-type strains. The RNA was electrophoresed and
transferred to a nylon membrane. (A) The blot was probed first with a 32P-labelled
Synechocystis sp. PCC 6803 petJ gene specific probe. (B) As a control, the membrane
was stripped of radioactivity and then hybridized with a 32P-labelled Synechococcus sp.
PCC 7002 petJ1 gene specific probe. Sizes of the ribosomal RNA markers are shown on
the right.
155
A.
2.82.31.5
0.5
B.
wild-ty
pe
wild-ty
pe
petJ
1::a
phII,
pet
J1, 6
803
petE
petJ
1::a
phII,
pet
J1, 6
803
petJ
wild-ty
pe
wild-ty
pe
petJ
1::a
phII,
pet
J1, 6
803
petE
petJ
1::a
phII,
pet
J1, 6
803
petJ
156
Figure 32. Immunoblot analysis of the Synechocystis sp. PCC 6803 PetE
(plastocyanin) protein produced in the petJ1::aphII/petJ1 merodiploid strain of
Synechococcus sp. PCC 7002. Immunoblot analysis of crude protein extracts from
various cyanobacterial strains was performed. 40 µg of total protein was isolated from
whole cells of the following strains and electrophoresed on a SDS polyacrylamide gel:
Lane 1, wild-type Synechococcus sp. PCC 7002, Lane 2, wild-type Synechocystis sp.
PCC 6803, and Lane 3, Synechococcus sp. PCC 7002 petJ::aphII/petJ1, pLAT2-petEtrc
,. The crude extracts were subjected to SDS-PAGE and blotted onto a nitrocellulose
membrane. The membrane was probed with polyclonal rabbit antiserum against
Synechocystis sp. PCC 6803 plastocyanin. Plastocyanin was detected in the
Synechococcus sp. PCC 7002 petJ1::aphII/petJ1/pLAT2-petEtrc merodiploid strain and
in the whole-cell extracts from the Synechocystis sp. PCC 6803 wild-type strain.
1 2 3
PC
157
3.4.4 petJ-2
The petJ2 gene was identified on a 3.7-kb BamHI-HincII fragment downstream of
the ccmK-1 gene, which encodes a protein involved in carbon uptake. Sequence analysis
of the Synechococcus sp. PCC 7002 petJ2 predicts a mature protein of 88 amino acids
and a pI of 9.29.
The Synechococcus sp. PCC 7002 PetJ2 amino acid sequence was subjected to
analysis with the SignalP server (http://www.cbs.dtu.dk/services/SignalP/index.html) to
determine if there were a predicted signal sequence (Nielsen et al., 1997). Analysis of the
sequence using the algorithms for the prediction of gram-negative and gram-positive
signal sequences was used to identify a potential signal sequence AFG-AD, where (-) is
the predicted cleavage site. The potential signal peptide is 27 amino acids
(MNKRLVQVIVFVMIVLLLVPLLATPAFG) at the N-terminus of the predicted
protein.
A ClustalW alignment of PetJ2 with cytochrome c6 proteins is shown in Figure
33. The PetJ2 protein from Synechococcus sp. PCC 7002 is 40% identical to and 55%
similar to the PetJ1 of Synechococcus sp. PCC 7002, 37% identical and 51% similar to
the PetJ protein from Synechocystis sp. PCC 6803 (accession number BAA17354)
(Kaneko et al., 1996), 47% identical and 59% similar to the PetJ protein from Anabaena
sp. PCC 7120 (accession number I39601) (Ghassemian et al., 1994), 48% identical to and
63% similar to the PetJ1 protein from Synechococcus sp. PCC 7942 (accession number
158
Figure 33. ClustalW alignment of the Synechococcus sp. PCC 7002 PetJ2 amino
acid sequence with PetJ amino acid sequences from other cyanobacteria. Gray high-
lighted regions indicate identical amino acids while lighter shading indicates conserved
amino acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. The consensus sequence is shown beneath all other sequences. Percent
identity and percent conserved amino acids determined by the ClustalW alignment tool
(Thompson et al., 1994b).
7002 PetJ27002 PetJ16803 PetJ7120 PetJS. maxima PetJ
10 20 30 40 50M N K R L V Q V I V F V M I V L L L V P L L A T P A F G A D L D Q G A Q I F E A H C A
M K K L L A I A L T V L A T V F A F G T P A F A A D A A A G A Q V F A A N C AM F K L F N Q A S R I F F G I A L P C L I F L G G I F S L G N T A L A A D L A H G K A I F A G N C A
M K K I F S L V L L G I A L F T F A F S S P A L A A D S V N G A K I F S A N C AG D V A A G A S V F S A N C A
. . L . L . F . . P A A A D . A G A . I F . A N C A
7002 PetJ27002 PetJ16803 PetJ7120 PetJS. maxima PetJ
60 70 80 90 100G C H L N G G N I V R R G K N L K K R A M A K N G Y T S - - - - - - - V E A I A N L V T Q G K G N MA C H A G G N N A V M P T K T L K A D A L K T Y L A G Y K D G S K S L E E A V A Y Q V T N G Q G A MA C H N G G L N A I N P S K T L K M A D L E A N G K N S - - - - - - V A A I V A Q I T N - G N G A MS C H A G G K N L V Q A Q K T L K K A D L E K Y G M Y S - - - - - - - A E A I I A Q V T N G K N A MA C H M G G R N V I V A N K T L S K S D L A K Y L K G F D D - - - D A V A A V A Y Q V T N G K N A MA C H G G N . V K T L K K . D L K Y G . S E A V A Q V T N G K G A M
7002 PetJ27002 PetJ16803 PetJ7120 PetJS. maxima PetJ
110 120 130 140 150S A Y G D K L S S E Q I Q A V S Q Y V L Q Q S Q T D - W K SP A F G G R L S D A D I A N V A A Y I A D Q A E N N K WP G F K G R I S D S D M E D V A A Y V L D Q A E K G - WP A F K G R L K P E Q I E D V A A Y V L G K A D A D - W KP G F N G R L S P K Q I E D V A A Y V V D Q A E K G - WP A F . G R L S Q I E D V A A Y V L D Q A E . W
159
P25935) (Laudenbach et al., 1990), and is 46% identical and 59% similar to the PetJ
protein from Anabaena sp. PCC 7937 (accession number P28597) (Bovy et al., 1992).
3.4.5 Interposon mutagenesis of the petJ2 gene in Synechococcus sp. PCC
7002
The petJ2 gene was interrupted by the insertion of a BamHI fragment containing
the aphII gene, which confers kanamycin resistance, into a unique BglII site within the
coding sequence of the petJ2 open reading frame. The segregation of the petJ2::aphII
and petJ2 alleles was checked by PCR analysis (Figure 34). A 0.4-kb PCR, which
corresponds to the wild-type copy of petJ2, product was produced when wild-type
Synechococcus sp. PCC 7002 chromosomal DNA was used as the template. In strains
transformed with the pPETJ2 plasmid, a PCR product of 1.7-kb is produced that
corresponds to the size expected for the insertionally inactivated petJ2::aphII allele and
no product from the wild-type copy of the petJ2 gene is observed (Figure 34). This
analysis reveals that the wild-type copy was fully replaced by the petJ2::aphII mutant
allele. Therefore, the petJ2 gene does not have an essential function in electron transport
under normal, photoautotrophic growth conditions in Synechococcus sp. PCC 7002.
160
Figure 34. Interposon mutagenesis of the petJ2 gene. (A) Physical map of the
Synechococcus sp. PCC 7002 petJ2 gene. The petJ2 gene was interrupted by inserting a
BamHI fragment containing the aphII gene into a unique BglII site within the coding
sequence of petJ2. The plasmid pPETJ2 was used to transform the wild-type strain of
Synechococcus sp. PCC 7002. (B) PCR analysis of the petJ2 locus. Primers 5C62 (5’-
TCC CTG CCC GAA TTA CCG A-3’) and 3C62 (5’-TTA AGA TTT CCA GTC GGT
TTG GGA TTG C-3’) were used to amplify the petJ2 gene from genomic DNA isolated
from wild-type and kanamycin-resistant strains of Synechococcus sp. PCC 7002. The
position of the primers is indicated in (A). Two PCR products were produced, a 0.4 kb
product corresponding to the wild-type petJ2 allele and a 1.7 kb product corresponding to
the petJ2::aphII allele.
161
A. B.
petJ2
::aph
II
wild ty
pe
λHE
petJ-2
aphII
petJ-2
Hin
cII
Hin
cII
Bgl
II
Ssp
IS
spI
Sph
I
Pst
I
Bgl
II
Bcl
I100 bp
1.7 kb
0.4 kb
162
3.4.6 Growth analysis of petJ2::aphII
The doubling time of the petJ2::aphII strain of Synechococcus sp. PCC 7002
under standard growth conditions was 4.0 ± 0.6 hours, which is roughly the same
doubling time as for the wild-type strain. In the related species, Synechococcus sp. PCC
7942, a deletion in the petE gene exacerbates a chilling effect on photoinhibition (Clarke
and Campbell, 1996). By comparing the growth rates of wild-type and mutant strains of
Synechococcus sp. PCC 7002 when exposed to chilling stress, it was hoped that any
exacerbations of photoinhibition in mutant strains would translate to a growth-limiting
phenotype. Therefore, attempts were made to characterize the mutant strain by exposing
the strain to a temperature-shift from 38°C to 22°C. However, the results of the
temperature-shift from 38°C to 22°C (Figure 35) reveal that there is no significant
difference between the growth rates after a temperature-shift from 38°C to 22°C of the
wild-type strain and petJ2::aphII strain of Synechococcus sp. PCC 7002.
163
Figure 35. Temperature-shift effects on wild-type and petJ2::aphII strains of
Synechococcus sp. PCC 7002. Exponentially growing cells were either maintained at
38°C or transferred to a 22°C water bath. (A) Temperature-shift effects on the
petJ2::aphII strain of Synechococcus sp. PCC 7002. The petJ2::aphII cells were grown
at 38°C () and at 22°C (). (B) Temperature-shift effects on the wild-type strain of
Synechococcus sp. PCC 7002. Wild-type cells grown at 38°C () and at 22°C (). The
data shown are from a single experiment, which was repeated three times, and the results
were consistent and reproducible.
164
0.01
0.1
1
10
OD
550
0 10 20 30 40 50
Time (hours)
∆petJ2 (22˚C)
∆petJ2 (38˚C)
OD
550
0.01
0.1
1
10
0 10 20 30 40 50
Time (hours)
WT (22˚C)WT(38˚C)
A
B
165
3.4.7 cytM
The cytM gene, encoding cytochrome cM was originally identified in
Synechocystis sp. PCC 6803 by Malakhov et al. (1994). The function of this cytochrome
is unknown, but it has been proposed to be a substitute for PetE or PetJ in cells grown
under high-light or chilling-stress conditions in Synechocystis sp. PCC 6803 (Malakhov
et al., 1999). The cytM gene from Synechococcus sp. PCC 7002 was identified on contig
638 from the ongoing Synechococcus sp. PCC 7002 genome project. A comparison of
the gene organization around the cytM genes from Synechococcus sp. PCC 7002 and
Synechocystis sp. PCC 6803 is shown in Figure 36. The genes flanking the cytM genes
are very different between Synechococcus sp. PCC 7002 and Synechocystis sp. PCC
6803, and the surrounding genes do not give any hints as to the function of the cytM gene
or its gene product. A ClustalW alignment of cytochrome cM proteins is shown in Figure
37. The results reveal that the CytM proteins have a high degree of sequence similarity
among cyanobacterial species.
The Synechococcus sp. PCC 7002 CytM amino acid sequence was subjected to
analysis with the SignalP server (http://www.cbs.dtu.dk/services/SignalP/index.html) to
determine if there were a predicted signal sequence (Nielsen et al., 1997). Analysis of the
sequence using the algorithms for the prediction of gram-negative and gram-positive
signal sequences was used to identify a potential signal sequence TQA-SD, where (-) is
the cleavage site. The potential signal peptide is 38 amino acids
166
Figure 36. Gene organization around the cytM gene of Synechococcus sp. PCC 7002
and Synechocystis sp. PCC 6803. Arrows represent direction of transcription and size of
genes. Maps are individually scaled and numbers below each represent bp. Gene names
and predicted product names are indicated above genes (arrows). Restriction sites are as
indicated on maps.
167
300 600 900 1200 1500 1800 2100 2400 2700
sll1704 csgA (cell-cell signaling factor C)
slr1215
sll1434-penicillin binding pr… cytM
NcoI
StyI
PvuI
StyIPvuI
PvuI
PvuIDraI
PvuI
BglII
XbaI
StyI
PvuI
PvuI
StyI
PvuI
StyI PvuI
StyI
StyI
Synechococcus sp. PCC 7002 cytM restriction map
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600
slr1351 (murF) s l r1353rpl9 (50S ribosomal protein L9)
cy tM
SspI
BglII
NcoI
StyI
StuI
PvuI
PvuIPvuI
AvaI
PvuIPvuI NcoI
StyIStyI
PvuIStyI
DraI
StyI
StyI
PvuI
DraI
KpnI
DraI
PvuI
EcoRV
StyI
EcoRV
HincII
NcoI
StyI NcoI
StyI
Synechococcus sp. PCC 6803 cytM restriction map
168
Figure 37. ClustalW alignment of CytM proteins from cyanobacteria. Gray high-
lighted regions indicate identical amino acids while lighter shading indicates conserved
amino acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. The alignment of the CytM proteins was determined by the ClustalW
alignment tool (Thompson et al., 1994b).
CytM ClustalW Amino Acid Alignment
7002 CytMA. variabilis CytM7942 CytM6803 CytM
10 20 30 40 50V A N S S E L S V P N F S K L L F V I I L V L A I A A L GM D N Q I T K P E I L I Q R I A L V A L V I L L A I P L G
M I I L R H V L Q T T D S L S P A I A S A V E Q S L S K S I E S R A A Q T L G WL L A T A V M V A IM A P V I E K S P T V A T V N A S P T G I WI M A G I V S L V I L
. . . S . . . . . . . . . . . . . . .
7002 CytMA. variabilis CytM7942 CytM6803 CytM
60 70 80 90 100I F G V Y S T Q A S D P Y I Q Q V L A L Q G D E L R G N A I F Q I N C A G C H G P Q A D G N V G P SF F G V Q L V K A S D P Y V K S V L A MK G D P I Q G H A I F Q I N C A G C H G L E A D G R V G P SG L V V T L I R P A D P Y V S T V L N L P G N A E R G Q A I F Q I N C A G C H G P E G R G L V G P DA V A L F S F M N F D P Y V S Q V L A L K G D A D R G R A I F Q A N C A V C H G I Q A D G Y I G P S. . V D P Y V V L A L G D R G A I F Q I N C A G C H G A D G V G P S
7002 CytMA. variabilis CytM7942 CytM6803 CytM
110 120 130 140 150L R A V A Q R K S D V R L I Q Q V I S G K T P P M P K F Q P A P Q E M A D L L S Y L R T L NL Q A V S K R K S K Y G L I H Q V I S G D T P P M P K F Q P N T Q E M A D L L S F L E T LL A N V S N R K S R K D L I R Q V T T G E T P P M P K F Q P S P E T M A D L L R Y L E T LL WG V S Q R R S Q S H I I H Q V V S G Q T P P M P Q F E P N P Q E M A D L L N Y L K T L NL . V S . R K S L I . Q V . S G T P P M P K F Q P P Q E M A D L L Y L T L
169
(VANSSELSVPNFSKLLFVIILVLAIAALGIFGVYSTQA) at the N-terminus of the
predicted protein. Interestingly, neither the CytM protein from Synechocystis sp. PCC
6803 nor the CytM protein from Synechococcus sp. PCC 7942 are predicted to have a
signal sequence. However, analysis of the CytM protein from A. variabilis predicts that
the protein is processed at the VKA-SD motif, where (-) represents the processing site.
3.4.8 bcpA
The bcpA gene was identified as an open reading frame on a 2.5-kb EcoRI
fragment in close proximity to the glnB gene. Sequence analysis of the Synechococcus
sp. PCC 7002 bcpA gene predicts a protein of 106 amino acids with a pI of 6.06. A
ClustalW alignment of BcpA with the Anabaena sp. PCC 7120 BcpA sequence is shown
in Figure 38A. The predicted BcpA protein from Synechococcus sp. PCC 7002 is 62%
identical and 75% similar to the BcpA protein from Anabaena sp. PCC 7120 (Kazusa
DNA Research Institute, 2000). Figure 38B shows a ClustalW alignment of
cyanobacterial plastocyanin proteins with the BcpA proteins. The BcpA protein from
Synechococcus sp. PCC 7002 has low similarity to the plastocyanins from cyanobacteria,
with the most significant similarity occurring near the putative Cu-binding ligands in the
carboxy-terminus.
In order to assess whether the bcpA gene was being transcribed, RNA blot
hybridization was performed using RNA isolated from wild-type Synechococcus sp. PCC
7002 grown under standard conditions with petJ1, petJ2, and bcpA gene specific probes
170
Figure 38. (A) BcpA ClustalW alignment. The predicted BcpA amino acid
sequences of Synechococcus sp. PCC 7002 and Anabaena sp. PCC 7120 were aligned
using the ClustalW alignment program from MacVector v. 6.5. The consensus amino
acid sequence is located underneath. Gray high-lighted regions are identical amino acids
and lighter shading indicates a conserved amino acid. A dash is equivalent to a gap in the
sequence. Percent identity and percent conserved amino acids was determined by the
ClustalW program (Thompson et al., 1994a) and is shown in Table 4.
(B) BcpA/PetE ClustalW amino acid alignment. PetE and BcpA proteins were aligned
using the ClustalW alignment program from Mac Vector v. 6.5. The consensus amino
acid sequence is located underneath. Gray high-lighted regions are identical amino acids
and lighter shading indicates a conserved amino acid. A dash is equivalent to a gap in the
sequence.
171
BcpA ClustalW Amino Acid Alignments
7002-BcpA7120-BcpA
10 20 30 40 50M V F H K Y F I H G C R A L V L L G L I WF C L T P A A I A L P V P A Q Q P - - - - - - - L Q E M Q
M I S L L S P I V R Q I C V V F T L L L C F T F T N T N S V L A A K E S S D L L K Q P V S E I T. R . . . . C T . A . E
7002-BcpA7120-BcpA
60 70 80 90 100I H L G T T S G A L R F V P D Q L E F V A G Q R Y K L L L D N P S N Q K H Y F T A K D F A D T S WTV S L G N S A N E L K F E P N N L E L V A G K R Y L L H L N N P S Q L K H Y F T A K D F A D G I WT. L G . L . F P . L E V A G R Y L L N P S . K H Y F T A K D F A D WT
7002-BcpA7120-BcpA
110 120 130 140 150Q K V E A G Q G G S K R G D P R T R T Q A R A I A E WI L I P - Q K T G K F E L H C S V P G H A A AQ K V E A G K V E I K G A I H E L E L K P G A E A E WV L V A I K - P G K Y G L R C P I P G H T E AQ K V E A G . K . A A E W. L . G K . L . C . P G H A
7002-BcpA7120-BcpA
160 170 180 190 200G M V G T I Q V I A D SG M T G E I V I N PG M G I .
A.
B. BcpA/PetE ClustalW Amino Acid Alignments
7002-BcpA7120-BcpA1PetE-6803PetE-7942PetE-7120PetE-Spinacia oleracea
10 20 30 40 50M V F H K Y F I H G C R A L V L L G L I WF C L T P A A I A L P V P A Q Q P - - - - - - - L Q E M Q
M I S L L S P I V R Q I C V V F T L L L C F T F T N T N S V L A A K E S S D L L K Q P V S E I TM S K K F L T I L A G L L L V V S S F F L S V S P A A A A N - - - - - - - - - - A T
M K V L A S F A R R L S L F A V A A V L C V G S F F L S A A P A S A Q T - - - - - - - - - - V AM K L I A A S L R R L S L A V L T V L L V V S S F A V F T P S A S A E T - - - - - - - - - - Y T
V E. R . . . . . L . . . A .
7002-BcpA7120-BcpA1PetE-6803PetE-7942PetE-7120PetE-Spinacia oleracea
60 70 80 90 100I H L G T T S G A L R F V P D Q L E F V A G Q R Y K L L L D N P S N Q K H Y F T A K D F A D T S WTV S L G N S A N E L K F E P N N L E L V A G K R Y L L H L N N P S Q L K H Y F T A K D F A D G I WTV K M G S D S G A L V F E P S T V T I K A G E - E V K WV N N - K L S P H N I V F A A D G - - - - -I K M G A D N G M L A F E P S T I E I Q A G D - T V Q WV N N - K L A P H N V V V E G Q - - - - - -V K L G S D K G L L V F E P A K L T I K P G D - T V E F L N N - K V P P H N V V F D A A L N - - - -V L L G G D D G S L A F L P G D F S V A S G E - E I V F K N N - A G F P H N V V F D E D E I - - - -V . L G D G L . F E P . . A G . . . . N N . P H N . V .
7002-BcpA7120-BcpA1PetE-6803PetE-7942PetE-7120PetE-Spinacia oleracea
110 120 130 140 150Q K V E A G Q G G S K R G D P R T R T Q A R A I A E WI L I P - Q K T G K F E L H C S V P G H A A AQ K V E A G K V E I K G A I H E L E L K P G A E A E WV L V A I K - P G K Y G L R C P I P G H T E A- - - V D A D T A A K L S H K G L A F A A G E S F T S T F T E - - - P G T Y T Y Y C E P - - H R G A- - - - - - - - - P E L S H K D L A F S P G E T F E A T F S E - - - P G T Y T Y Y C E P - - H R G A- P A K S A D L A K S L S H K Q L L M S P G Q S T S T T F P A D A P A G E Y T F Y C E P - - H R G A- P S G V D A A K I S M S E E D L L N A P G E T Y K V T L T E - - - K G T Y K F Y C S P - - H Q G A
. . . S . L . P G . T G Y . Y C P H G A
7002-BcpA7120-BcpA1PetE-6803PetE-7942PetE-7120PetE-Spinacia oleracea
160 170 180 190 200G M V G T I Q V I A D SG M T G E I V I N PG M V G K V V V EG M V G K I V V QG M V G K I T V A GG M V G K V T V NG M V G K I V
172
(Figure 39). Results reveal that the petJ1 gene transcript is present at the highest
levels, petJ2 has a very low level of transcript, and the bcpA transcript is present at an
intermediate level compared to petJ1 and petJ2. Although bcpA is expressed in
Synechococcus sp. PCC 7002 (Figure 39), it is clear that the bcpA gene product cannot
functionally substitute for PetJ1 under normal growth conditions since attempts to
segregate the petJ1 and petJ1::aphII alleles were not successful. It is not surprising that
petJ2 also cannot functionally substitute for PetJ1 based its low transcript levels in
Synechococcus sp. PCC 7002 cells grown under normal conditions.
3.4.9 Interposon mutagenesis of the bcpA gene from Synechococcus
sp. PCC 7002
The bcpA gene was interrupted by the insertion of a 1.3-kb SmaI fragment
containing the aphII gene into a unique Klenow filled-in NcoI site within the coding
sequence of the bcpA gene. The resulting plasmid, pBCPA was used to transform the
wild-type strain of Synechococcus sp. PCC 7002 (Figure 40A). Segregation of the bcpA
allele was verified by PCR analysis (Figure 40B). This result indicates that the
bcpA::aphII locus has been fully segregated and that the bcpA gene is not essential for
the viability of Synechococcus sp. PCC 7002 under the conditions tested.
173
Figure 39. RNA transcript analysis of petJ1, petJ2, and bcpA. Total RNA was isolated
from wild-type Synechococcus sp. PCC 7002. Total RNA (20 µg) was loaded per lane in
a formaldehyde-agarose gel and electrophoresed as described in Materials and Methods,
transferred to a nylon membrane and probed with either a petJ1-specific, petJ2-specific,
or bcpA-specific 32P-labelled probe. Results shown are for three, identically treated,
separately probed membranes. Ribosomal RNA marker sizes (kb) are indicated at right.
2.82.31.5
0.5
petJ1petJ2
bcpA
174
Figure 40. Interposon mutagenesis of the Synechococcus sp. PCC 7002 bcpA gene. (A)
Physical map of the bcpA gene. The bcpA gene was interrupted by insertion of a 1.3-kb
SmaI fragment containing the aphII gene into a Klenow-treated, blunt NcoI site within
the bcpA coding sequence. Primers (5BcpA 5’-ATG GAG AAA ATC CAT GGT ATT
TCA TAA ATA-3’ and 3BcpA 5’-TTG AGG GAG CTT AGC TTA CGA ATC AGC
AAT-3’) were used to amplify the bcpA gene for PCR analysis. Primer positions are
indicated below the bcpA gene. (B) PCR analysis of the bcpA locus. PCR products are
as indicated on gel. The sizes of the PCR products are given on the left and right of the
figure.
175
Bam
HI
bcpA
Eco
RI
Hin
cII
Nco
I
Nco
I
aphII
wild
type
bcpA
::aph
IIbc
pA::a
phII
464 bp
λHE
1.8 kbp
A
B
176
3.4.10 Growth rate analysis of bcpA- strain of Synechococcus sp.
PCC 7002
Under normal growth conditions, the bcpA- strain of Synechococcus sp. PCC 7002
has a doubling time of 4.0 ± 0.8 hours, which is nearly identical to that of the wild-type.
Therefore, interruption of the bcpA locus has no effect on the growth of Synechococcus
sp. PCC 7002 under standard conditions. In a previous study by (Clarke and Campbell,
1996), it was shown that inactivation of the petE gene in Synechococcus sp. PCC 7942
experienced photoinhibition at low temperatures. In order to determine whether or not
inactivation of bcpA would have a similar effect in Synechococcus sp. PCC 7002, growth
of the wild-type and bcpA- strains of Synechococcus sp. PCC 7002 was monitored
following a temperature shift-down. Figure 41 shows the temperature-shift analysis of
the wild-type and bcpA- strains of Synechococcus sp. PCC 7002. These results indicate
that there is no difference in growth rate before and after a temperature-shift from 38°C
to 22°C in either the bcpA- or wild-type strains of Synechococcus sp. PCC 7002.
Therefore, the bcpA locus plays no role in low-temperature adaptation for Synechococcus
sp. PCC 7002.
3.4.11 Overproduction of the BcpA protein in E. coli.
The bcpA gene of Synechococcus sp. PCC 7002 predicts a novel blue-copper
protein that could have unique structural and biochemical traits. To investigate these
177
Figure 41. Temperature-shift effects on wild-type and bcpA::aphII strains of
Synechococcus sp. 7002. Exponentially growing cells were either maintained at 38°C or
transferred to a 22°C water bath. The bcpA::aphII cells were grown at 38°C () and
22°C () . (B) Temperature-shift effects on the wild-type strain of Synechococcus sp.
PCC 7002. Wild-type cells grown at 38°C () and 22°C (). The data shown are from
a single experiment, which was repeated three times, and the results were consistent and
reproducible.
178
0.01
0.1
1
10O
D55
0
0 10 20 30 40 50
Time (hours)
∆bcpA (22˚C)
∆bcpA (38˚C)O
D55
0
0.01
0.1
1
10
0 10 20 30 40 50
Time (hours)
WT (22˚C)WT(38˚C)
A
B
179
possibilities, two plasmids were made to overproduce the recombinant BcpA protein.
The first rBcpA plasmid was made using the following primers to amplify the bcpA gene
from Synechococcus sp. PCC 7002: BcpNcoI5’: 5’-CCC AGC GGC CAT GGC CTT
ACC CGT-3’ and BcpBlpI3': 5’- TTG AGG GAG CTT AGC TTA CGA ATC AGC
AAT –3’. The restriction sites for NcoI in the BcpNcoI5’ primer and BlpI for the
BcpBlpI3’ primer are underlined. These primers were used to amplify a PCR product
that eliminates the first 30 amino acids of the protein that encode a potential prokaryotic
signal sequence according to the rules of von Heijne (von Heijne, 1986). The PCR
product was digested with NcoI and BlpI and inserted into the pET16 vector for protein
overproduction.
The following primers were used to amplify the bcpA gene from Synechococcus
sp. PCC 7002: BcpNde5': 5'-CCT GCC CAA CAG CAT ATG CAG GAA ATG CAG-3'
(NdeI restriction site underlined) and the previously defined BcpBlpI3' primer for the
second rBcpA plasmid. The BcpNde5' primer was designed to truncate the amino
terminus of the BcpA at amino acid 39, and the primer also changed the leucine at
position 39 to a methionine. This portion of the protein was eliminated since the
remaining nine N-terminal amino acids were hydrophobic and possibly causing the
misfolding observed from the recombinant protein generated by the first plasmid. The
primers were used to amplify a product encoding a truncated bcpA gene, which was
digested with NdeI and BlpI. This DNA fragment was cloned into the NdeI and BlpI site
of the E. coli expression vector pET30C+. Overproduction of the protein led to the
formation of inclusion bodies. The rBcpA inclusion bodies were purified as described in
180
the Materials and Methods section. After purification, the proteins were checked by
SDS-PAGE (Figure 42). Attempts to refold the recombinant protein isolated from
inclusion bodies produced by either pET plasmid were unsuccessful under a number of
conditions (Table 7).
3.4.12 Immunoblot analysis of rBcpA
The rBcpA protein was purified as described in Materials and Methods and was
sent to the antibody facility at The Pennsylvania State University for the production of
rBcpA antisera. The anti-rBcpA antisera from two rabbits were assayed by immunoblot
analysis (Figure 43). Results indicate that the antisera cross-reacts with the rBcpA
protein and with a protein of similar size in the soluble extract from wild-type
Synechococcus sp. PCC 7002. The cross-reacting protein from Synechococcus sp. PCC
7002 is present in very low quantities and may represent the native BcpA protein. The
BcpA immunoblot results along with the result of the RNA blot analysis suggest that the
bcpA gene is transcribed and translated at low levels in wild-type Synechococcus sp. PCC
7002 grown under standard growth conditions.
181
Figure 42. Overproduction of the rBcpA protein. The plasmid pET30C+BCP was
used to transform E. coli BLR. Four transformants were picked to start cultures in 5 ml
LB kanamycin. These cultures were grown overnight and three cultures (15 ml) was
transferred to a flask containing LB medium (1L) containing 30 µg ml-1 kanamycin and
grown for 3 hours at 37°C and BcpA expression was induced by addition of 0.5 mM
IPTG at 3 hours and the cells were grown for another 3 hours at which time they were
harvested in 250 ml centrifuge bottles in a GSA rotor (4100×g) for 10 min at 4°C. The
cell pellets were washed with 50 mM Tris, pH 8.0 and re-centrifuged at 5000×g in a total
volume of 25 ml. The pellet was collected and resuspended in 25 ml 50 mM Tris, pH
8.0. These cells were broken by passage through an SLM-AMINCO French Pressure cell
as described in the Materials and Methods. The eluent was centrifuged at 3000×g for 10
min and an off-white pellet was isolated from the supernatant. The inclusion body pellet
was washed with 20 ml of 50 mM Tris, pH 8.0 and the soluble fraction was saved. The
inclusion body pellet was resuspended in 5 ml of 50 mM Tris, pH 8.0 and 25 µl was
added to 25 µl of 4×SDS loading buffer. The same volume was used for the soluble
fraction. Samples were heated to 70°C for 5 min and electrophoresed on an SDS-
polyacrylamide. Lanes 1 and 2: 5 µl of resuspended BCP inclusion body pellet. Lane 3:
5 µl of soluble E. coli BLR pET30C+BCP fraction. Lane 4: low MW ladder from
BioRAD. Lanes 5 and 6: 10 µl of resuspended BCP inclusion body pellet. Lanes 7 and
8: 10 µl of soluble E. coli BLR pET30C+BCP fraction. The rBcpA protein is indicated
with an arrow.
183
Table 7. Protein refolding conditions for rBcpA. Attempts to refold the recombinant
BcpA proteins were made in the following buffers: 50 mM Tris, pH 8.0; 50 mM Tris, pH
8.0, 100 mM NaCl; 50 mM Tris, pH 8.0, 150 mM NaCl, 50 mM MES, pH 6.0; 50 mM
MES, pH 6.0, 100 mM NaCl; and 5 mM phosphate, pH 7.0, 2 mM EDTA. Attempts
were done under the conditions described in the table. Fast dilution indicates that the
chaotrope-solubilized protein was diluted with buffer rapidly (50:1-100:1 (v/v)) and then
concentrated with an Amicon filter. Slow dilution indicates that the chaotrope-
solubilized protein was diluted slowly with buffer (0.1 ml h-1). Dialysis was carried out
overnight against 4000-fold dilution of buffer.
Chaotrope dilution method aerobic copper result
urea fast dilution yes yes precipitateurea fast dilution yes no precipitateurea fast dilution no yes precipitateurea fast dilution no no precipitateurea dialysis yes yes precipitateurea dialysis yes no precipitateurea dialysis no yes precipitateurea dialysis no no precipitateurea slow dilution yes yes precipitateurea slow dilution yes no precipitate
guanidine-HCl fast dilution yes yes precipitateguanidine-HCl fast dilution yes no precipitateguanidine-HCl fast dilution no yes precipitateguanidine-HCl fast dilution no no precipitateguanidine-HCl dialysis yes yes precipitateguanidine-HCl dialysis yes no precipitateguanidine-HCl slow dilution yes yes precipitateguanidine-HCl slow dilution yes no precipitate
184
Figure 43. Immunoblot analysis of using rBcpA antisera. Immunoblot analysis was
performed as described in Materials and Methods. rBcpA antisera from two rabbits (E
and F) were used for the experiment. The primary antibodies were diluted as described
above each blot. All gels were loaded in the same manner as follows: lane 1: 100 ng of
rBcpA, lane 2: 50 ng of rBcpA, lane 3: 100 µg protein from the membrane fraction of
Synechococcus sp. PCC 7002, lane 4: 100 µg protein from the soluble fraction of
Synechococcus sp. PCC 7002. The cross-hybridizing signal from rBcpA/BcpA is
indicated at right by the arrow (12.6 kDa).
185
1:1000 1:10,000
Antisera from Rabbit F
1 2 3 4 1 2 3 4
1:1000 1:10,000
Antisera from Rabbit E
1 2 3 4 1 2 3 4
12.6 kDa
12.6 kDa
186
3.5 Heme-Copper Oxidases in Synechococcus sp. PCC 7002
Two separate gene clusters encoding putative heme-copper oxidases in
Synechococcus sp. PCC 7002 were identified in this study. One apparent operon
(ctaCIDIEI) represents the primary heme-copper cytochrome oxidase. The second
apparent operon (ctaCIIDIIEII) represents a secondary heme-copper quinol oxidase.
Unlike Synechocystis sp. PCC 6803, no evidence for the occurrence of a cydAB-type
quinol oxidase in Synechococcus sp. PCC 7002 was obtained.
3.5.1 Screening and Cloning of the ctaI and ctaII gene clusters from
Synechococcus sp. PCC 7002
A BamHI fragment of 4.5-kb was isolated from a partial genomic library of
Synechococcus sp. PCC 7002 DNA by cross-hybridization with a ctaDI PCR probe
amplified from Synechocystis sp. PCC 6803 genomic DNA. This clone contained the
sequence of the ctaDI gene. The Synechococcus sp. PCC 7002 ctaDII gene cluster was
found by screening a cosmid genomic library with a PCR product probe derived from the
slr2082 (Kaneko et al., 1996) open reading frame encoding the ctaDII gene in
Synechocystis sp. PCC 6803. The cosmid 2B1 was identified as a positive clone and a
3.5-kb HincII fragment was subcloned. Nucleotide sequence analysis of this 3.5-kb
HincII fragment revealed two open reading frames with homology to ctaCII and ctaDII
from Synechocystis sp. PCC 6803 as well as the open reading frames sll1485 and sll1486.
187
Primers were designed to sequence cosmid 2B1 outside of the 3.5-kb HincII
fragment. Sequencing downstream from ctaDII, it was possible to identify and sequence
the ctaEII gene. Primers were also made to generate a PCR product corresponding to the
Synechocystis sp. PCC 6803 cydA gene, which encodes one subunit of the cytochrome
bd-quinol oxidase, and the resulting PCR product was used to screen the Synechococcus
sp. PCC 7002 genomic library for the presence of a quinol oxidase. No cross-hybridizing
DNA fragments corresponding to the cydA gene from Synechocystis sp. PCC 6803 were
found in Synechococcus sp. PCC 7002 genomic DNA Southern blot hybridizations.
Moreover, no genes with similarity to the cydABDC genes have been found in the
genome sequencing project from Synechococcus sp. PCC 7002. These results indicate
that Synechococcus sp. PCC 7002 has two terminal oxidase operons, ctaCIDIEI and
ctaCIIDIIEII, but does not have appear to an operon encoding a CydAB-like quinol
oxidase.
A comparison of the gene organization for the ctaCIDIEI operon from
Synechococcus sp. PCC 7002 with the ctaCIDIEI operons from Synechocystis sp. PCC
6803 and Anabaena sp. PCC 7120 is shown in Figure 44. These data show that the gene
organization for the ctaCIDIEI operon is identical for all three cyanobacterial strains.
Figure 45 shows a comparison of the gene organization of the ctaCIIDIIEII gene clusters
from Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803, and Anabaena sp. PCC
7120. This figure reveals that the organization of the ctaCIIDIIEII gene cluster is differs
for the three cyanobacterial strains. In Synechococcus sp. PCC 7002, the ctaCIIDIIEII
genes occur in a single operon. However, arrangements of the ctaCII, ctaDII, and ctaEII
188
Figure 44. Gene organization of the ctaCIDIEI genes from cyanobacteria. The gene
organization of the ctaCIDIEI operons of Synechococcus sp. PCC 7002, Synechocystis
sp. PCC 6803, and Anabaena sp. PCC 7120 are depicted in this figure. Maps are
individually scaled and numbers below each represent bp. Gene names and predicted
product names are indicated above genes (arrows).
400 800 1200 1600 2000 2400 2800 3200 3600 4000 4400 4800
ctaDI ctaEIctaCI ORF8
600 1200 1800 2400 3000 3600 4200 4800 5400 6000 6600 7200 7800 8400
sll1084 ctaDI slr1140ctaCI nrtD (sll1082)ctaEI
trxA (slr1139)
ctaEI
700 1400 2100 2800 3500 4200 4900
ctaDIctaCIsll1898 slr1542
Cyanobacterial ctaI gene organization
Synechococcus sp. PCC7002
Synechocystis sp. PCC6803
Anabaena sp. PCC7120
189
Figure 45. Gene organization of the ctaCIIDIIEII genes from cyanobacteria. The
gene organization of the ctaCIIDIIEII apparent operon of Synechococcus sp. PCC 7002,
and the organization of the ctaCIIDIIEII genes from Synechocystis sp. PCC 6803, and
Anabaena sp. PCC 7120 are depicted in this figure. Maps are individually scaled and
numbers below each represent bp. Gene names and predicted product names are
indicated above genes (arrows).
500 1000 1500 2000 2500 3000 3500 4000 4500 5000
ctaDIIctaCIIsll1486 ctaEIIsll1485 sldI
Gene organization of cyanobacterial ctaII gene clusters
500 1000 1500 2000 2500 3000 3500 4000 4500 5000 5500 6000 6500 7000 7500 8000 8500 9000 9500 10000 10500
sll1987 (katG)sll0236 ctaDIIslr2079 sll1988tyrA ctaEIIslr2080ssr3532
500 1000 1500 2000 2500 3000 3500 4000 4500 5000 5500 6000
ctaDII ctaCIIsll1486sll1485uvrC (sll0865) ORF
500 1000 1500 2000 2500 3000 3500 4000 4500 5000 5500 6000
ptxB rpsD ctaEII
Synechococcus sp. PCC7002
Synechocystis sp. PCC6803
500 1000 1500 2000 2500 3000 3500 4000 4500 5000 5500 6000 6500 7000 7500
sll0813 (ctaCII)sll0816 (trpX)sll0817 sll0814sll0818 sll0815 slr0812slr1080 ssl1520
Anabaena sp. PCC7120
190
genes are different in the other cyanobacteria. The ctaDII and ctaEII are clustered
together in Synechocystis sp. PCC 6803, whereas the ctaCII gene is located elsewhere in
the genome of this organism. Preliminary analysis of the nucleotide sequence of contigs
from the Anabaena sp. PCC 7120 genome sequencing project (Kazusa DNA Research
Institute, 2000) reveals that the ctaCII and ctaDII genes are clustered together, whereas
the ctaEII gene is found elsewhere in the genome in Anabaena sp. PCC 7120.
3.5.2 Insertional Mutagenesis of c taDI and ctaDII from
Synechococcus sp. PCC 7002
To characterize the role of the two cytochrome oxidases in electron transport
pathways, interposon mutagenesis of the ctaDI and ctaDII genes was performed. A
unique BglII site in ctaDI was used to introduce a 1.3-kb BamHI fragment containing the
aphII gene, which confers kanamycin resistance. Two plasmids were made with the
aphII gene interrupting ctaDI. The first plasmid, pctaDI 1.1 had ctaDI interrupted with
the aphII gene in the same transcriptional orientation. The second plasmid, pctaDI 2.2
had the ctaDI interrupted with aphII in the opposite transcriptional orientation (Figure
46). Both pctaDI 1.1 and pctaDI 2.2 were independently used to transform wild-type
Synechococcus sp. PCC 7002 cells and ctaDII- strains (see below). Figure 46 shows that
all transformants were homozygous for the ctaDI::aphII locus. The ctaDII gene was
interrupted by insertion of a 2-kb SmaI fragment that had the aadA gene which confers
spectinomycin resistance (Prentki and Krisch, 1984) into a unique EcoRV site within the
191
Figure 46. Interposon mutagenesis of the ctaDI gene from Synechococcus sp. PCC
7002. (A) Physical map of the 4.5-kb BamHI fragment encoding the ctaCIDIEI gene
cluster of Synechococcus sp. strain PCC 7002 and disruption of the ctaDI locus by
interposon mutagenesis. Arrows indicate the direction of transcription and size of the
gene. The aphII gene, which encodes aminoglycoside 3'-phosphotransferase II and
confers kanamycin resistance, was inserted into a BglII site within the coding region of
ctaDI in both orientations to create the plasmids pctaDI 1.1 and pctaDI 2.2. These
plasmids were used to transform the wild-type and ctaDII- strains of Synechococcus sp.
PCC 7002. (B) PCR analysis of the ctaDI locus. PCR analysis was performed on
genomic DNA isolated from the wild-type and mutant strains of Synechococcus sp. PCC
7002 using the primers ctaDI.I1 and ctaDI.I2 as indicated in (A). The sizes of the PCR
products are indicated on the left. The 1.3-kb increase in size of the PCR product in the
insertion mutants corresponds to the size of the aphII insert. Lane 1 represents the 2.0-kb
wild-type ctaDI PCR product. Lane 2: Total DNA isolated from a kanamycin resistant
strain that had been transformed with pctaDI 1.1 was used as the template for PCR. The
3.3-kb ctaDI::aphII PCR product and no wild-type product are indicative of successful
segregation of the mutant and wild-type alleles. Lane 3: As lane 2 except that the wild-
type strain was originally transformed with pctaDI 2.2. Lanes 4 and 5: Total DNA
isolated from a kanamycin-spectinomycin strain had been transformed with either pctaDI
1.1 or pctaDI 2.2 were used as templates for PCR. This was done to check for reversion
to a wild-type ctaDI after transformation with the pctaDII Ω plasmid (Figure 48). The
results indicate that the double-mutant strains are still homozygous for ctaDI::aphII.
192
ctaDI(2
.2)::
aphI
I
ctaDII:
:Ω
wild ty
pecta
DI(1.1
)::ap
hII
ctaDI(2
.2)::
aphI
I
ctaDI(1
.1)::
aphI
I
ctaDII:
:Ω
2.0 kb3.3 kb
A.
sll1898
ctaCI
500 bp
ctaDI ctaEI
slr1999
slr1542
Bam
HI
Bam
HI
Eco
RV
Bgl
II
Hin
cII
Hin
cII
Hin
cII
Stu
I
Pst
I
ctaDI.I1 ctaDI.I2
aphII
aphII
Pst I
1.1
2.2
B.
193
ctaDII coding sequence. The resulting plasmid, pctaDII Ω was used to transform
wild-type Synechococcus sp. PCC 7002 as well as the ctaDI- strains of Synechococcus sp.
PCC 7002 (Figure 47A). Attempts to PCR amplify the ctaDII::Ω locus were
unsuccessful, thus the segregation of the ctaDII and ctaDII::Ω alleles was determined by
Southern blot hybridization analysis (Figure 47B). The results of the Southern blot
analysis indicate that the ctaDII::Ω locus was also completely segregated in all genetic
backgrounds that were transformed with pctaDII. These results indicate that the ctaDI
and ctaDII genes have been successfully interrupted and are not essential for viability
under standard growth conditions for Synechococcus sp. PCC 7002.
3.5.3 Expression of the cta gene clusters in Synechococcus sp. PCC
7002
To assess whether or not the cta genes were expressed in Synechococcus sp. PCC
7002, the level of mRNA and the size of the transcripts of the ctaI and ctaII apparent
operons were examined by Northern blot hybridization analysis. Specific DNA probes
were made for all cloned and sequenced cta genes (Table 8). A 3.5-kb RNA species was
detected for ctaCI, ctaDI, and ctaEI on the RNA blot (Figure 48). This indicates that
this gene cluster is transcribed as a polycistronic mRNA encoding all three genes. No
hybridization signal was detected by the ctaII-specific probes on RNA blots from wild-
type Synechococcus sp. PCC 7002, indicating that the level of mRNA transcript
194
Figure 47. Interposon mutagenesis of the ctaDII gene from Synechococcus sp. PCC
7002. (A) Physical map of the 3.5-kb HincII fragment encoding ctaCIIDII and disruption
of the ctaDII locus by interposon mutagenesis. Arrows indicate the direction of
transcription and size of the gene. The Ω fragment, conferring spectinomycin resistance,
was inserted into an EcoRV site within the coding region of ctaDII. The plasmid pctaDII
Ω was used to transform the wild-type and ctaDI- strains of Synechococcus sp. PCC
7002. Transformants were selected for spectinomycin resistance as described in
Materials and Methods. (B) Southern blot analysis of the ctaDII locus. Genomic DNA
was digested with HincII and probed with a 32P-labeled PCR product for ctaDII. The
sizes of the hybridizing DNA fragments are indicated on the left. Lane 1: Hybridizing
fragment of 3.5-kb corresponds to the size of the wild-type copy of ctaDII. Lane 2:
Total DNA isolated from a spectinomycin resistant transformant was digested with
HincII. The 2.0 kb increase in size of the hybridizing ctaDII specific fragment
corresponds to the insertion of the 2.0-kb Ω fragment which confers spectinomycin
resistance. The absence of a 3.5-kb hybridizing fragment is indicative of the full
segregation of the ctaDII and ctaDII::Ω alleles. Lanes 4 and 5: The ctaDI- strains made
previously (Figure 46) were transformed with pctaDII Ω. Total DNA isolated from the
kanamycin spectinomycin resistant transformants was digested with HincII. Results
indicate that the strains were homozygous for ctaDII::Ω.
195
00 bp
ctaDII ctaCII
Hin
cII
Hin
cII
Eco
RV
Nco
IN
coI
Nde
I
sll1486
Ω
Sph
I
Bam
HI
Hin
d III
Hin
d III
Bam
HI
.5 kb
.5 kb
ctaDI(2
.2)::
aphI
I
ctaDII:
:Ωcta
DI(1.1
)::ap
hII
ctaDII:
:Ω
wild ty
pecta
DII::Ω
.
.
196
Table 8. Cytochrome oxidase oligonucleotide list
Gene primer name nucleotide sequence (5’-3’) PCR
product size
(bp)
ctaCI ctaCI.1 GTG AAT ATT CCC AAT AGC ATC 929
ctaCI.2 CCC CAT CTC CTC GGC GTA GGC
ctaDI ctaDI.1 ATG AGT GAC GCG ACA ATA CAC 1133
ctaDI.2 TAG GTG TCA TGG ACA TGG
ctaEI ctaEI.1 TAC GGC GAT CGC CAC CGA 566
ctaEI.1 CAA GGC CGA ACA GGA TAA TTC
ctaCII ctaCII.1 CAC TTT CGG CGA TCG CCC TAC TTT TGG GGG 868
ctaCII.2 GGG ATG ATA ATT CAC CAC
ctaDII ctaDII.1 CCA TGA CCC AAG CTC CC 1257
ctaDII.2 CGG GAA CCA GTG GTA CAC CGC
ctaEII ctaEII.1 GAC TGC CAT CAA TGA AAC C 588
ctaEII.2 ATT GCC AGA GAT AAA TCA GCC
197
Figure 48. RNA blot hybridization analysis of the ctaCIDIEI gene cluster of
Synechococcus sp. PCC 7002. PCR products specific for the ctaEI, ctaDI and ctaCI
genes were made and used as probes for RNA blot hybridization analysis. Total RNA
(20µg) was electrophoresed per lane of the gel. The size of the major hybridizing RNA
species is indicated on the left. This 3.5-kb hybridizing fragment corresponds to the size
of ctaCIDIEI operon. Smaller, non-specific hybridizing species may represent
breakdown products of the main transcript. Sizes of the ribosomal RNAs are indicated
on the right.
199
accumulation for ctaII was below the detection limit of Northern blot hybridization
analysis.
Because the transcripts of the ctaCII, ctaDII, and ctaEII genes could not be
detected by Northern blot hybridization, attempts to detect the ctaII transcripts were
made using reverse transcriptase (RT)-PCR. ctaII gene specific primers and template
RNA isolated from wild-type Synechococcus sp. PCC 7002 cells were used for RT
reactions. Three different cDNA templates were synthesized from total RNA isolated
from wild-type Synechococcus sp. PCC 7002 using the following primers: ctaEII.2,
ctaDII.2, and ctaCII.2. Primer ctaEII.2 corresponds to the 3' end of the ctaEII gene.
Primer ctaDII.2 corresponds to a region approximately 300 nucleotides downstream from
the 3' end of the ctaDII gene. Primer ctaCII.2 corresponds to the 3' end of the ctaCII
gene. The RT reaction with primer ctaEII.2 produced the cDNA template ctaEII.2T. The
RT reaction with primer ctaDII.2 produced the cDNA template ctaDII.2T. The RT
reaction with primer ctaCII.2 produced the cDNA template ctaCII.2T. Primers
corresponding to the 5' ends of the ctaCIIDIIEII genes are labeled as follows: ctaEII.1 (5'
end of the ctaEII gene), ctaDII.1 (5' end of the ctaDII gene), and ctaCII.1 (5' end of the
ctaCII gene). These 5’ primers were used in conjunction with the 3' primers, ctaEII.2,
ctaDII.2, and ctaEII.2 for PCR amplifications of the cDNA templates. As a control, total
RNA used for the RT cDNA synthesis was used as a template for all PCR reactions. No
PCR products were produced when the total RNA was used as a template with any of the
primers . This indicates that all of the PCR products from the reactions with the cDNA
templates are the result of amplification of the cDNA template and not genomic DNA.
200
When the cDNA ctaEII.2T was used as a template and primers ctaEII.1 and ctaEII.2
were used for the subsequent PCR reaction a 589-bp product was detected by agarose gel
electrophoresis (Figure 49). When the cDNAs ctaEII.2T and ctaDII.2T were used as
templates and primers ctaDII.1 and ctaDII.2 were used for the subsequent PCR reaction,
a 1655-bp product which corresponds to the ctaDII gene, was detected by agarose gel
electrophoresis (Figure 49). When the cDNAs ctaEII.2T, ctaDII.2T and ctaCII.2T were
used as templates and primers ctaCII.1 and ctaCII.1 were used for the subsequent PCR to
amplify the RT-PCR products, an 868-bp product was detected by standard agarose gel
electrophoresis, which corresponds to the ctaCII gene (Figure 49). These results indicate
that the mRNA species covers the entire region of the ctaII gene cluster and that the three
genes are co-transcribed. However, the inability to detect this mRNA species by RNA
blot analysis indicates that the transcript abundance of the ctaII genes was low under
standard growth conditions for Synechococcus sp. PCC 7002.
201
Figure 49. RT-PCR analysis of the ctaII gene cluster of Synechococcus sp. PCC
7002. (A). Gene arrangement of the ctaII gene cluster. The small numbered arrows
below the physical map represent primers used for PCR amplification. Primer sequences
are given in Table 8. (B). cDNA products from the reverse transcriptase reactions.
Primer ctaEII.2 was used to make the ctaEII.2T cDNA product. Primer ctaDII.2 was
used to make the ctaDII.2T cDNA product. Primer ctaCII.2 was used to make the
ctaCII.2T cDNA product. These cDNA products were used as templates for PCR. (C)
Electrophoretic analysis of PCR products amplified from templates listed above the lanes.
Primers ctaCII.1 and ctaCII.2 were used for lanes 1 to 4. Primers ctaDII.1 and ctaDII.2
were used for lanes 5 to 7. Primers ctaEII.1 and ctaEII.2 were used for lanes 8 and 9.
202
ctaDIItaCIIll1486 taEIIsll1485 sldI
ctaCII.1 taEII.2ctaDII.2taDII.1 ctaEII.1taCII.2
cDNA ctaDII.2TctaEII.2T
ctaCII.2T
68655
589
p bp
8681655
589
ctaE
II.2T
ctaD
II.2T
ctaD
II.2T
ctaC
II.2T
ctaE
II.2T
ctaE
II.2T
7002
DNA
7002
DNA
7002
DNA
500 bp
868 bp 1655 bp 589 bp
203
3.5.4 Respiratory activity and oxygen evolution activity in Synechococcus
sp. PCC 7002 wild-type, ctaDI-, and ctaDII- strains
Table 9 shows respiratory rates of whole cells of the wild-type and various
mutant strains grown under standard growth conditions. Strains in which the ctaDI gene
was insertionally inactivated displayed a 3-fold decrease in oxygen uptake compared to
the wild-type strain, while the CtaDII-deficient strain had the same rate of oxygen uptake
as the wild-type strain rate. The ctaDI- ctaDII- double mutant exhibited a decrease in
respiratory rate that was essentially identical to the ctaDI- mutant strain when compared
to wild-type cells. Although no genetic evidence could be found for a Cyd AB quinol-
type oxidase, the enzyme could still be present in Synechococcus sp. PCC 7002. To
determine the existence of a possible third oxidase activity, a known inhibitor of terminal
oxidase activity, KCN was added. All strains were sensitive to the addition of KCN to a
final concentration of 1 mM, and only low rates of oxygen uptake were seen in the
presence of KCN. This level of oxygen uptake can be attributed to the amount of oxygen
consumed by the electrode itself, since this rate was observed even in the absence of live
cells during the assay. The oxygen uptake activity observed in the ctaDI- and ctaDI-
ctaDII- strains is within the range of error for oxygen consumed by the electrode. These
results indicate that under the conditions tested, ctaDI encodes a subunit of a functional
204
terminal oxidase responsible for the respiratory activity of the cell and the CtaII
enzyme has no significant oxygen uptake activity.
Table 9. Oxygen Consumption Rates of Photoautotrophically Grown Synechococcus sp.
PCC 7002 Strains in Darkness. Values shown represent the averages ± the standard
deviation of five independent experiments.
Strain oxygen uptake(µmol of O2 (mg of chl)-1 h-1)
no additions
oxygen uptake(µmol of O2 (mg of chl)-1 h-1)
+ 1mM KCN
oxygen uptake(µmol of O2 (mg of chl)-1 h-1)
KCN sensitive activitywild-type 24 ± 6 2.5 ± 0.2 21 ± 6
ctaDI- 8 ± 2 2.5 ± 0.4 5 ± 2
ctaDII- 21 ± 6 2.4 ± 0.2 18 ± 6
ctaDI- ctaDII- 7 ± 1 2.5 ± 0.1 4 ± 1
Differences in oxygen evolution activities were not significant between mutant
and wild-type strains (Table 10). For cells of all strains grown under 150 to 250 µE m-2
s-1 constant illumination, the oxygen evolving activities were virtually identical. There
was a marked decrease in oxygen evolving capacity in all strains when they were grown
under high to extremely high light intensities (700 to 4500 µE m-2 s -1) compared to cells
grown under lower to standard constant illumination (Table 10). The oxygen evolving
activity was not determined for the ctaDI- strain at 4.5 mE m-2 s-1 because the strain did
not grow under these conditions. The differences in oxygen evolving activities observed
205
Table 10. Doubling times, oxygen evolution rates and Fv’/Fm’ for wild-type, ctaDI-,
ctaDII-, and ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002 grown under various
light intensities. All values represent the averages ± standard deviations of 5 independent
experiments. inviable: cells were unable to grow under these conditions.
Strain LightIntensity
(µE m-2 s-1)
doublingtime
(hours)
oxygen evolution
(µmol of O2 (OD550nm)-1 h-1)
Fv’/Fm’
wild-type 150 4.0 ± 0.4 1394 ± 70 ND
ctaDI- 150 4.2 ± 0.4 1360 ± 40 ND
ctaDII- 150 4.1 ± 0.4 1460 ± 70 ND
ctaDI- ctaDII- 150 3.9 ± 0.4 1460 ± 90 ND
wild-type 250 3.6 ± 0.4 1428 ± 70 0.47 ± 0.1
ctaDI- 250 4.0 ± 0.4 1410 ± 90 0.38 ± 0.1
ctaDII- 250 3.8 ± 0.4 1440 ± 90 0.46 ± 0.1
ctaDI- ctaDII- 250 4.1 ± 0.4 1392 ± 90 0.33 ± 0.1
wild-type 700 4.0 ± 0.4 360 ± 15 ND
ctaDI- 700 4.0 ± 0.4 351 ± 15 ND
ctaDII- 700 4.0 ± 0.4 375 ± 30 ND
ctaDI- ctaDII- 700 4.0 ± 0.4 294 ± 60 ND
wild-type 4500 4.5 ± 0.4 340 ± 50 ND
ctaDI- 4500 inviable inviable ND
ctaDII- 4500 5.0 ± 0.6 339 ± 70 ND
ctaDI- ctaDII- 4500 5.0 ± 0.4 320 ± 50 ND
206
between cells grown under standard constant illumination versus the oxygen evolving
activities of cells grown under constant light stress correspond to an environmentally
stimulated response within the cyanobacterial cells that causes a change in ratio of the
photosystems as well as a change in the structural and functional activities of the
photosystems (see results for 77K fluorescence).
3.5.5 Growth analysis of Synechococcus sp. PCC 7002 wild-type,
ctaDI-, ctaDII-, and ctaDI- ctaDII- strains
The growth rates of the mutant and wild-type strains of Synechococcus sp. PCC
7002 were analyzed for cells grown under several different continuous light intensities
(150 µE m-2 s-1, 250 µE m-2 s-1, 700 µE m-2 s-1, and 4.5 mE m-2 s-1). Under moderately low
and normal light intensities (150 µE m-2 s-1 and 250 µE m-2 s -1) and under moderate light
stress (700 µE m-2 s-1), the mutant strains could grow exponentially and the doubling
times were nearly indistinguishable from the doubling time of the wild-type strain (Table
10). This indicates that under these conditions, cytochrome oxidase activity does not
significantly contribute to the growth physiology and energetics of the cells.
In order to examine the effects of extremely high light intensity on the
Synechococcus sp. PCC 7002 strains, the cells were grown under a very high light
intensity (4.5 mE m-2 s-1) while the temperature and CO2 concentration were held
constant. The growth yield of the ctaDI- strain was severely reduced compared to the
207
wild-type strain under these conditions (Figure 50). Additionally, although the
growth rates are slightly slowed for the wild-type, ctaDII- and ctaDI- ctaDII- strains, the
growth rate of the ctaDI- single mutant was drastically slowed when compared to the
wild-type strain (Figure 51B). These results indicate that, although cytochrome oxidase I
is dispensable for growth at light intensities from 150 µE m-2 s-1 to 700 µE m-2 s-1, it is
important when the cells are grown under very high light stress (4.5 mE m-2 s-1).
However, under the same light intensity stress (4.5 mE m-2 s-1), the ctaDII- and ctaDI-
ctaDII- strains had growth rates similar to the wild-type strain of Synechococcus sp. PCC
7002 (Figure 51ACD). These results suggest two things. First, the wild-type strain of
Synechococcus sp. PCC 7002 is able to tolerate extremely high light intensities as long as
the temperature is carefully maintained and high CO2 levels are provided. The second
conclusion is that the absence of ctaDII allows the cells to grow at 4.5 mE m-2 s-1 even in
the absence of ctaDI. Thus, the absence of ctaDII suppresses the phenotype observed
when ctaDI is inactivated alone.
208
Figure 50. Effect of extreme high light intensity on growth of Synechococcus sp.
PCC 7002 wild-type (PR6000) and ctaDI- cells. Cells were grown under standard
conditions until they reached exponential growth phase. The cells were then diluted to an
OD550 = 0.05 and grown under either standard light conditions (250 µE m-2 s-1) or extreme
high light conditions (4.5 mE m-2 s-1) for 24 hours. (S) standard light conditions; (H)
extreme high light conditions.
WT ctaDI-
S H S H
209
Figure 51. Effect of extreme high light intensity on growth rates of wild-type, ctaDI-,
ctaDII- and ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002. All cells were grown
under standard conditions until they reached the exponential phase of growth. The cells
were then diluted to OD550 = 0.05 and grown at either 250 µE m-2 s-1 or 4.5 mE m-2 s-1
continuous illumination. Open circles () represent cells grown under a continuous
illumination at 250 µE m-2 s-1. Closed circles () represent cells grown under a
continuous illumination at 4.5 mE m-2 s-1. Typical results of one of seven independent
growth experiments are shown. Panel (A) wild-type. Panel (B) ctaDI-. Panel (C)
ctaDII-. Panel (D) ctaDI- ctaDII-.
210
0.01
0.1
1
10
0 5 10 15 20 25
Time (h)
OD
550
Time (h)
0.01
0.1
1
10
0 5 10 15 20 25
A.O
D55
0
0.01
0.1
1
10
0 5 10 15 20 25
Time (h)
C.
0.01
0.1
1
10
0 5 10 15 20 25
Time (h)
B.
D.
211
3.5.6 Chlorophyll and carotenoid contents of wild-type, ctaDI-,
ctaDII- and ctaDI ctaDII- strains
Table 11 shows chlorophyll a and carotenoid contents of wild-type and cta
mutant strains grown under different light intensities. For cells grown under low to
standard constant illumination (150 to 250 µE m-2 s-1) there was virtually no difference in
the chlorophyll and carotenoid content between the mutant and wild-type strains. For
cells grown under moderate to very high light stress (400 to 4500 µE m-2 s-1), the
chlorophyll contents decrease in all strains compared to the chlorophyll concentrations
from cells grown under normal light intensities. The exception is for the ctaDI- mutant
strain, which fails to grow under extreme high light stress. These results suggest that in
all strains there is a change in the structure of the photosynthetic apparatus when cells are
grown under constant light stress, denoted by a decrease in the chlorophyll and
carotenoid contents. There is, however, no significant difference between the wild-type
and mutant strains in overall chlorophyll or carotenoid content under any of the light
intensities. The exception to this pattern is the ctaDI- strain. The ctaDI- strain is unable
to grow under conditions with constant extreme light stress (4.5 mE m-2 s-1) and thus
chlorophyll and carotenoid concentrations are impossible to determine under these
conditions. These observations are consistent with both oxygen evolution activity data
212
Table 11. Chlorophyll a and carotenoid contents for wild-type, ctaDI-, ctaDII-, and
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002 grown under various light
intensities. All values represent the averages of at five independent experiments ±
standard deviations. inviable: cells were unable to grow under these conditions.
Strain Light Intensity(µE m-2 s-1)
Chlorophyll a (µg ml-1 OD550 nm-1)
Carotenoids(µg ml-1 OD550 nm-1)
wild-type 150 3.7 ± 0.4 0.8 ± 0.2
ctaDI- 150 3.7 ± 0.3 0.8 ± 0.1
ctaDII- 150 3.6 ± 0.6 0.8 ± 0.1
ctaDI- ctaDII- 150 3.3 ± 0.3 0.8 ± 0.1
wild-type 250 3.4 ± 0.6 0.8 ± 0.1
ctaDI- 250 3.0 ± 0.5 0.8 ± 0.1
ctaDII- 250 3.4 ± 0.5 0.8 ± 0.2
ctaDI- ctaDII- 250 3.2 ± 0.7 0.9 ± 0.1
wild-type 400 3.4 ± 0.6 0.8 ± 0.1
ctaDI- 400 3.2 ± 0.6 0.8 ± 0.1
ctaDII- 400 3.4 ± 0.5 0.8 ± 0.2
ctaDI- ctaDII- 400 3.1 ± 0.5 0.8 ± 0.1
wild-type 700 1.5 ± 0.1 0.7 ± 0.3
ctaDI- 700 1.3 ± 0.1 0.7 ± 0.1
ctaDII- 700 1.5 ± 0.4 0.7 ± 0.3
ctaDI- ctaDII- 700 1.4 ± 0.6 0.7 ± 0.2
wild-type 1500 1.3 ± 0.1 0.7 ± 0.3
ctaDI- 1500 1.2 ± 0.1 0.7 ± 0.1
ctaDII- 1500 1.4 ± 0.4 0.7 ± 0.3
ctaDI- ctaDII- 1500 1.3 ± 0.2 0.7 ± 0.2
wild-type 4500 1.5 ± 0.1 0.7 ± 0.3
ctaDI- 4500 inviable inviable
ctaDII- 4500 0.6 ± 0.4 0.6 ± 0.2
ctaDI- ctaDII- 4500 0.7 ± 0.3 0.6 ± 0.2
213
that show a decrease in the oxygen evolving activity with an increase in light induced
stress in Synechococcus sp. PCC 7002 and the 77K fluorescence data (see below).
3.5.7 Photoinhibition of the whole chain electron transport activity
in Synechococcus sp. PCC 7002 wild-type, ctaDI-, ctaDII-, and
ctaDI- ctaDII- strains
The photoinhibition of the whole chain electron transport activity of wild-type
and the Cta-deficient strains of Synechococcus sp. PCC 7002 grown under standard
conditions was examined by incubation of the different strains under standard conditions
(250 µE m-2 s-1, 38°C, 10 mM NaHCO3), or under high light conditions (4.5 mE m-2 s-1,
38°C, 10 mM NaHCO3) for fixed time periods after which the oxygen evolving activities
of each of the samples were assayed. All strains were unaffected in their ability to evolve
oxygen when incubated under standard light intensity conditions for any amount of time
up to an hour (Figure 52). Differences were observed when the cells were incubated for
various times under very high light intensity conditions. For the first 40 min of
incubation under high light conditions, there was little effect on the oxygen evolving
capacity the wild-type strain of Synechococcus sp. PCC 7002 compared to the first 40
min of incubation of the wild-type strain under standard conditions. However, after 1
hour of high light treatment the wild-type cells had an oxygen evolution rate that was
214
Figure 52. Effects of high light intensity on whole chain electron transport in wild-
type, ctaDI- and ctaDII- strains of Synechococcus sp. PCC 7002. Cells were grown under
standard conditions until they reached the exponential phase of growth. Cells were
supplemented with 10 mM NaHCO3 and were treated at 38°C with either high light of 4.5
mE m-2 s-1 (wild-type ( ),ctaDI- (), ctaDII- (), ctaDI- ctaDII- (∇)), or standard light
intensity of 250 µE m-2 s-1 for all strains (). Photosynthetic oxygen evolution from
whole cells was measured at 38°C. The data shown are the averages ± standard deviation
(error bars) of five independent experiments.
0
20
40
60
80
100
% O
xyge
n ev
olut
ion
0 20 40 60
Time (minutes)
215
40% that of the wild-type cells incubated under standard conditions. After 1 hour of
high light incubation, the ctaDII- strain had nearly the same level of oxygen evolving
activity as the ctaDII- strain incubated under standard conditions. The ctaDI-/ctaDII-
strain also showed little variance between incubations under standard light conditions and
high light conditions for the first 40 min. The ctaDI- ctaDII- strain exhibited a 30%
decrease in oxygen evolving activity after incubation for 1 hour under high light
conditions. The ctaDI- strain displayed a roughly exponential decrease after exposure to
high light (Figure 52). These results suggest that the CtaDI-deficient strain is much
more sensitive to photoinhibition than the wild-type strain when the cells are exposed to
high light stress. However, the absence of the ctaDII gene product can somehow
compensate for the absence of the ctaDI gene product (see data for the ctaDI- ctaDII-
strain) and cause the cells to be less sensitive to photoinhibition.
3.5.8 Fluorescence emission at 77K of wild-type, ctaDI-, ctaDII- and
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Fluorescence emission spectra at 77K were collected for mutant and wild-type
strains in order to examine possible differences in PSI and PSII levels. Figure 53 shows
the 77K fluorescence emission spectra of whole cells of the wild-type strain, ctaDI-,
ctaDII-, and ctaDI- ctaDII- strains for cells grown at 100 µE m-2 s-1, 400 µE m-2 s-1, and
1500 µE m-2 s-1. Cells grown under moderately low light (100 µE m-2 s-1) exhibit little or
no difference in fluorescence emission maxima (Figure 53A). This implies that the
216
structure and ratio of both photosystems remains unchanged between the mutant
strains and the wild-type strain with regards to the reaction-center-associated
chlorophylls. When cells from both the wild-type and mutant backgrounds are grown
under moderate light stress (400 µE m-2 s-1), all strains show a shift in the ratio of PSI to
PSII, with a relative decrease in PSI and increase in PSII associated chlorophylls (Figure
53B). There is a decreased PSII content in the ctaDI- and ctaDII- single mutant strains
and a more pronounced reduction in fluorescence in the ctaDI ctaDII double mutant strain
when compared to the wild-type strain. This is easily visualized by examining the
decrease in fluorescence emission at 685 nm and 695 nm respectively of the mutant
strains compared to the wild-type strains (Figure 53B). Although both emission peaks at
685 nm and 695 nm are indicative of relative PSII levels, because of the light-activated
chlorophyll binding proteins fluorescence at 685 nm, the emission peak at 695 nm is
more indicative of the PSII content (Dr. Gaozhong Shen, personal communication).
When all the strains are grown under high light intensities (1500 µE m-2 s-1) there is a
relative decrease in PSII emission levels for all of the strains, but this is especially
evident in the mutant strains (Figure 53C). These results indicate that Synechococcus sp.
PCC 7002 changes the ratio of PSI to PSII during changes in growth light intensity.
These effects are more drastic in the mutant strains for which especially the PSI and PSII
emission levels are greatly decreased relative to the wild-type strain. Therefore, the
structural integrity of the photosystem complexes may be compromised at higher light
intensities and the lack of cytochrome oxidases further exacerbates this effect.
217
Figure 53. Fluorescence emission spectra at 77K of whole cells of Synechococcus
sp. PCC 7002 wild-type and cytochrome oxidase mutant strains. Samples were
normalized to an OD730nm = 1.0 for all strains and were excited at a wavelength of 446 nm.
Strains used are indicated in the figure. (A) Cells grown at 100 µE m-2 s-1. (B) Cells
grown at 400 µE m-2 s-1. (C) Cells grown at 1500 µE m-2 s-1.
218
30x103
25
20
15
10
5Flu
ores
cenc
e A
mpl
itude
800750700650600
Wavelength (nm)
Cells grown under 100 µE m-2 s-1
Wild type ctaDI-
ctaDII-
ctaDI- ctaDII-
80x103
60
40
20
0
Flu
ores
cenc
e A
mpl
itude
800750700650600
Wavelength (nm)
12x103
10
8
6
4
2Flu
ores
cenc
e A
mpl
itude
800750700650600
Wavelength (nm)
A
B
C
Wild type ctaDI-
ctaDII-
ctaDI- ctaDII-
Cells grown under 400 µE m-2 s-1
Cells grown under 1500 µE m-2 s-1
Wild type ctaDI-
ctaDII-
ctaDI- ctaDII-
219
3.5.9 P700+ Reduction Kinetics
To determine the effect that the cytochrome oxidases have on photosynthetic
electron transport, P700+ reduction kinetics were examined. Light-induced oxidation and
dark-induced reduction of P700 are readily monitored by a single-beam
spectrophotometer using an 830 nm measuring beam because of the complete absence of
additional components undergoing absorption changes in the same spectral region (Yu et
al., 1993). In order to obtain full reduction of P700+ in this experiment, 15-second dark
incubation times were used. This was followed by a 22-second light exposure to induce
the complete oxidation of P700. These light-dark cycles were repeated a total of 128
times per sample and the results were averaged using a Nicolet 4095 digital oscilloscope
as described in the Materials and Methods. These data were transferred to a Macintosh
G3 computer, with which IGORPRO v 3.5 was used to determine a best-curve fit, from
which the half-times for the reduction of P700+ were determined. The experiments were
repeated a minimum of three times and differences between the half-times determined
were less than 10%. The half-times of P700+ for the wild-type and mutant strains are
tabulated in Table 12.
220
Table 12. Half-times (ms) for P700 reduction in Synechococcus sp. PCC 7002.
Values represent averages ± standard deviations for three independent experiments.
Strain no inhibitors 10 mM DCMU 10 mM DCMU,1 mM KCN
wild-type 120 ± 8 330 ± 10 237 ± 10
ctaDI- 70 ± 5 130 ± 10 146 ± 10
ctaDII- 87 ± 7 262 ± 10 203 ± 10
ctaDI- ctaDII- 53 ± 4 127 ± 10 160 ± 10
DCMU: 3-(3,4-dichlorophenyl)-1,1-dimethylurea, KCN: potassium cyanide
In the absence of any inhibitors in the ctaDI- strain the reduction of P700+ is 1.7-
fold faster than in the wild-type strain. In the ctaDII- strain, P700+ reduction is 1.4-fold
faster than the wild-type strain. For the ctaDI- ctaDII- strain, P700+ reduction is 2.2-fold
faster than in the wild-type strain. These results indicate that in the absence of terminal
oxidases, there is an increase in the flow of electrons from the cytochrome b6f complex
for P700+ reduction.
The inhibitor 3-(3,4-dichlorophenyl)-1,1-dimethylurea (DCMU) inhibits electron
flow from PSII; therefore in the presence of DCMU the electron flow must be derived
predominantly from the NADH dehydrogenase and from cyclic electron flow around PSI
(Yu et al., 1993). In the case of the wild-type strain, the addition of DCMU to a final
concentration of 10 mM increased the half-time of P700+ reduction to 330 ms as opposed
to 120 ms in the absence of DCMU. The cta mutant strains all have faster half-times as
compared to the wild-type strain when DCMU was added (Table 12). In the presence of
221
10 mM DCMU, ctaDI- had a 2.5-fold faster half-time compared to the wild-type
strain under the same conditions. The ctaDII- strain had a 1.7-fold faster half-time
compared to the wild-type strain in the presence of 10 mM DCMU and the ctaDI- ctaDII-
strain has a 2.6-fold faster reduction of P700+ compared to the wild-type strain. These
results suggest that in the absence of Cta complexes, the majority of cyclic photosynthetic
electron transport goes through PSI, but when Cta complexes are present, some of the
electron flow is output through the terminal oxidases, resulting in a slower reduction of
P700+.
As a control, the cytochrome oxidase inhibitor KCN was added to all strains and
the P700+ reduction kinetics were measured. It had been noted previously that the
addition of KCN alone had no appreciable affect on the rate of P700+ reduction (Yu et al.,
1993). Therefore, these experiments were carried out in the presence of both KCN and
DCMU as described by (Yu et al., 1993). It was expected that in the presence of KCN,
the half-time for P700+ reduction would decrease for the wild-type and ctaDII- strains
relative to the same strains treated with DCMU alone. The half-time of P700+ reduction
was not expected to change significantly in the ctaDI- strains upon the addition of KCN.
This is what was observed (Table 12). The addition of KCN to a final concentration of 1
mM in addition to the 10 mM DCMU caused a 1.4-fold decrease in the half-time in the
wild-type strain compared to DCMU alone. The half-times for the ctaDI- and ctaDI-
ctaDII- mutant strains did not decrease, but the half-time for P700+ reduction in the
ctaDII- strain decreased 1.3-fold.
222
The faster half-times of P700+ reduction observed in the ctaDI- strains are
indicative of a bifurcation in electron flow from cytochrome c6 to PSI and cytochrome
oxidase. The faster half-times observed in the ctaDII- strain are likely due to the changes
in PSI stoichiometry discovered in the 77K fluorescence experiments (see section 3.5.8).
3.5.10 Pulse-Amplitude Modulated Fluorescence Measurements
Pulse-amplitude modulated (PAM) fluorescence measurements allow one to
measure the number of functional photosystem II complexes in whole cells. In order to
examine the electron flow around PSII, PAM fluorescence studies were performed at
room temperature. By examining the ratio of FV’/FM’ one can determine the total number
of active PSII sites compared to total chlorophyll content. Results indicate that there is a
decrease in this ratio for the strains harboring the ctaDI- mutation for cells grown at 250
µE m-2 s-1 (Table 10). Therefore, the ctaDI- and ctaDI- ctaDII- strains have a reduced
number of active PSII complexes, which correlates with the decreased fluorescence
emission at 685 and 695 nm at 77K (Figure 53).
223
3.5.11 Oxidative Stress and Cell Viability of Synechococcus sp.
PCC 7002 wild-type and ctaD mutant strains
Methyl viologen (MV) is a herbicide that can be reduced through a single-electron
transfer from reduced Fe-S clusters associated with PSI. This results in the formation of
a stable cation radical causing the subsequent reduction of oxygen, which forms the toxic
superoxide radical anion. In order to examine the effects of oxidative stress on
Synechococcus sp. PCC 7002, wild-type and cytochrome oxidase mutant strains were
grown in the presence and absence of MV and their growth rates were compared.
Tolerance to 50 µM of MV was observed for the ctaDII- and ctaDI- ctaDII- strains of
Synechococcus sp. PCC 7002 (Figure 54), which were able to grow in the presence of 50
µM MV unlike the wild-type and ctaDI- strains. The ctaDII- and ctaDI- ctaDII- strains
grown in the presence of MV had slightly lower chlorophyll contents but similar
carotenoid contents (ctaDII-, 2.9 ± 0.6 µg chl ml-1 OD550–1, 0.8 ± 0.3 µg car ml-1 OD550
–1, n
= 4 and ctaDI- ctaDII-, 3.0 ± 0.7 µg chl ml-1 OD550–1, 0.8 ± 0.4 µg car ml-1 OD550
–1, n = 4)
compared to cells grown without MV (see Table 11).
Figure 55 shows the growth curves of the Synechococcus sp. PCC 7002 wild-type
and cta mutant strains grown in the presence and absence of 50µM MV. These growth
curves indicate that the strains harboring the ctaDII- mutation are able to grow in the
presence of 50 µM MV at rates nearly equal to cultures grown without MV (Figure 55C
and 55D). This is in stark contrast to the growth rates of the wild-type strain or the
ctaDI- strain in the presence of 50 µM MV (Figure 55A and 55B). These
224
Figure 54. Effects of methyl viologen on cell death of Synechococcus sp. PCC 7002
wild-type and ctaDII- strains. Cells were grown at standard growth conditions until they
reached exponential growth phase. These cells were then diluted to an OD550nm = 0.05.
The cells were then grown in the presence (+) or absence (-) of 50 µM methyl viologen
for 24 hours.
225
Figure 55. Effects of MV on growth rates of wild-type, ctaDI- and ctaDII- strains of
Synechococcus sp. PCC 7002. Cells were grown under standard growth conditions until
they reached exponential growth phase. These cells were then diluted to an OD550 = 0.05
and grown in the presence or absence of 50 µM MV under otherwise identical conditions.
Open circles () represent cells grown without 50 µM MV, and closed circles ()
represent cells grown in the presence of 50 µM MV. Results represented are typical of
one of seven independent growth experiments. Panel (A): wild type. Panel (B): ctaDI-.
Panel (C): ctaDII-. Panel (D): ctaDI- ctaDII-.
226
Time (h)
0 5 10 15 20 25
OD
550
0.1
1
10A.
Time (h)
0.1
1
10
0 5 10 15 20 25
B.
0.1
1
10
0 5 10 15 20 25
Time (h)
C.
0.1
1
10
0 5 10 15 20 25
Time (h)
D.
OD
550
227
results indicate that strains harboring the ctaDII- mutation must have a mechanism to
counteract the increase in superoxide generated by methyl viologen.
The tolerance of the ctaDII- and ctaDI- ctaDII- strains of Synechococcus sp. PCC
7002 toward MV was further verified by cell viability counts (Table 13). Viability
counts reveal that there was a 45% decrease in the number of wild-type colony forming
units (CFU) per OD550 after a 4-hour incubation under standard conditions (250 µE m-2 s-
1, 38°C, 1.5% CO2/air) in the presence of 50 µM MV. Similarly, there was a 40%
decrease in the number of ctaDI- CFUs per OD550 after a 4 hour incubation under standard
growth conditions (250 µE m-2 s-1, 38°C, 1.5% CO2/air) in the presence of 50 µM MV.
However, there was no discernable difference in the number of CFUs for ctaDII- strains
incubated with or without 50 µM MV. The total number of CFUs decreases for all
strains when they were incubated in the dark compared to any of the strains incubated
under standard conditions without MV (Table 13). However, there were no discernable
differences in the number of CFUs for any of the strains incubated with or without 50 µM
MV in the dark when that particular strain was compared to itself. These results indicate
that MV most likely forms the cation radical as described by (Epel and Neumann, 1973)
during photosynthesis, but these results also indicate that MV is not very toxic when the
cells are incubated in the dark.
228
Table 13. Cell viability after incubation with or without 50 µM MV, in either light or
darkness. Conditions are described in Materials and Methods. Results are given as
number of colony forming units per OD550. Values presented are the average of three
independent experiments ± standard deviation.
Strains of Synechococcus sp. PCC 7002
treatment wild-type ctaDI- ctaDII- ctaDI- ctaDII-
none 8.0 ± 0.3 × 107 5.0 ± 0.3 × 107 7.4 ± 0.3 × 107 5.8 ± 0.5 × 107
50 µM MV 3.6 ± 0.1 × 107 2.0 ± 0.1 × 107 7.7 ± 0.4 × 107 6.0 ± 0.4 × 107
dark 4.8 ± 0.4 × 107 4.6 ± 0.3 × 107 6.6 ± 0.2 × 107 4.0 ± 0.3 × 107
dark + 50 µM MV 4.4 ± 0.4 × 107 4.6 ± 0.3 × 107 6.8 ± 0.3 × 107 3.7 ± 0.4 × 107
229
3.5.12 Superoxide Dismutase (SOD) Activity Assays
One possible explanation for the tolerance toward MV observed in the ctaDII- and
ctaDI- ctaDII- strains is that these strains have increased levels of enzymes (superoxide
dismutase (SOD), peroxidase, and catalase) related to the alleviation of oxidative stress.
In order to test this hypothesis, the wild-type and ctaDII- strains were assayed for
differences in various enzyme levels. Both soluble and membrane fractions derived from
the ctaDII- strain of Synechococcus sp. PCC 7002 have an increased level of SOD
activity compared to the fractions from the wild type strains (see Table 14). The SOD
activity of the membrane fractions for the ctaDII-strain was significantly higher than for
the wild-type (>2-fold difference). These results suggest that ctaDII gene product is
somehow involved in regulating the amount of membrane bound superoxide dismutase
activity and in the absence of ctaDII, there are greater amounts of functional, membrane-
bound superoxide dismutase. Thus, the higher SOD activity observed in the ctaDII- and
ctaDI- ctaDII- strains may be responsible for the tolerance these strains exhibit towards
MV.
230
Table 14. Superoxide dismutase activity in cell extracts of Synechococcus sp. PCC
7002. Values shown represent the averages of five separate experiments ± standard
deviation.
Strain WT ctaDI- ctaDII- ctaDI- ctaDII-
soluble proteins (mg) 5 ± 1 5 ± 1 6 ± 1 5 ± 1membrane-associatedproteins (mg)
5 ± 1 5 ± 1 6 ± 1 5 ± 1
total proteins (mg) 10 ± 2 10 ± 1 12 ± 2 10 ± 1soluble SOD units 173 ± 27 176 ± 20 220 ± 37 196 ± 16membrane-associatedSOD units
64 ± 12 67 ± 15 148 ± 30 134 ± 6
total SOD units 237 ± 33 243 ± 18 370 ± 63 330 ± 15% of total activity(soluble)
73% 72% 60% 59%
% of total activity(membrane)
27% 28% 40% 39%
fold-increase inactivity compared toWT (soluble)
1.0* 1.02 1.27 1.13
fold-increase inactivity compared toWT (membrane)
1.0* 1.04 2.31 1.84
fold-increase inactivity compared toWT (total)
1.0* 1.03 1.56 1.39
*normalized SOD activity, WT=1.0
231
3.5.13 Hydroperoxide levels in Synechococcus sp. PCC 7002
Another indicator of oxidative stress in cells is the concentration of
hydroperoxides in cells. Hydrogen peroxide is the by-product of SOD dismutation of
superoxide. In order to examine the effects of the mutations in the ctaDI and ctaDII loci,
the total level of hydroperoxides were measured in both wild-type and mutant strains
grown under standard conditions in the presence and absence of 50 µM MV. The results
are summarized in Table 15. The results indicate that the wild-type and ctaDI- strains
have similar levels of hydroperoxides both in the presence and absence of methyl
viologen. The ctaDII- strain has a 31% higher level of hydroperoxides compared to the
wild-type strain in the absence of methyl viologen and a 24% higher level of
hydroperoxides when treated with methyl viologen. The ctaDI- ctaDII- double mutant
had similarly increased levels of hydroperoxides as the ctaDII- strain. H2O2 is a direct by
product of superoxide dismutase activity (Fridovich and Hassan, 1979) and the presence
of higher concentrations of hydroperoxides in the ctaDII- mutant strains implies that there
is an increase in the level of activity of superoxide dismutase. However, this does not
correspond to an increase in the activity of catalases or peroxidases in the ctaDII- strains
(see below).
232
Table 15. Hydroperoxide levels in Synechococcus sp. PCC 7002 and cta mutant
strains.
Values shown represent the averages ± standard deviations of at least three separate
experiments.
Strain 50 µM methylviologen treatment
Hydroperoxides
(nmol ml-1 OD550 nm-1 )% peroxidesrelative towild-type
wild-type no additions 58 ± 3 100wild-type 50 µM MV
treatment54 ± 9 100
ctaDI- no additions 60 ± 9 103
ctaDI- 50 µM MVtreatment
54 ± 6 100
ctaDII- no additions 76 ± 9 131
ctaDII- 50 µM MVtreatment
67 ± 2 124
ctaDI- ctaDII- no additions 74 ± 1 128
ctaDI- ctaDII- 50 µM MVtreatment
70 ± 4 130
233
3.5.14 Catalase and peroxidase activity in Synechococcus sp. PCC
7002
To determine the whether or not the cta mutations had an effect on oxidative
stress enzymes other than SOD, catalase and peroxidase activities for the wild-type and
cta- strains were assayed. The results shown in Table 16 indicate that there is no
appreciable difference in catalase activity between the wild-type, ctaDI-, ctaDII-, or
ctaDI- ctaDII- strains. The results in Table 16 further reveal that there is no difference in
peroxidase activity between the wild-type and mutant strains. These results support the
previous observation from 3.5.13 where the levels of hydroperoxides were higher in the
ctaDII- and ctaDI- ctaDII- strains compared to the wild-type and ctaDI- strains.
Therefore, although interruption of ctaDII has an effect on the levels of SOD activity,
inactivation of the ctaDII gene has no effect on the levels of catalase or peroxidase
activity compared to the wild-type strain. These results imply that the interruption of
ctaDII only affects the SOD activity in these cells.
234
Table 16. Catalase and peroxidase activity in Synechococcus sp. PCC 7002 and cta
mutant strains. Values shown represent the averages ± standard deviations of at five
separate experiments. One unit results in the decomposition of one micromole of
hydrogen peroxide per minute at 25°C and pH 7.0 per OD550 nm.
Strain units of catalase ml-1
OD550 nm-1units of peroxidase ml-1
OD550 nm-1
wild-type 0.05 ± 0.01 1.7 ± 0.4ctaDI- 0.05 ± 0.01 1.7 ± 0.4ctaDII- 0.06 ± 0.02 1.7 ± 0.4ctaDI- ctaDII- 0.06 ± 0.02 1.7 ± 0.4
Chapter 4
DISCUSSION AND CONCLUSIONS
4.1 Genes and gene organization in cyanobacteria
The completely sequenced genome of Synechocystis sp. PCC 6803 revealed 16
genes that encode subunits for the type I NADH dehydrogenase complex (Kaneko et al.,
1996). Synechococcus sp. PCC 7002 has homologs to all of these subunits. There were
no cross-hybridizing fragments for the cydAB genes, which encode the quinol oxidase of
Synechocystis sp. PCC 6803, and these genes have not been detected in the
Synechococcus sp. PCC 7002 genome sequencing project. Synechococcus sp. PCC 7002
also lacks the hupLS genes, which encode the uptake hydrogenase in Anabaena sp. PCC
7120 (Carrasco et al., 1995).
This study reveals that most of the electron transfer genes have the highest
sequence similarity to their respective counterparts in the freshwater cyanobacterial strain
Synechocystis sp. PCC 6803 (Kaneko et al., 1996) or the filamentous, nitrogen fixing
cyanobacteria, Anabaena sp. PCC 7120 (Kazusa DNA Research Institute, 2000). The
comparisons with Anabaena sp. PCC 7120 are still incomplete since a final, complete
genomic sequence of that organism is still unavailable. However, the completely
236
sequenced genome of Synechocystis sp. PCC 6803 shows that there are many similarities
between the gene organization of electron transfer genes between Synechocystis sp. PCC
6803 and Synechococcus sp. PCC 7002. Examples of apparent operon organization
throughout cyanobacteria include the ndhAIGE cluster, the ndhF5 and ndhF2 genes, the
ndhCKJ cluster, and the ctaCIDIEI operon. However, there are some genes for which the
retention of operon structure appears to be more important in Synechococcus sp. PCC
7002 than in Synechocystis sp. PCC 6803. Examples of these apparent operons found in
Synechococcus sp. PCC 7002 but not found in Synechocystis sp. PCC 6803 or Anabaena
sp. PCC 7120 include the hydrogenase gene cluster and the ctaCIIDIIEII operon.
In Synechococcus sp. PCC 7002, the genes for both the structural components of
the hydrogenase enzyme, as well as the genes encoding hydrogenase assembly proteins,
occur in a single operon. However, in the other cyanobacterial species examined in this
study (Synechocystis sp. PCC 6803 and Anabaena sp. PCC 7120), although the genes for
the hydrogenase subunits are in relatively close proximity to one another, the genes for
the hydrogenase-assembly proteins are scattered throughout their respective genomes. In
the case of the ctaCIIDIIEII genes, Synechococcus sp. PCC 7002 appears to be unique in
that these genes form an operon; In contrast, although the ctaCII and ctaDII genes are in
close physical proximity to one in another in Synechocystis sp. PCC 6803, the ctaEII
gene is located elsewhere in the genome.
Why is there such diversity of gene organization in cyanobacteria? One possible
explanation is the varied transposon activity within different cyanobacterial species.
Many (106) transposon-like elements have been identified in the genome of
237
Synechocystis sp. PCC 6803 (Kaneko et al., 1996). This number is considerably higher
than the number of transposon-like elements identified in Synechococcus sp. PCC 7002
(5) thus far. Transposase activity may have been responsible for many of the gene
rearrangements in Synechocystis sp. PCC 6803. Conversely, the lack of these
transposon-like elements may explain the extensive retention of apparent operons in
Synechococcus sp. PCC 7002.
4.1.1 Future directions of the genome sequencing project in Synechococcus
sp. PCC 7002
Currently, the entire genome of Synechococcus sp. PCC 7002 is being sequenced
and assembled by the Bryant lab in collaboration with the lab of Dr. Jindong Zhao at
Beijing University and The Institute for Genetics, Beijing, China. The sequence data
gathered in this study are being used to orient and assemble contigs for the complete
sequence determination of Synechococcus sp. PCC 7002. The nucleotide sequences from
this study have also been used to fill in gaps from the sequencing project when they
overlap with other contigs.
238
4.2 Type I NADH Dehydrogenase
There are 14 subunits that constitute NDH-1 complex in E. coli, R. capsulatus, P.
denitrificans, and T. thermophilus (Yagi et al., 1998). These 14 subunits are homologous
to subunits of the mitochondrial type I NADH dehydrogenase of B. taurus and are
considered the minimal number of subunits necessary for the NADH-quinone
oxidoreductase activity. In contrast to these bacteria, plant chloroplasts and
cyanobacteria have homologs to only 11 of the 14 subunits found in other bacteria.
These subunits have been collectively termed the “alien complex” by (Friedrich et al.,
1995). An observation from this and another study (Friedrich et al., 1995) and is that the
chloroplast and cyanobacterial genes encoding subunits for the type I NADH
dehydrogenase are more closely related to one another than are mitochondrial and
chloroplast genes for type I NADH dehydrogenases, even if mitochondrial and plastic
NDH-1 gene sequences are derived from the same plant. The cyanobacterial type I
NADH dehydrogenase genes are also less similar to the bacterial genes encoding type I
NADH dehydrogenases than they are to chloroplast encoded type I NADH
dehydrogenase genes. These observations suggest a break in lineage or a separate lineage
in the origins of the NDH-1 genes from cyanobacteria and the chloroplast from those of
bacteria and the mitochondria. Noticeably absent in chloroplast and cyanobacteria are
the genes nuoF and nuoE. These genes encode the subunits harboring the FMN cofactor
and at least 4 Fe-S clusters. Furthermore, these subunits comprise the NADH
dehydrogenase (diaphorase) portion of the enzyme in purple bacteria (Friedrich et al.,
239
1995; Friedrich et al., 1990). This suggests that another donor/acceptor besides NADH is
utilized by the type I NADH dehydrogenase in chloroplasts and cyanobacteria. A
plausible electron donor/acceptor would be ferredoxin because of the predicted close
proximity of the type I NADH dehydrogenase to PSI (Berger et al., 1993).
The names of the genes and proteins for the NDH-1 subunits in cyanobacteria
have been assigned according to the nomenclature used for the chloroplast homologs and
a similar system of nomenclature has been used in this study. The Synechococcus sp.
PCC 7002 NDH-1 genes are found in several locations throughout its genome and their
organization is similar to the gene arrangements found in Synechocystis sp. PCC 6803.
The following NDH-1 genes were found in this study: ndhAIGE, ndhB, ndhD1, ndhD2,
ndhD3, ndhD4, ndhF1, ndhF2, ndhF3, ndhF4, ndhF5, ndhCKJ, ndhH, and ndhL. The
genes for two of the type I NADH dehydrogenase subunits (the ndhF1 gene (slr0844
homolog) (Schluchter et al., 1993) and the ndhD3 (slr1732 homolog) and ndhF3 (slr1733
homolog) genes (Klughammer et al., 1999) were characterized in previous studies.
Single-copy genes encoding putative subunits of the type I NADH dehydrogenase
in Synechococcus sp. PCC 7002 are represented by the ndhA, ndhI, ndhG, ndhE, ndhB,
ndhC, ndhK, ndhJ, ndhH, and ndhL genes. The phenotypes of mutations at several of
these loci have been investigated previously in Synechocystis sp. PCC 6803 (Ogawa,
1990; Ogawa, 1991). The ndhL open reading frame identified in Synechocystis sp. PCC
6803 is predicted to encode a hydrophobic protein of 80 amino acids. Deletion of the
ndhL gene in Synechocystis sp. PCC 6803 revealed that this strain is limited in its ability
to transport CO2 and HCO3- (Ogawa, 1990). Thus far, inactivation of the single copy ndh
240
genes in cyanobacteria consistently leads to defects in their abilities to uptake CO2 and
HCO3-. Therefore, similar phenotypes would be expected for such mutations in
Synechococcus sp. PCC 7002. Under the conditions presented in 3.1.1, the ndhB::cat
allele could not be segregated from the wild-type copy in Synechococcus sp. PCC 7002.
This may seem surprising given that these genes could be inactivated in Synechocystis sp.
PCC 6803 (Ogawa, 1991) and Synechococcus sp. PCC 7942 (Marco et al., 1993).
However, the results of (Ogawa, 1991) have recently come into question, since attempts
to repeat Ogawa's inactivation of the ndhB gene in Synechocystis sp. PCC 6803 by other
groups have also been unsuccessful (Dr. Wim Vermaas, Arizona State University,
personal communication). Thus, the initial observations made by Ogawa may have been
due to a secondary mutation that rescues the phenotype of an essential single-copy gene.
Mutational analysis of the ndhD3 and the ndhF3 genes in Synechococcus sp. PCC
7002 has revealed that these mutants, like the ndhB- mutants of Synechocystis sp. PCC
6803 (Ogawa, 1991) and Synechococcus sp. PCC 7942 (Marco et al., 1993), also require
high levels of CO2 (Klughammer et al., 1999). A similar phenotype was observed in
ndhD3- Synechocystis sp. PCC6803 strains (Price et al., 1998). However, mutations in
other ndhD genes are able to concentrate Ci at levels similar to the wild-type strain of
Synechocystis sp. PCC 6803 (Ohkawa et al., 2000). However, mutation analysis of the
ndhF1gene in Synechococcus sp. PCC 7002 revealed that the gene product of ndhF1 is
involved in cyclic electron transport and respiratory function (Schluchter et al., 1993) and
not in inorganic carbon transport (Sültemeyer et al., 1997b). Therefore, it has been
241
proposed that the different ndhD and ndhF gene products are involved with multiple,
functionally distinct, NDH-1-like complexes in cyanobacteria.
The phylogenetic analysis of the NdhD and NdhF protein sequences shows that
certain genes were probably misnamed upon their initial discovery (i.e. NdhF2 and
NdhF5). The names for the NdhD and NdhF proteins from Synechocystis sp. PCC 6803
cyanobacteria have been changed in this report to reflect the results of this phylogenetic
analysis. The close physical proximity and high degree of sequence similarity (>60%)
observed with the ndhF2 and ndhF5 genes throughout all of the cyanobacteria examined
in this study may be the result of a recent gene duplication at one of these loci. The
function of these genes and their gene products is still unknown.
4.2.1 Future directions for research of the type I NADH dehydrogenase
genes in Synechococcus sp. PCC 7002
To further investigate the function of the type I NADH dehydrogenase in
cyanobacteria, attempts should be made to inactivate the ndh genes discovered in this
study by interposon mutagenesis. Restriction maps have been provided in the
APPENDIX so designing these experiments would be relatively straightforward. Since a
plasmid with an interrupted ndhB gene has been constructed, and further attempts could
be made to isolate an NdhB-deficient strain under different growth conditions in order to
find out more about its relationship with the type I NADH dehydrogenase in
Synechococcus sp. PCC 7002.
242
4.3 Type II NADH dehydrogenases
There are two genes (ndbA and ndbB) that are predicted to encode type II NADH
dehydrogenases in Synechococcus sp. PCC 7002. However, the function of these gene
products has not been examined. Attempts to identify a homolog to ndbC of
Synechocystis sp. PCC 6803 by heterologous gene hybridization and examination of the
Synechococcus sp. PCC 7002 genome sequence data have thus far been unsuccessful.
The ndbA and ndbB genes have not yet been inactivated in Synechococcus sp. PCC 7002,
and thus their function is unknown.
Analyses of the predicted amino acid sequences of the NDH-2 proteins from
Synechococcus sp. PCC 7002 revealed that they all have putative NADH and FAD
binding motifs. These motifs consist of a β-sheet-α-helix-β-sheet structure, a glycine-
rich phosphate binding consensus sequence (GXGXG), a conserved negatively charged
residue (D or E) at the end of the second β-sheet, and six positions typically occupied by
small hydrophobic residues. Four of these hydrophobic residues are present in the
NADH binding motif, while the other two precede the conserved D or E residue. In
NADPH binding proteins, the third G in the GXGXXG motif is replaced by S, A, or P
and the negative charge at the end of the second β-sheet is missing (Howitt et al., 1999).
Since these features are absent in the predicted NDH-2 proteins from Synechococcus sp.
PCC 7002, it is likely that these enzymes bind NADH and not NADPH.
243
Both of the NDH-2 proteins predicted from the Synechococcus sp. PCC 7002
sequence also have the conserved nucleotide-binding domain for FAD and a secondary
motif downstream that is characteristic of FAD binding motifs (Howitt et al., 1999). This
motif contains the conserved Asp that forms a hydrogen bond with FAD. These
characteristics, along with the potential ability to bind NADH, are hallmarks of NDH-2
proteins.
4.3.1 Future directions for research of the type II NADH dehydrogenase
genes in Synechococcus sp. PCC 7002
(Howitt et al., 1999) proposed that the NDH-2 proteins act as redox sensors
within cyanobacteria. Now that the nucleotide sequence for the ndbA and ndbB genes is
available from Synechococcus sp. PCC 7002, it would be interesting to inactivate the
ndbA and ndbB genes in Synechococcus sp. PCC 7002 strains in which the PSI genes
have also been inactivated. If the high-light tolerance found in strains of Synechocystis
sp. PCC 6803 where both ndb genes and PSI genes have been inactivated (Howitt et al.,
1999) could be reproduced in Synechococcus sp. PCC 7002, it would be attractive to see
if the inactivation of these genes contributes to generate higher levels of proteins
designed to deal with oxidative stress (i.e. SOD, peroxidase, or catalase) similar to the
ctaII mutants. Additionally, it would be interesting to determine if the ndb- strains of
Synechococcus sp. PCC 7002 display the same tolerance to MV as the ctaDII- strains of
Synechococcus sp. PCC 7002. The Synechococcus sp. PCC 7002 NdbA and NdbB
244
proteins are predicted to be hydrophilic and it may be of interest to generate recombinant
versions of these proteins in E. coli for the generation of NdbA-specific and NdbB-
specific antibodies for use in physiological characterization of the Ndb proteins in
Synechococcus sp. PCC 7002. These recombinant Ndb proteins could also be used for
biochemical and biophysical characterization.
4.4 Bi-directional hydrogenase
An interesting finding in this study is that the genes for the bi-directional
hydrogenase are organized in a unique manner in Synechococcus sp. PCC 7002.
Compared to the organization of hydrogenase genes in other cyanobacterial species, all of
the hydrogenase genes are in close physical proximity to each other, possibly forming an
operon. There are many assembly proteins associated with the maturation of the
hydrogenase enzyme in bacteria (Jacobi et al., 1992). These assembly proteins may be
necessary for proper maturation of the hydrogenase enzyme. Synechococcus sp. PCC
7002 is also unique because the genes predicted to encode hydrogenase assembly proteins
(hyp genes) are clustered with the genes predicted to encode the hydrogenase itself. The
other cyanobacterial species examined in this study (Synechocystis sp. PCC 6803 and
Anabaena sp. PCC 7120) have the hyp genes located elsewhere in their respective
genomes.
Mutants of Synechococcus sp. PCC 7002 in which either the hoxH gene or both
the hoxH and hoxF genes have been inactivated show no apparent growth phenotypes
245
under normal growth conditions. These results are similar to results in Synechocystis sp.
PCC 6803, in which the hoxF gene was inactivated (Howitt and Vermaas, 1999). A
popular proposal was that the hoxF, hoxE, and hoxU subunits were the missing
cyanobacterial homologs of the type I NADH dehydrogenase genes nuoF, nuoE, and
nuoG (Appel and Schulz, 1996). However, this hypothesis seems unlikely, since the
mutant strains in both Synechocystis sp. PCC 6803 (Howitt and Vermaas, 1999) and
Synechococcus sp. PCC 7002 lack distinguishing phenotypes that could be attributed to
the absence of a functional type I NADH dehydrogenase. If the hoxF, hoxE, and hoxU
gene products were indeed functionally substituting for the nuoF, nuoE and nuoG gene
products, one would predict that the absence of a functional HoxF subunit would exhibit
a phenotype similar to that of strains in which a gene encoding a unique subunit of the
type I NADH dehydrogenase had been inactivated. However, the hoxF mutant strain has
no significant phenotypes when grown under standard growth conditions. It remains
possible that HoxF functions in conjunction with the type I NADH dehydrogenase under
alternative growth conditions (e. g. anaerobiosis).
4.4.1 Future directions for research of the bi-directional hydrogenase and
hyp genes in Synechococcus sp. PCC 7002
Although there has been no phenotype associated with the inactivation of the
HoxH and HoxF subunits of the hydrogenase in Synechococcus sp. PCC 7002, there may
be conditions that have not been assayed that will elucidate a function for the
246
hydrogenase in cyanobacteria. Hydrogenase expression in lithoautotrophic bacteria is
dependent on the level of hydrogen present. In R. eutropha, hydrogenases are produced
in the presence of H2 as well as during growth on poor carbon sources. Hydrogenase
synthesis is blocked during growth on preferentially utilized carbon sources, such as
succinate and pyruvate (Schwartz et al., 1998). In Bradyrhizobium japonicum, the
expression of hydrogenase is repressed by oxygen and carbon substrates and hydrogenase
expression is stimulated by hydrogen and carbon dioxide (Merberg et al., 1983). In
cyanobacteria, hydrogenase activity levels in heterocysts and vegetative cells of A.
variabilis and in the unicellular cyanobacteria, A. nidulans, were increased by incubation
of the cultures under anaerobic conditions and the addition of molecular H2 (Houchins
and Burris, 1981; Papen et al., 1986). Growth of Synechococcus sp. PCC 7002 under
anaerobic or microaerophilic conditions may also increase expression of the hydrogenase
enzyme and give some insight as to the physiological function of the bi-directional
hydrogenase in cyanobacteria. A simple way to conduct this experiment would be to
bubble cultures in liquid media with a 5% CO2 95% N2 gas mixture which would
effectively lower the O2 levels.
The Hyp proteins have been shown to be necessary for proper assembly and
function of the mature hydrogenase in other bacterial species (Jacobi et al., 1992;
Magalon and Bock, 2000; Olson et al., 1997; Olson and Maier, 1997). The hydrogenase
assembly factors have also been implicated in the proper assembly of other Ni-containing
enzymes in bacteria (Olson et al., 2001). These assembly factors may be necessary for
the assembly of other Ni containing enzymes in Synechococcus sp. PCC 7002. The hypB
247
gene may be a good target for inactivation to test the hypothesis of whether or not the
corresponding polypeptide is necessary for the assembly of Ni-containing enzymes in
Synechococcus sp. PCC 7002. It is known that Synechococcus sp. PCC 7002 has a urease
which utilizes Ni for its catalytic activity (Sakamoto et al., 1998). Inactivation of the
hypB gene could lead to the inability of Synechococcus sp. PCC 7002 to grow on urea.
Although the structures for other members of the hydrogenase family have been
solved, no structural data exist for the cyanobacterial bi-directional hydrogenases. The
subunits of bi-directional hydrogenase from Synechococcus sp. PCC 7002 are predicted
to be hydrophilic and it may be beneficial to try and produce recombinant forms of the
proteins for structural studies. Production of recombinant forms for the hydrogenase
subunits could also be used for antibody production. Hox-specific antibodies would be
useful to determine the location of the hydrogenase in the cell as well as for physiological
studies.
4.5 Mobile electron carriers
It is clear from this study that Synechococcus sp. PCC 7002 has some unique
genes compared to other cyanobacteria. Synechococcus sp. PCC 7002 has the genetic
potential to encode two soluble c-type cytochromes, petJ1 and petJ2. The petJ2 gene is
predicted to encode a second cytochrome c6 protein that would be different in primary
amino acid sequence from the cytochrome cm proteins found in Synechocystis sp. PCC
6803 and Synechococcus sp. PCC 7002. At this point, there is evidence that petJ1 is the
248
essential cytochrome c6 and there is no evidence that the petJ2 gene product can
functionally replace the petJ1 gene product. Synechococcus sp. PCC 7002 also has the
potential to synthesize a blue-copper protein encoded by the bcpA gene. The bcpA gene
is also found in Anabaena sp. PCC 7120, but it is not present in the genome of
Synechocystis sp. PCC 6803.
The failure to segregate the petJ1 gene in Synechococcus sp. strain PCC 7002 is
quite surprising, since inactivation of the petJ gene was achieved in both Synechocystis
sp. PCC 6803 and Synechococcus sp. PCC 7942 (Laudenbach et al., 1990; Zhang et al.,
1994). However, both Synechocystis sp. PCC 6803 and Synechococcus sp. PCC 7942
have a petE gene, encoding a plastocyanin. No evidence for the presence of the petE
gene in Synechococcus sp. strain PCC 7002 was obtained by Southern hybridization
using heterologous probes, and thus far, examination of the genome sequence of
Synechococcus sp. PCC 7002 has failed to reveal the presence of a petE gene.
Plastocyanin has never been isolated from nor detected in Synechococcus sp. strain PCC
7002. Caution must be exercised when making an argument for the absence of
plastocyanin, since heterologous hybridization analysis for the petE gene in
Synechococcus sp. strain PCC 7942 had also failed to reveal the presence of the gene
sequence or the gene product even though it was present (Clarke and Campbell, 1996).
However, the failure to obtain segregation of petJ1/petJ1::aphII under various growth
conditions points towards the absence of either plastocyanin or another mobile electron
carrier that could substitute for PetJ1, in Synechococcus sp. strain PCC 7002. The
absence of alternative electron carriers that could substitute for the PetJ1 gene product is
249
supported by an observation from (Manna and Vermaas, 1997). They observed that in
Synechocystis sp. strain PCC 6803, if the petE gene is deleted, the petJ gene cannot be
deleted and vice-versa. It is presumed, therefore, that these two mobile electron carriers
can substitute for one another, but a functional copy of one of these genes must always be
present in cyanobacteria for viability. Therefore, if there were only one gene for a
functional mobile electron carrier (petJ1) in Synechococcus sp. PCC 7002, attempts to
inactivate the petJ1 gene in Synechococcus sp. PCC 7002 would be similar to attempts to
inactivate both the petE gene and petJ gene in other cyanobacteria. This observation
supports the idea that Synechococcus sp. PCC 7002 has only one mobile electron carrier,
namely cytochrome c6, and that there is no plastocyanin in this species.
An alternative explanation for the failure to obtain a petJ1- strain would be that, if
plastocyanin or another mobile electron carrier is present in Synechococcus sp. strain
PCC 7002, it is unable to functionally substitute for cytochrome c6 under the conditions
tested. It is clear that Synechococcus sp. PCC 7002 has a copy of the cytM gene,
encoding cytochrome cM, at least one other gene predicted to encode a c- type
cytochrome, petJ2, as well as the gene encoding the potential blue-copper protein BcpA.
It is also clear that none of these genes can functionally substitute for the petJ1 gene
product. The attempted substitution of the Synechococcus sp. PCC 7002 petJ1 gene with
either the Synechocystis sp. PCC 6803 petE or petJ gene was also unsuccessful. These
heterologous genes were expressed in Synechococcus sp. PCC 7002 and a processed
Synechocystis sp. PCC 6803 petE gene product was detected by immunoblot analysis.
Since neither of these gene products could substitute for the Synechococcus sp. PCC 7002
250
PetJ1, it must be concluded that the interaction of PetJ1 with its redox partners is very
specific. Whether it is the reductive (cytochrome b6f) or oxidative (PSI or cytochrome
oxidase) side, or both that require such apparent specificity is unknown at this point. Of
special interest on the oxidative side of this electron transfer reaction is the psaF gene
product. The psaF gene product has a function as a docking protein for cytochrome c6 or
plastocyanin, allowing electron transfer between cytochrome c6 and P700 of PSI. In a
mutant strain of the cyanobacterium Synechococcus elongatus, Hippler et al. (1999)
generated a strain that expressed a hybrid Chlamydomonas reinhardtii/S. elongatus
version of the PSI subunit, PsaF. This hybrid PsaF was incorporated in vivo into the
cyanobacterial PSI and showed an increase by 2-3 orders of magnitude in the ability for
the C. reinhardtii mobile electron carriers to reduce P700+ (Hippler et al., 1999). Thus,
although the PsaF proteins from Synechococcus sp. PCC 7002 and Synechocystis sp. PCC
6803 are very similar (60% identical, 73% similar), the interactions of the PsaF proteins
with their mobile electron transport proteins are very specific, enough so that the
Synechocystis sp. PCC 6803 mobile electron carriers cannot functionally substitute for
PetJ1 in Synechococcus sp. PCC 7002.
Specificity of interaction of electron donors and acceptors is a reasonable
hypothesis as to why neither of the putative secondary electron carrier gene products,
BcpA or PetJ2, or for that matter the heterologous gene products from Synechocystis sp.
PCC 6803 cannot functionally substitute for PetJ1 in Synechococcus sp. PCC 7002. A
similar situation has been observed in the purple non-sulfur bacteria R. sphaeroides and
R. capsulatus. In R. capsulatus, there are two electron carriers between the cytochrome
251
bc1 complex and the reaction center, cytochrome c2 and cytochrome c y that can
functionally substitute for one another (Jenney and Daldal, 1993; Jenney et al., 1996). If
cytochrome c2 is deleted in R. capsulatus, this bacterium can still grow photosynthetically
using cytochrome cy (Jenney et al., 1996). The case for R. sphaeroides is different even
though the two organisms are closely related. Deletion of the cytochrome c2 gene results
in a non-photosynthetic mutant in R. sphaeroides (Hochkoeppler et al., 1995). This is in
spite of the fact that R. sphaeroides has a cytochrome cy gene. However, if the R.
capsulatus cytochrome cy gene is expressed in the R. sphaeroides cytochrome c2-deficient
strain, the cells again become photosynthetic (Jenney et al., 1996). Perhaps
Synechococcus sp. strain PCC 7002 does have a plastocyanin as R. sphaeroides has a
copy of the gene encoding cytochrome cy but this gene product is unable to substitute for
the main mobile electron carrier, which in Synechococcus sp. strain PCC 7002 is
cytochrome c6. This may point to a functional difference based upon specific structural
interactions between cytochrome c6 and any of its putative redox partners within the
thylakoid membrane (cytochrome b6f, PSI, or cytochrome oxidase). Cytochrome c6 in
Synechococcus sp. strain PCC 7002 may also be serving an alternative and unique
function that is different than that described in other cyanobacterial species.
252
4.5.1 Future directions for research of the mobile electron carrier genes in
Synechococcus sp. PCC 7002
Thus far, here have not been any phenotypes associated with the inactivation of
either the petJ2 or bcpA genes in Synechococcus sp. PCC 7002. Further physiological
characterization of the petJ2- and bcpA- strains under different growth conditions may
uncover a phenotype for these deficiencies. These genes may be involved in anaerobic or
microaerophilic metabolism, and therefore, no phenotype would have been observed
under the growth conditions tested. In Synechococcus sp. PCC 7002, the unique genes
present possibilities for new electron transfer pathways. An sqr gene encoding sulfide-
quinone oxidoreductase has been found on contig 450 from the Synechococcus sp. PCC
7002 genome sequencing project and presents the possibility that Synechococcus sp. PCC
7002 may be able to use hydrogen sulfide as an electron source. Sulfide-quinone
oxidoreductases have also recently been found in the filamentous cyanobacteria
Oscillatoria limnetica and Anabaena sp. PCC 7120 (Bronstein et al., 2000). Thus, it is
possible that the mobile electron-transport gene products may be involved with electron
transfer to and from the sulfide-quinone oxidoreductase.
Attempts to refold the rBcpA protein produced in E. coli have been unsuccessful.
The initial design of this plasmid was made with the belief that the mature protein would
be processed at a signal-sequence site in the amino terminus based on the ideas of ((von
Heijne, 1986). Since these truncated plasmids have not worked, perhaps a new
rBcpA/pET plasmid should be made with the full-length, unprocessed predicted protein,
253
if any further attempts will be made to overproduce this recombinant protein for
biochemical characterization.
4.6 Terminal Oxidases Present in Synechococcus sp. PCC 7002
It has been demonstrated previously that there are three gene clusters that encode
ORFs similar to known quinol and cytochrome oxidases in Synechocystis sp. PCC 6803
(Alge and Peschek, 1993a; Alge and Peschek, 1993b; Howitt and Vermaas, 1998). This
study reports the identification of two cta gene clusters similar to the two heme-copper
oxidase operons found in the related cyanobacterium, Synechocystis sp. PCC 6803
(Howitt and Vermaas, 1998). The CtaD protein is the largest subunit of the bacterial
cytochrome oxidase. Synechococcus sp. PCC 7002 has two genes that potentially
encode CtaD subunits, and these genes have been named ctaDI and ctaDII. The ctaDI
gene is predicted to encode a 549 amino acid polypeptide that would have a molecular
mass of 60.9 kDa and an estimated pI of 5.62. This polypeptide also contains the
putative Mg2+
binding site. The ctaDII gene is predicted to encode a 551 amino acid
protein with a molecular mass of 61 kDa and an estimated pI of 6.37. Alignments in the
region of the Mg2+
binding site show a high degree of identity with their counterparts
from Synechocystis sp. PCC 6803 (Figure 56). As in Synechocystis sp. PCC 6803, the
CtaDII subunit of Synechococcus sp. PCC 7002 is divergent in the region of the Mg2+
binding site.
254
The CtaC subunit of the bacterial cytochrome oxidase contains the putative CuA
binding motif. Synechococcus sp. PCC 7002 has two genes that potentially encode CtaC
subunits, and these genes have been named ctaCI and ctaCII. The ctaCI gene is
predicted to encode a 362 amino acid polypeptide that would have a molecular mass of
39.6 kDa and an estimated pI of 4.55. The ctaCII gene is predicted to encode a 298
amino acid protein with a molecular mass of 33.8 kDa and an estimated pI of 6.50.
Figure 56 also shows an alignment of the CuA binding motif for the Synechococcus sp.
PCC 7002 and Synechocystis sp. PCC 6803 CtaCI proteins. The CuA binding motifs of
the CtaCII proteins from both Synechococcus sp. PCC 7002 and Synechocystis sp. PCC
6803 are similarly divergent in this binding motif.
255
Figure 56. Oxidase Mg2+ and CuA binding motif alignments. A. Alignments of the
heme-copper oxidase Mg2+ binding region in the cyanobacterial CtaD subunit with the
large subunits of other cytochrome oxidases. Putative binding ligands are in bold. In the
cyanobacterial CtaDII subunits, the conserved histidine and aspartic acid have been
replaced by asparagines. B. Alignments of the heme copper oxidase CuA binding region
in the cyanobacterial CtaC subunit with subunit II from other cytochrome oxidases.
Putative binding ligands are in bold. In the cyanobacterial CtaCII subunits, the conserved
cysteine is replaced by aspartic acid. The conserved glutamic acid is replaced by
glutamine. The conserved histidine is replaced by phenylalanine. A single methionine is
conserved in all CtaC subunits of heme copper oxidases.
CtaDI 7002 368 APFDIHVHDTYFVVGHFHYVCtaDI 6803 371 VPFDIHVHDTYFVVGHFHYVCtaDII 7002 382 VPVDIHVNNTYFVVGHFHYVCtaDII 6803 361 APFDLHVNNTYFVVGHFHYVBos taurus 361 SSLDIVLHDTYYVVAHFHYVS. cerevisiae 361 ASLDVAFHDTYYVVGHFHYVP. denitrificans 396 GSLDRVYHDTYYIVAHFHYVB. subtilis 368 AAADYQFHDTYFVVAHFHYV
A. Alignment of the Mg2+ binding motif from CtaD
CtaCI 7002 274 PVICAELCGSYHGGMKTTMTVETAEGYDQW CtaCI 6803 245 PVICAELCGAYHGGMKSVFYAHTPEEYDDWCtaCII 7002 211 RIRDSQFSGTYFAAMQADVVVESQEDYQTWCtaCII 6803 230 KLHDSQFSGTYFAVMTAPVVVQSLSDYQAWBos taurus 193 YGQCSEICGSNHSFMPIVLELVPLKYFEKW S. cerevisiae 221 YGACSELCGTGHANMPIKIEAVSLPKFLEW P. denitrificans 213 FGQCSELCGINHAYMPIVVKAVSQEKYEAW B. subtilis 214 FGKCAELCGPSHALMDFKVKTMSAKEFQGW
B. Alignment of the CuA binding motif from CtaC
256
From these observations, it may be concluded that, as in Synechocystis sp. PCC
6803, the CtaDII and CtaCII subunits are similar to the bo-type quinol oxidase subunits,
whereas the counterparts CtaDI and CtaCI are similar to the subunits of the aa3 type
cytochrome oxidase (Howitt and Vermaas, 1998).
Both the ctaDI and ctaDII genes of Synechococcus sp. PCC 7002 are dispensable
for growth under normal to moderately high light conditions. As in Synechocystis sp.
PCC 6803, the primary cytochrome oxidase is encoded by the ctaCIDIEI gene cluster in
Synechococcus sp. PCC 7002, and the secondary cytochrome oxidase, encoded by the
ctaCIIDIIEII gene cluster, seems to contribute little to respiratory processes under the
conditions tested. However, unlike Synechocystis sp. PCC 6803, which has a functional
quinol oxidase encoded by the cydAB operon, no cydAB-like genes were detected in
Synechococcus sp. PCC 7002 by heterologous Southern blot hybridization using
Synechocystis sp. PCC 6803 cydA or cydB specific probes. The absence of the cydAB
genes in Synechococcus sp. PCC 7002 is not surprising based upon the oxygen-uptake
data in this study. In Synechocystis sp. PCC 6803, mutagenesis of either cydAB or
ctaDIEI singly had no effect on respiratory rates when compared to Synechocystis sp.
PCC 6803 wild type respiratory rates, while a double mutation in these loci had a
dramatic effect on oxygen consumption (Howitt and Vermaas, 1998). In marked
contrast, the single mutant, ctaDI- in Synechococcus sp. PCC 7002 drastically reduces the
rates of oxygen consumption when compared to the Synechococcus sp. PCC 7002 wild-
257
type strain. This is further functional evidence that Synechococcus sp. PCC 7002 lacks a
cydAB-type quinol oxidase.
4.6.1 Effects of Cytochrome Oxidase on Electron Flow Around PSII
It is clear from the change in ratios of FV’/FM’ between the wild-type and the
ctaDI- and ctaDI- ctaDII- mutant strains of Synechococcus sp. PCC 7002 that electron
flow around PSII has been modified. However, this ratio is virtually unchanged in the
strain with the ctaDII- single mutation. FV’/FM’ reflects the photochemical efficiency of
open PSII centers in higher plants under a given light acclimation status and represents
the combined regulation of PSII via both reversible non-photochemical quenching and
photoinhibitory inactivation of PSII (Campbell et al., 1998). However, in cyanobacteria,
changes in FV’/FM’ also combine non-photochemical influences on PSII function as well
as photoinhibitory inactivation of PSII (Campbell et al., 1998). It has been documented
that cyanobacteria have a high capacity to remove electrons from PSII with oxygen as a
terminal acceptor for electron flow from water (Campbell et al., 1998). This electron
flow is low under low light conditions, variable but significant at the standard growth
light intensity, and large under conditions of excess light (Campbell et al., 1998). This
activity is sensitive to cyanide in other cyanobacteria, which also suggests that there is a
contribution from the activities from cytochrome oxidase (Schubert et al., 1995). Thus,
with the evidence presented here, it is logical to conclude that the primary oxidase,
CtaCIDIEI serves as a buffer from excess excitation of PSII by removing electrons as
258
necessary in Synechococcus sp. PCC 7002. These conclusions are in complete agreement
with our physiological results with the ctaDI- mutants and are reflected in the ratio of
FV’/FM’.
4.6.2 Cytochrome Oxidase and its Role in High Light Stress
Under low to moderate light stress conditions, the mutations in the cta loci cause
little or no physiological duress to Synechococcus sp. PCC 7002. However, the P700+
reduction kinetics of the mutants were strongly affected, with the half-times of P700+
reduction decreasing dramatically in the ctaDI- strain. This suggests that cytochrome
oxidase plays a role within cyanobacterial cells to process the electrons generated by PSII
and competes with PSI for electrons. There are no significant differences in the total
amount of chlorophyll from the cells of the Cta-deficient mutants when compared to the
chlorophyll levels in wild type cells under many different light conditions. An exception
to this pattern was observed in the ctaDI- single mutant strain, which failed to grow under
extremely high-light stress (4.5 mE m-2 s-1, 38°C, 1.5% CO2/air). The absence of the
primary cytochrome oxidase from these cells places an increased amount of redox stress
on the other proteins in the electron transport chain. The resulting increased amount of
reductant may have a deleterious effect and cause an overall reduction in the number of
functional PSII complexes in the ctaDI- strain at higher light intensities because of
oxidative damage. The inability of the ctaDI- cells to manage an increased amount of
light stress becomes more evident when the cells are exposed to increasing light
259
intensities. As the light intensities increased, the cells had a gradual loss of PSII as
reflected by a decrease in the 77K fluorescence emission peaks at 685 nm and 695 nm.
These observations lead to the conclusion that the CtaI complex acts as an electron sink,
removing excess electrons to insure the fidelity of the other proteins in the electron
transport chain.
4.6.3 Tolerance of Methyl Viologen and High Light Stress in ctaDII- and
ctaDI- ctaDII- strains of Synechococcus sp. PCC 7002
Unlike the ctaDI- strain of Synechococcus sp. PCC 7002, the ctaDII- strain is able
to tolerate growth at 4.5 mE m-2 s-1. This suggests that the CtaII complex does not play
the same role as the CtaI complex in Synechococcus sp. PCC 7002. Interestingly, the
tolerance to extreme light intensities is observed even in the ctaDI- ctaDII- double mutant
strain. This observation is unusual because ctaDI- strain is sensitive to high light.
Therefore, the ctaDII- strain suppresses one defect associated with the ctaDI mutant. The
ctaDI- ctaDII- strain, similar to the ctaDI mutant strain, is severely affected in its ability
to uptake oxygen. Therefore, it is not simply an increased level of respiratory activity
that accounts for the increased tolerance to light stress observed in the ctaDII- mutant
strains, but rather, some unknown mechanism.
An additional observation was that the ctaDII- strains showed a high tolerance
towards oxidative stress based on the tolerance these cells exhibit towards the addition of
methyl viologen to the growth medium. Methyl viologen (Paraquat) is a herbicide that
260
causes oxidative stress by increasing the amount of superoxide (O2-) in cells. Mutations
in the ctaDII locus are resistant to high levels (50 µM) of MV in the growth medium,
whereas wild-type and ctaDI- cells are both sensitive to the addition of MV to the growth
medium.
Superoxide dismutase (SOD) is an enzyme that catalyzes the destruction of the
superoxide (O2-) radical. SOD has been widely implicated in protecting cells against the
harmful effects of active oxygen (Asada, 1999; Storz and Imlay, 1999). There are three
types of SOD enzymes that can be distinguished by their metal cofactors at the active
site: iron (Fe-SOD), manganese (Mn-SOD), and copper/zinc (Cu/Zn-SOD) (Asada,
1999). Two of the enzymes have been found in cyanobacteria: the iron (Fe-SOD)
enzyme and the manganese (Mn-SOD) enzyme (Herbert et al., 1992; Okada et al., 1979).
The SOD activity can be divided into a soluble activity that most likely represents the
constitutively expressed Fe-SOD enzyme activity and a membrane fraction containing the
Mn-SOD enzyme (Okada et al., 1979). The genes for both the Fe-SOD (contig 344) and
Mn-SOD (contig 384) have been identified in the sequencing project for the
Synechococcus sp. PCC 7002 genome. The SOD activity assays indicate that there are
significantly higher levels of superoxide dismutase activity in both of the ctaDII- strains
of Synechococcus sp. PCC 7002. Higher SOD activity would allow cells to tolerate the
excess generation of superoxide anion from the MV added to the media. Thus, higher
levels of SOD activity partially explain the tolerance of the ctaDII- strains exhibit in the
presence of MV.
261
Examination of the activity of oxidative stress enzymes (SOD, catalase, and
peroxidase) as well as the accumulation of total hydroperoxides has indicated that only
SOD activity is significantly increased in the ctaDII- strains. Therefore, it can be implied
that although the other enzymes linked to oxidative stress may be important to the cells,
they are nonetheless not the major determinants for the tolerance to prolonged exposure
to high-light stress or to a sudden oxidative stress, specifically the generation of
superoxide by MV.
The nucleotide sequence surrounding the ctaDII gene cluster reveals two open
reading frames immediately upstream of the ctaCII gene that are homologs of the sll1485
and sll1485 open reading frames in Synechocystis sp. PCC 6803. The function of these
open reading frames is unknown at this time. Immediately downstream of the ctaEII
gene, there is a partially sequenced open reading frame with similarity to the
Gluconobacter oxydans sldI gene. The sldI gene encodes the large subunit of the sorbitol
dehydrogenase. The ctaDII gene was interrupted with the Ω fragment (Prentki and
Krisch, 1984), which has two transcriptional terminators flanking the aadA gene
sequence. Thus, substantial increase in the transcription of the open reading frames
immediately upstream or downstream of the ctaDII loci is unlikely.
The unusual phenotype of oxidative stress tolerance observed in the ctaDII-
strains may be due to an interruption in a signaling pathway in which the secondary
cytochrome oxidase is involved. If the CtaDII enzyme is somehow involved in the
down-regulation of superoxide dismutase, the absence of CtaDII in the ctaDII- strains
could result in cells that contain higher basal levels of the SOD. These higher levels of
262
SOD could be responsible for the tolerance of oxidative stress. It has been shown here
that the ctaDII- strains have increased levels of membrane bound SOD activity, but that
the levels of other enzymes associated with oxidative stress (catalase and peroxidase) are
not affected by this mutation. The use of an oxidase as a signaling transducer is not
unprecedented in prokaryotes. An aerotaxis transducer has been found in archaebacteria
(Brooun et al., 1998). In addition, R. sphaeroides contains a cbb3-type cytochrome that is
responsible for the repressing expression of photosynthesis gene in the presence of
oxygen (O'gara and Kaplan, 1997) and is believed to play a role in aerotaxis signaling
(Armitage et al., 1985). It remains to be determined whether there are other enzymes
involved in regulating the levels of membrane bound SOD activity in Synechococcus sp.
PCC 7002.
4.6.4 Future research directions for cta- mutants in Synechococcus sp. PCC
7002
Further characterization of the heme-copper oxidase mutant strains may reveal
additional insight into the functions of these enzymes in vivo. If the CtaCIIDIIEII
enzyme is acting as a redox response regulator, it is important to identify its redox
partners and to determine how it regulates the expression of other genes within
cyanobacteria. Now that the genes encoding subunits from three large electron transport
enzymes have been cloned, interposon mutagenesis may be performed to help elucidate
the physiological function of these enzymes in cyanobacteria.
263
It has been recently found that the ratios of specific carotenoids are altered in
response to oxidative stress in plants (Carol and Kuntz, 2001) and R. sphaeroides (O'gara
and Kaplan, 1997; O'gara et al., 1998). In both plants and bacteria, these physiological
changes are mediated by specific oxidases. In R. sphaeroides, this event is controlled by
the cbb3 oxidase (O’gara and Kaplan, 1997; O’gara et al., 1998) and in Arabidopsis
thaliana carotenoid contents are controlled by the plastidal terminal oxidases (PTOX)
(Carol and Kuntz, 2001). Although the overall carotenoid content did not change in the
wild-type and cta mutant strains of Synechococcus sp. PCC 7002, the ratios of the
different species of carotenoids may have changed. In addition to the production of
oxidative stress related enzymes (SOD, peroxidase, catalase), changes in the ratios of
carotenoids are also important for adaptation to oxidative stress (Carol and Kuntz, 2001).
The future addition of a photodiode array detector to the Bryant lab will facilitate the
determination of differences in the carotenoid contents of the wild-type and cta mutant
strains by HPLC analysis.
Figure 57 shows an alternative explanation to the toxic effects observed when
ctaDI is inactivated. In the wild-type strain, the genes encoding both CtaCIDIEI and
CtaCIIDIIEII are transcribed and the proteins are presumably produced. When the wild-
type cells are grown under 4.5 mE m-2 s-1 constant illumination, no toxicity effect is
observed (Figure 57A). Similarly, no toxicity effects are observed in the ctaDII- and
ctaDI- ctaDII- strains (Figure 57C and Figure 57D). However, the absence of CtaDI
264
Figure 57. Model for toxicity in ctaD mutant strains grown under 4.5 mE m-2 s-1 constant
illumination. (A) The wild-type strain of Synechococcus sp. PCC 7002 is able to produce
complete ctaCIDIEI and ctaCIIDIIEII transcripts and products. It is possible that the
presence of the CtaCIDIEI complex counteracts any toxic effects from a CtaCIIDIIEII
complex. (B) Inactivation of the ctaDI gene still allows the possibility for the expression
of the ctaCI gene and production of a hybrid CtaCIDIIEII complex as well as the native
CtaCIIDIIEII complex. Either of these could be toxic to the cell. (C) Inactivation of the
ctaDII gene still allows the possibility for the expression of the ctaCII gene and
production of a hybrid CtaCIIDIEI complex as well as the native CtaCIDIEI complex.
Either the CtaCIIDIEI complex is not toxic to the cells or the CtaCIDIEI enzyme is able
to compensate for toxic effects generated by the hybrid CtaCIIDIEI enzyme. (D)
Inactivation of both the ctaDI and ctaDII genes eliminates the possibility of making
either complex, therefore no toxic effect is observed.
265
A. WT
B. ctaDI-
C. ctaDII-
D. ctaDI- ctaDII-
possible transcripts
possible products
toxic
CII DII EII
CI DI EI CI DI EI
CII DII EII
CI DI EI
CII DII EII
CI DI EI
CII DI EI
CII DII EII
CI DI EIno products?
CI DI EI
CII DII EII
CI DII EII
CII DII EII
266
singularly is toxic to cells grown under 4.5 mE m-2 s-1 constant illumination. Two
possible explanations of these phenomena are: (1) the inactivation of ctaDI eliminates the
production of the entire CtaCIDIEI enzyme which, when present, counteracts a toxic
effect generated by the CtaCIIDIIEII enzyme or (2) the inactivation of the ctaDI gene
eliminates only the CtaDI subunit, but the capacity to produce the CtaCI subunit still
exists (Figure 57B).
It is clear from this study that the levels of detectable transcript for ctaCIDIEI are
much higher than those of ctaCIIDIIEII. Thus, one could assume, if ctaCI were
expressed, that the CtaCI protein could be produced in higher concentrations than the
CtaCII subunit. Based on the similarity between the Cta subunits, one might imagine that
the CtaCI subunit could interact and form complexes with the CtaDII and CtaEII
subunits. If the CtaCI subunit were functional in these hybrid complexes, it would
presumably be able to accept electrons from cytochrome c6. This could possibly place a
higher level of reducing equivalents through what is normally a low activity enzyme
(CtaCIIDIIEII). Perhaps it is the higher levels of reducing equivalents through a hybrid
CtaCIDIIEII enzyme that causes the toxicity observed in the ctaDII- strain.
In the ctaDII- strain, it is impossible to produce either the CtaCIIDIIEII enzyme or
the CtaCIDIIEII hybrid enzyme, thus no toxic effects are observed (Figure 57C) and in
the ctaDI- ctaDII- strain it is impossible to produce either of the CtaCIIDIIEII or the
CtaCIDIIEII enzyme, therefore, no toxic effects are observed in this strain either (Figure
57D). An experiment where the ctaCI and ctaCII genes were inactivated could answer
whether the proposed hybrid CtaCIDIIEII enzyme was toxic to the cells. Inactivation of
267
the ctaCI gene would eliminate the possibility of a CtaCI protein interacting with the
CtaDII and CtaEII subunits, therefore preventing the formation of the hybrid CtaCIDIIEII
enzyme. If growth under 4.5 mE m-2 s-1 constant illumination no longer proved fatal to a
ctaCI- strain, it would lend credence to the idea that presence of a CtaCIDIIEII enzyme
causes toxicity within the cells. However, if growth under constant extreme light
intensity were still fatal to the ctaCI- strain, it would lend support to the idea that the
CtaCIIDIIEII enzyme or the product of its reaction is toxic to the cell. It would also
suggest that the CtaCIDIEI enzyme is able to counteract the toxicity caused by the
CtaCIIDIIEII enzyme.
This study has established protocols to perform experiments where cyanobacterial
cells could be exposed a constant level of either extremely high light stress (growth under
4.5 mE m-2 s-1) or oxidative stress in the presence of methyl viologen. It will be
interesting to determine the effects of these stresses on other electron transport mutants,
which may now be created based on genomic sequence information. Experiments such
as these will allow us to better characterize the stress responses of cyanobacterial cells.
4.7 Concluding Remarks
Information provided by the sequencing of genomes from different organisms
allows us to compare the types of genes, number of genes as well as the gene
organization of different species. The value of comparative analysis of both gene
sequences and organization is evident from this study. Comparing the sequences of
268
predicted gene products allows us to make predictions about the functions of the gene
products. Examination of gene organization may also provide a means to identify the
function of the genes, especially if these genes are organized in apparent operons. This
information provides an invaluable starting point for experimentation.
Although many (40) genes predicted to encode electron transport proteins were
identified in this study, it likely does not represent the total number of electron transport
proteins within the cell. There are still many open reading frames with unassigned
function that have been identified in Synechocystis sp. PCC 6803 and homologs to many
of these open reading frames have also been found in Synechococcus sp. PCC 7002.
Some of these unassigned open reading frames may encode electron transport proteins.
Further studies of electron transport proteins will allow the opportunity to discover new
functions for these proteins. In this study, phenotypes discovered from the inactivation of
the ctaD genes of Synechococcus sp. PCC 7002 have given us new insights to the
function of heme-copper oxidases in cyanobacteria. It will be exciting to find out what
other electron proteins may be acting in oxidative stress response pathways.
This study has shown that it is dangerous to hold a single organism as the model
for all others simply because genomic information is readily available. Two genes
characterized in this study (petJ1, ndhB) were found to be essential for viability in
Synechococcus sp. PCC 7002 whereas they were non-essential in other cyanobacteria. It
is important to recognize that these lab ‘workhorses’ were once isolated from different
environments, and the adaptation to these environments has allowed these organisms to
acquire some unique proteins. In studying the differences and similarities of the proteins
269
between organisms, we will better understand how nature uses similar scaffolding to
create proteins which fulfill different needs in the cell.
270
References
Abramson, J., Riistama, S., Larsson, G., Jasaitis, A., Svensson-Ek, M., Laakkonen, L.,
Puustinen, A., Iwata, S., and Wikstrom, M. (2000). The structure of the ubiquinol oxidase
from Escherichia coli and its ubiquinone binding site, Nat Struct Biol 7, 910-7.
Adams, M. W., Mortenson, L. E., and Chen, J. S. (1980). Hydrogenase, Biochimica et
Biophysica Acta 594, 105-176.
Alex, L. A., Reeve, J. N., Orme-Johnson, W. H., and Walsh, C. T. (1990). Cloning,
sequence determination, and expression of the genes encoding the subunits of the nickel-
containing 8-hydroxy-5-deazaflavin reducing hydrogenase from Methanobacterium
thermoautotrophicum ∆H, Biochemistry 29, 7237-7244.
Alge, D., and Peschek, G. A. (1993a). Characterization of a cta/CDE operon-like
genomic region encoding subunits I-III of the cytochrome c oxidase of the
cyanobacterium Synechocystis PCC 6803, Biochem Mol Biol Int 29, 511-25.
Alge, D., and Peschek, G. A. (1993b). Identification and characterization of the ctaC
(coxB) gene as part of an operon encoding subunits I, II, and III of the cytochrome c
oxidase (cytochrome aa3) in the cyanobacterium Synechocystis PCC 6803, Biochem
Biophys Res Commun 191, 9-17.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local
alignment search tool, Journal of Molecular Biology 215, 403-410.
Anderson, S. L., DeBruijin, M. H. L., Coulson, A. R., Eperon, I. C., Sanger, F., and
Young, I. G. (1982). Complete sequence of bovine mitochondrial DNA, Journal of
Molecular Biology 156, 683-717.
271
Appel, J., and Schulz, R. (1996). Sequence analysis of an operon of a NAD(P)-reducing
nickel hydrogenase from the cyanobacterium Synechocystis sp. PCC 6803 gives
additional evidence for direct coupling of the enzyme to NAD(P)H-dehydrogenase
(complex I), Biochim Biophys Acta 1298, 141-7.
Arizmendi, J. M., Runswick, M. J., Skehel, J. M., and Walker, J. E. (1992a).
NADH:ubiquinone oxidoreductase from bovine heart mitochondria. A fourth nuclear
encoded subunit with a homologue encoded in chloroplast genomes, FEBS Letters 301,
237-242.
Arizmendi, J. M., Skehel, J. M., Runswick, M. J., Fearnley, I. M., and Walker, J. E.
(1992b). Complementary DNA sequences of two 14.5 kDa subunits of
NADH:ubiquinone oxidoreductase from bovine heart mitochondria. Completion of the
primary structure of the complex?, FEBS Letters 313, 80-84.
Armitage, J. P., Ingham, C., and Evans, M. C. (1985). Role of proton motive force in
phototactic and aerotactic responses of Rhodopseudomonas sphaeroides, J Bacteriol 161,
967-72.
Asada, K. (1999). The Water-Water Cycle in Chloroplasts: Scavenging of Active
Oxygens and Dissipation of Excess Photons, Annual Review of Plant Physiology and
Plant Molecular Biology 50, 601-639.
Beers, R., and Sizer, I. (1952). A spectrophotmetric method for measuring the breakdown
of hydrogen peroxide by catalase, Journal of Biological Chemistry 195, 133.
Berg, J. M., and Holm, R. H. (1982). Sulfur protein clusters and their synthetic analogs
(New York, John Wiley and Sons).
272
Berger, S., Ellersiek, U., Westhoff, P., and Steinmüller, K. (1993). Studies on the
expression of NDH-H, a subunit of the NAD(P)H-plastoquinone-oxidoreductase of
higher-plant chloroplasts, Planta 190, 25-31.
Birnboim, H. C., and Doly, J. (1979). A rapid alkaline extraction procedure for screening
recombinant plasmid DNA, Nucleic Acids Research 7, 1513-1523.
Blattner, F. R., Plunkett III, G., Bloch, C. A., Perna, N. T., Burland, V., Riley, M.,
Collado-Vides, J., Glasner, J. D., Rode, C. K., Mayehew, G. F., et al. (1997). The
complete genome sequence of Escherichia coli K-12, Science 277, 1453-1461.
Boison, G., Schmitz, O., Mikheeva, L., Shestakov, S., and Bothe, H. (1996). Cloning,
molecular analysis and insertional mutagenesis of the bidirectional hydrogenase genes
from the cyanobacterium Anacystis nidulans, FEBS Lett 394, 153-8.
Boison, G., Schmitz, O., Schmitz, B., and Bothe, H. (1998). Unusual gene arrangement
of the bidirectional hydrogenase and functional analysis of its diaphorase subunit HoxU
in respiration of the unicellular cyanobacterium Anacystis nidulans, Curr Microbiol 36,
253-8.
Bovy, A., de Vrieze, G., Borrias, M., and Weisbeek, P. (1992). Isolation and sequence
analysis of a gene encoding a basic cytochrome c-553 from the cyanobacterium
Anabaena SP. PCC 7937, Plant Molecular Biology 19, 491-492.
Briggs, L. M., Pecoraro, V. L., and McIntosh, L. (1990). Copper-induced expression,
cloning, and regulatory studies of the plastocyanin gene from the cyanobacterium
Synechocystis sp. PCC 6803, Plant Molecular Biology 15, 633-642.
273
Bronstein, M., Schutz, M., Hauska, G., Padan, E., and Shahak, Y. (2000). Cyanobacterial
sulfide-quinone reductase: cloning and heterologous expression, J Bacteriol 182, 3336-
44.
Brooun, A., Bell, J., Freitas, T., Larsen, R. W., and Alam, M. (1998). An archaeal
aerotaxis transducer combines subunit I core structures of eukaryotic cytochrome c
oxidase and eubacterial methyl-accepting chemotaxis proteins, Journal of Bacteriology
180, 1642-1646.
Brosius, J. (1989). Superpolylinkers in cloning and expression vectors, DNA 8, 759-777.
Bryant, D. A., de Lorimier, R., Guglielmi, G., and Stevens, S. E. (1990). Structural and
compositional analyses of the phycobilisomes of Synechococcus sp. PCC 7002. Analyses
of the wild-type strain and a phycocyanin-less mutant constructed by interposon
mutagenesis, Arch Microbiol 153, 550-60.
Bryant, D. A., and Tandeau de Marsac, N. (1988). Isolation of genes encoding
components of the photosynthetic apparatus, Methods of Enzymology 167, 755-765.
Campbell, D., Hurry, V., Clarke, A. K., Gustafsson, P., and Öquist, G. (1998).
Chlorophyll fluorescence analysis of cyanobacterial photosynthesis and acclimation,
Microbiology and Molecular Biology Reviews 62, 667-683.
Carol, P., and Kuntz, M. (2001). A plastid terminal oxidase comes to light: implications
for carotenoid biosynthesis and chlororespiration, Trends Plant Sci 6, 31-36.
Carrasco, C. D., Buettner, J. A., and Golden, J. W. (1995). Programmed DNA
rearrangement of a cyanobacterial hupL gene in heterocysts, Proceedings of the National
Academy of Sciences, USA 2, 791-795.
274
Chapman, G. V., Colman, P. M., Freeman, H. C., Guss, J. M., Murata, M., Norris, V. A.,
Ramshaw, J. A., and Venkatappa, M. P. (1977). Preliminary crystallographic data for a
copper-containing protein, plastocyanin., Journal of Molecular Biology 110, 187-189.
Church, G. M., and Gilbert, W. (1984). Genomic sequencing, Proceedings of the National
Academy of Sciences, USA 81, 1991-1995.
Clarke, A. K., and Campbell, D. (1996). Inactivation of the petE Gene for plastocyanin
lowers photosynthetic capacity and exacerbates chilling-induced photoinhibition in the
cyanobacterium Synechococcus, Plant Physiology 112, 1551-1561.
Colbeau, A., Elsen, S., Tomiyama, M., Zorin, N. A., Dimon, B., and Vignais, P. M.
(1998). Rhodobacter capsulatus HypF is involved in regulation of hydrogenase synthesis
through the HupUV proteins, Eur J Biochem 251, 65-71.
Colbeau, A., Richaud, P., Toussaint, B., Caballero, F. J., Elster, C., Delphin, C., Smith,
R. L., Chabert, J., and Vignais, P. M. (1993). Organization of the genes necessary for
hydrogenase expression in Rhodobacter capsulatus. Sequence analysis and identification
of two hyp regulatory mutants, Molecular Microbiology 8, 15-29.
Dayhoff, M. O., and National Biomedical Research Foundation. (1965). Atlas of protein
sequence and structure, Vol - (Silver Spring, Md., National Biomedical Research
Foundation.).
Deckert, G., Warren, P. V., Gaasterland, T., Young, W. G., Lenox, A. L., Graham, D. E.,
Overbeek, R., Snead, M. A., Keller, M., Aujay, M., et al. (1998). The complete genome
of the hyperthermophilic bacterium Aquifex aeolicus, Nature 392, 353-358.
275
Dernedde, J., Eitinger, M., and Friedrich, B. (1993). Analysis of a pleiotropic gene region
involved in formation of catalytically active hydrogenases in Alcaligenes eutrophus H16,
Archives of Microbiology 159, 545-553.
Drapal, N., and Bock, A. (1998). Interaction of the hydrogenase accessory protein HypC
with HycE, the large subunit of Escherichia coli Hydrogenase 3 during enzyme
maturation, Biochemistry 37, 2941-2948.
Dupuis, A., Peinnequin, A., Chevallet, M., Lundardi, J., Pierrard, B., Procaccio, V., and
Issartel, J. P. (1995). Identification of five Rhodobacter capsulatus genes encoding the
equivalent of ND subunit of the mitochondrial NADH-ubiquinone oxidoreductase., Gene
167, 99-104.
Elhai, J., and Wolk, C. P. (1988). A versatile class of positive-selection vectors based on
the nonviability of palindrome-containing plasmids that allows cloning into long
polylinkers, Gene 68, 119-138.
Ellersiek, U., and Steinmüller, K. (1992). Cloning and transcription analysis of the
ndh(A-I-G-E) gene cluster and ndhD gene of the cyanobacterium Synechocystis sp.
PCC6803, Plant Molecular Biology 20, 1097-1110.
Epel, B. L., and Neumann, J. (1973). The Mechanism of the Oxidation of Ascorbate and
Mn2+ by Chloroplasts The Role of The Radical Superoxide, Biochimica et Biophysica
Acta 325, 520-529.
Fridovich, I., and Hassan, H. M. (1979). Paraquat and the exacerbation of oxygen
toxicity, Trends in Biochemical Sciences 4, 113-115.
276
Friedrich, T., Steinmüller, K., and Weiss, H. (1995). The proton-pumping respiratory
complex I of bacteria and mitochondria and its homologue in chloroplasts, FEBS Letters
367.
Friedrich, T., Strohdeicher, M., Hofhaus, G., Preis, D., Sahm, H., and Weiss, H. (1990).
The same domain motif for ubiquinone reduction in mitochondrial or chloroplast NADH
dehydrogenase and bacterial glucose dehydrogenase., FEBS Letters 265, 37-40.
Friedrich, T., and Weiss, H. (1997). Modular evolution of the respiratory
NADH:ubiquinone oxidoreductase and the origin of its modules, Journal of Theoretical
Biology 187, 529-540.
Fu, C., and Maier, R. J. (1994). Nucleotide sequences of two hydrogenase-related genes
(hypA and hypB) from Bradyrhizobium japonicum, one of which (hypB) encodes an
extremely histidine-rich region and guanine nucleotide-binding domains, Biochimica
Biophysica Acta 1184, 135-138.
Fujinaga, J., and Meyer, J. (1993). The gene encoding [2Fe-2S]ferredoxin from
Clostridium pasteurianum, Biochemical and Biophysical Research Communications 192,
1115-1122.
Fukaya, M., Tayama, K., Tamaki, T., Ebisuya, H., Okumura, H., Kawamura, Y.,
Horinouchi, S., and Beppu, T. (1993). Characterization of a cytochrome a1 that functions
as a ubiquinol oxidase in Acetobacter aceti, Journal of Bacteriology 175, 4307-4314.
Garcia-Horsman, J. A., Barquera, B., Rumbley, J., Ma, J., and Gennis, R. B. (1994a). The
Superfamily of Heme-Copper Respiratory Oxidases, Journal of Bacteriology 176, 5587-
5600.
277
Garcia-Horsman, J. A., Berry, E., Salpleigh, J. P., Alben, J. O., and Gennis, R. B.
(1994b). A novel cytochrome c oxidase from Rhodobacter sphaeroides that lacks CuA,
Biochemistry 33, 3113-3119.
Garcin, E., Vernede, X., Hatchikian, E. C., Volbeda, A., Frey, M., and Fontecilla-Camps,
J. C. (1999). The crystal structure of a reduced [NiFeSe] hydrogenase provides an image
of the activated catalytic center, Structure Fold Des 7, 557-66.
Geerts, D., Bovy, A., de Vrieze, G., Borrias, M., and Weisbeek, P. (1995). Inducible
expression of heterologous genes targeted to a chromosomal platform in the
cyanobacterium Synechococcus sp. PCC 7942, Microbiology 141, 831-841.
Geider, R. J., and Osborne, B. A. (1992). Algal Photosynthesis (New York, Chapman and
Hall).
Ghassemian, M., Wong, B., Ferriera, F., Markley, J. L., and Straus, N. A. (1994).
Cloning, sequencing and transcriptional studies of the genes for cytochrome c-553 and
plastocyanin from Anabaena sp. PCC 7120, Microbiology 140, 1151-1159.
Gierasch, L. M. (1989). Signal sequences, Biochemistry 28, 923-930.
Golden, S. S., Brusslan, J., and Haselkorn, R. (1987). Genetic engineering of the
cyanobacterial chromosome, Methods in Enzymology 153, 215-231.
Grigorieva, G., and Shestakov, S. (1982). Transformation in the cyanobacterium
Synechocystis 6803, FEMS Microbiology Letters 13.
Hattori, M., and Sakaki, Y. (1986). Dideoxy sequencing method using denatured plasmid
templates, Anal Biochem 152, 232-8.
Henikoff, S., and Henikoff, J. G. (1992). Amino acid substitution matrices from protein
blocks, Proc Natl Acad Sci U S A 89, 10915-9.
278
Herbert, S. K., Samson, G., Fork, D. c., and Laudenbach, D. E. (1992). Characterization
of damage to photosystems I and II in a cyanobacterium lacking detectable iron
superoxide dismutase activity, Proceedings of the National Academy of Sciences, USA
89, 8716-8720.
Hippler, M., Drepper, F., Rochaix, J. D., and Muhlenhoff, U. (1999). Insertion of the N-
terminal part of PsaF from Chlamydomonas reinhardtii into photosystem I from
Synechococcus elongatus enables efficient binding of algal plastocyanin and cytochrome
c6., Journal of Biological Chemistry 274, 4180-4188.
Hiratsuka, J., Shimada, H., Whittier, R., Ishibashi, T., Sakamoto, M., Mori, M., Kondo,
C., Honji, Y., Sun, C. R., Meng, B. Y., and al, e. (1989). The complete sequence of the
rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct
tRNA genes accounts for a major plastid DNA inversion during the evolution of the
cereals., Molecular General Genetics 217, 185-194.
Ho, K. K., and Krogmann, D. W. (1984). Electron donors to P700 in cyanobacteria and
algae an instance of unusual genetic variability, Biochimica et Biophysica Acta 766, 310-
316.
Hochkoeppler, A., Jenney, J., F. E., Lang, S. E., Zannoni, D., and Daldal, F. (1995).
Membrane-associated cytochrome cy of Rhodobacter capsulatus is an electron carrier
from the cytochrome bc1 complex to the cytochrome c oxidase during respiration,
Journal of Bacteriology 177, 608-613.
Hosler, J. P., Fetter, J., Tecklenberg, M. M. J., Espe, M., Lerma, C., and Ferguson-Miller,
S. (1992). Cytochrome aa3 of Rhodobacter sphaeroides as a model for mitochondrial
279
cytochrome c oxidase: purification, kinetics, proton pumping and spectral analysis,
Journal of Biological Chemistry 267, 24264-24272.
Houchins, J. P., and Burris, R. H. (1981). Comparative characterization of two distinct
hydrogenases from Anabaena sp. strain 7120., Journal of Bacteriology 146, 215-221.
Howitt, C. A., Udall, P. K., and Vermaas, W. F. J. (1999). Type 2 NADH dehydrogenase
in the cyanobacterium Synechocystis sp. strain PCC 6803 are involved in regulation
rather than respiration, Journal of Bacteriology 181, 3994-4003.
Howitt, C. A., and Vermaas, W. F. J. (1998). Quinol and Cytochrome Oxidases in the
Cyanobacterium Synechocystis sp. PCC 6803, Biochemistry 37, 17944-17951.
Howitt, C. A., and Vermaas, W. F. J. (1999). Subunits of the NAD(P)-reducing nickel-
containing hydrogenase do not act as part of the type-1 NAD(P)H-dehydrogenase in the
cyanobacterium
Synechocystis sp. PCC 6803. In The Phototrophic Prokaryotes, G. A. Peschek, W.
Löffelhardt, and G. Schmetterer, eds. (New York, Kluwer Academic/Plenum Publishers),
pp. 595-601.
Howitt, C. A., Whelan, J., Price, G. D., and Day, D. A. (1996). Cloning, analysis and
inactivation of the ndhK gene encoding a subunit of NADH quinone oxidoreductase from
Anabaena PCC 7120, European Journal of Biochemistry 340, 173-180.
Kazusa DNA Research Institute. (2000). Anabaena sp. PCC 7120 genome sequence
(http://www.kazusa.or.jp/cyano/cyano.html).
Jacobi, A., Rossman, R., and Böck, A. (1992). The hyp operon gene products are required
for the maturation of catalytically active hydrogenase isoenzymes in Escherichia coli,
Archives of Microbiology 158, 44-451.
280
Jenney, J., F. E., and Daldal, F. (1993). A novel membrane-associated c-type cytochrome,
cyt cy, can mediate the photosynthetic growth of Rhodobacter capsulatus and
Rhodobacter sphaeroides, The EMBO Journal 12, 1283-1292.
Jenney, J., F. E., Prince, R. C., and Daldal, F. (1996). The membrane-bound cytochrome
cy of Rhodobacter capsulatus can serve as an electron donor to the photosynthetic
reaction center of Rhodobacter sphaeroides, Biochimica et Biophysica Acta 1273, 159-
164.
Kaneko, T., Sato, S., Kotani, H., Tanaka, A., Asamizu, E., Nakamura, Y., Miyajima, N.,
Hirosawa, M., Sugiura, M., Sasamoto, S., et al. (1996). Sequence analysis of the genome
of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II. Sequence
determination of the entire genome and assignment of potential protein-coding regions.,
DNA Research 3, 109-136.
Kelly, M. J. S., Poole, R. K., Yates, M. G., and Kennedy, C. (1990). Cloning and
mutagenesis of genes encoding the cytochrome bd terminal oxidase complex in
Azotobacter vinelandii: mutants deficient in the cytochrome d complex are unable to fix
nitrogen in air, Journal of Bacteriology 172, 6010-6019.
Kerfeld, C. A., Anwar, H. P., Interrante, R., Merchant, S., and Yeates, T. O. (1995). The
structure of chloroplast cytochrome c6 at 1.9 A resolution: evidence for functional
oligomerization., Journal of Molecular Biology 250, 627-47.
Kerfeld, C. A., and Krogmann, D. W. (1998). Photosynthetic Cytochromes c in
Cyanobacteria, Algae, and Plants, Annual Review of Plant Physiology, Plant Molecular
Biology 49, 397-425.
281
Klughammer, B., Sültemeyer, D., Badger, M. R., and Price, G. D. (1999). The
involvement of NAD(P)H dehydrogenase subunits, NdhD3 and NdhF3, in high-affinity
CO2 uptake in Synechococcus sp. PCC7002 gives evidence for multiple NDH-1
complexes with specific roles in cyanobacteria, Molecular Microbiology 32, 13051315.
Kummerle, R., Atta, M., Scuiller, J., Gaillard, J., and Meyer, J. (1999). Structural
similarities between the N-terminal domain of Clostridium pasteurianum hydrogenase
and plant-type ferredoxins., Biochemistry 38, 1938-1943.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head
of bacteriophage T4, Nature 227, 680-685.
Laudenbach, D. E., Herbert, S. K., McDowell, C., Fork, D. C., Grossman, A. R., and
Straus, N. A. (1990). Cytochrome c-553 Is not required for photosynthetic activity in the
cyanobacterium Synechococcus, The Plant Cell 2, 913-924.
Lauraeus, M., Haltia, T., Saraste, M., and Wikström, M. (1991). Bacillus subtilis
expresses two kinds of haem-A-containing terminal oxidases, European Journal of
Biochemistry 197.
Lockau, W. (1981). Evidence for a dual role of cytochrome c-553 and plastocyanin in
photosynthesis and respiration of the cyanobacterium, Anabaena variabilis., Archives of
Microbiology 128, 336-340.
Lutz, S., Jacobi, A., Schlensog, V., Bohm, R., Sawers, G., and Bock, A. (1991).
Molecular characterization of an operon (hyp) necessary for the activity of the three
hydrogenase isoenzymes in Escherichia coli, Mol Microbiol 5, 123-35.
282
Magalon, A., and Bock, A. (2000). Analysis of the HypC-HycE complex, a key
intermediate in the assembly of the metal center of the Escherichia coli hydrogenase 3.,
Journal of Biological Chemistry 275, 21114-211120.
Malakhov, M. P., Malakhova, O. A., and Murata, N. (1999). Balanced regulation of
expression of the gene for cytochrome cM and that of genes for plastocyanin and
cytochrome c6 in Synechocystis, FEBS Lett 444, 281-4.
Malki, S., Saimmaime, I., De Luca, G., Rousset, M., Dermoun , Z., and Belaich, J. P.
(1995). Characterization of an operon encoding an NADP-reducing hydrogenase in
Desulfovibrio fructosovorans, Journal of Bacteriology 177, 2628-2636.
Manna, P., and Vermaas, W. (1997). Lumenal proteins involved in respiratory electron
transport in the cyanobacterium Synechocystis sp. PCC 6803, Submitted.
Marco, E., Ohad, N., Schwarz, R., Lieman-Hurwitz, J., Gabay, C., and Kaplan, A.
(1993). High CO2 concentration alleviates the block in photosynthetic electron transport
in an
ndhB-inactivated mutant of Synechococcus sp. PCC 7942, Plant Physiology 101, 1047-
1053.
Massanz, C., Fernandez, V. M., and Friedrich, B. (1997). C-terminal extension of the H2-
activating subunit, HoxH, directs maturation of the NAD-reducing hydrogenase in
Alcaligenes eutrophus, Eur J Biochem 245, 441-8.
Massanz, C., Schmidt, S., and Friedrich, B. (1998). Subforms an In Vitro Reconstitution
of the NAD-Reducing Hydrogenase of Alcaligenes eutrophus, Journal of Bacteriology
180, 1023-1029.
283
Matsubayashi, T., Wakasugi, T., Shinozaki, K., Yamaguchi-Shinozaki, K., Zaita, N.,
Hidaka, T., Meng, B. Y., Ohto, C., Tanaka, M., Kato, A., and et al. (1987). Six
chloroplast genes (ndhA-F) homologous to human mitochondrial genes encoding
components of the respiratory chain NADH dehydrogenase are actively expressed:
determination of the splice sites in ndhA and ndhB pre-mRNAs, Mol Gen Genet 210,
385-93.
Maxwell, P. C., and Biggins, J. (1976). Role of cyclic electron transport in photosynthesis
as measured by the photoinduced turnover of P700 in vivo, Biochemistry 15, 3975-3981.
Merberg, D., O'hara, E. B., and Maier, R. J. (1983). Regulation of hydrogenase in
Rhizobium japonicum : Analysis of mutants altered in regulation by carbon substrates and
oxygen, Journal of Bacteriology 156, 1236-1242.
Merchant, S., Hill, K., Kim, J. H., Thompson, J., Zaitlin, D., and Bogorad, L. (1990).
Isolation and characterization of a complementary DNA clone for an algal pre-
apoplastocyanin, J Biol Chem 265, 12372-9.
Meyer, J., and Gagnon, J. (1991). Primary structure of hydrogenase I from Clostridium
pasteurianum, Biochemistry 30, 9697-704.
Minghetti, K. C., Goswitz, V. C., Gabriel, N. E., Hill, J. J., Barassi, C., Gerogiou, C. D.,
Chan, S. I., and Gennis, R. B. (1992). Modified, large-scale purification of the
cytochrome o complex of Escherichia coli yields a two heme/one copper terminal
oxidase with high specific activity, Biochemistry 31, 6917-6924.
Muñiz, W. H. (1993) Genetic engineering of Agmenellum quadruplicatum PR-6:
Deletion of the AquI restriction system and creation of a chromosomal cloning vector,
PhD, The Pennsylvania State University, University Park.
284
Muth, E., Morschel, E., and Klein, A. (1987). Purification and characterization of an 8-
hydroxy-5-deazaflavin-reducing hydrogenase from the archaebacterium Methanococcus
voltae, Eur J Biochem 169, 571-7.
Nelson, K. E., Clayton, R. A., Gill, S. R., Gwinn, M. L., Dodson, R. J., Haft, D. H.,
Hickey, E. K., Peterson, J. D., Nelson, W. C., Ketchum, K. A., et al. (1999). Evidence for
lateral gene transfer between Archaea and bacteria from genome sequence of
Thermotoga maritima, Nature 399, 323-329.
Nielsen, H., Engelbrecht, J., Brunak, S., and von Heijne, G. (1997). Identification of
prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites, Protein
Engineering 10, 1-6.
O'gara, J. P., Eraso, J. M., and Kaplan, S. (1998). A redox-responsive pathway for
aerobic regulation of photosynthesis gene expression in Rhodobacter sphaeroides 2.4.1,
Journal of Bacteriology 180, 4044-4050.
O'gara, J. P., and Kaplan, S. (1997). Evidence for the role of redox carriers in
photosynthesis gene expression and carotenoid biosynthesis in Rhodobacter sphaeroides
2.4.1, Journal of Bacteriology 179, 1951-1961.
Obinger, C., Knepper, J. C., Zimmermann, U., and Peschek, G. A. (1990). Identification
of a periplasmic c-type cytochrome as electron donor to the plasma membrane-bound
cytochrome oxidase of the cyanobacterium Nostoc mac, Biochem Biophys Res Commun
169, 492-501.
Ogawa, T. (1990). Cloning and inactivation of a gene essential to inorganic carbon
transport of Synechocystis PCC6803, Plant Physiology 96, 280-284.
285
Ogawa, T. (1991). A gene homologous to the subunit-2 gene of NADH dehydrogenase is
essential to inorganic carbon transport of Synechocystis PCC6803, Proceedings of the
National Academy of Sciences USA 88, 4275-4279.
Ohkawa, H., Price, G. D., Badger, M. R., and Ogawa, T. (2000). Mutation of the ndh
genes leads to inhibition of CO2 uptake rather than HCO3- uptake in Synechocystis sp.
strain PCC 6803, Journal of Bacteriology 182, 2591-2596.
Ohyama, K., Fukuzawa, H., Kohchi, T., Shirai, H., Sano, T., Sano, S., Umesono, K.,
Shiki, Y., Takeuchi, M., Chang, Z., et al. (1986). Chloroplast gene organization deduced
from complete sequence of liverwort Marchantia polymorpha chloroplast DNA, Nature
322, 572-574.
Okada, S., Kanematsu, S., and Asada, K. (1979). Intracellular Distribution of Manganese
and Ferric Superoxide Dismutases in Blue-Green Algae, FEBS Letters 103, 106-110.
Olson, J. W., Fu, C., and Maier, R. J. (1997). The HypB protein from Bradyrhizobium
japonicum can store nickel and is required for the nickel-dependent transcriptional
regulation of hydrogenase, Molecular Microbiology 24, 119-128.
Olson, J. W., and Maier, R. J. (1997). The sequences of hypF hypC and hypD complete
the hyp gene cluster required for hydrogenase activity in Bradyrhizobium japonicum,
Gene, 93-99.
Olson, J. W., Mehta, N. S., and Maier, R. J. (2001). Requirement of nickel metabolism
proteins HypA and HypB for full activity of both hydrogenase and urease in Helicobacter
pylori, Molecular Microbiology 39, 176-182.
286
Oxelfelt, F., Tamagnini, P., and Lindblad, P. (1998). Hydrogen uptake in Nostoc sp. PCC
73102. Cloning and characterization of a hupSL homologue, Archives of Microbiology
169, 267-274.
Papen, H., Kentemich, T., Schmulling, T., and Bothe, H. (1986). Hydrogenase activities
in cyanobacteria, Biochimie 68, 121-132.
Peschek, G. A., Wastyn, M., Trnka, M., Molitor, V., Fry, I. V., and Packer, L. (1989).
Characterization of the cytochrome c oxidase in isolated and purified plasma membranes
from the cyanobacterium Anacystis nidulans, Biochemistry 28, 3057-63.
Peters, J. W., Lanzilotta, W. N., Lemon, B. J., and Seefeldt, L. C. (1998). X-ray crystal
structure of the Fe-only hydrogenase (CpI) from Clostridium pasteurianum to 1.8
angstrom resolution, Science 282, 1853-8.
Prentki, P., and Krisch, H. M. (1984). In vitro insertional mutagenesis with a selectable
DNA fragment, Gene 29, 303-313.
Price, G. D., Sültemyer, D., Klughammer, B., Ludwig, M., and Badger, M. R. (1998).
The functioning of the CO2 concentrating mechanism in several cyanobacterial strains: a
review of general physiological characteristics, genes, proteins, and recent advances,
Canadian Journal of Botany 76.
Puustinen, A., Finel, M., Haltia, T., Gennis, R. B., and Wikström, M. (1991). Properties
of two terminal oxidases of Escherichia coli, Biochemistry 30, 3936-3942.
Redinbo, M. R., Cascio, D., Choukair, M. K., Rice, D., Merchant, S., and Yeates, T. O.
(1993). The 1.5-Å Crystal Structure of Plastocyanin from the Green Alga
Chlamydomonas reinhardtii, Biochemistry 32, 10560-10567.
287
Rey, L., Murillo, J., Hernando, Y., Hidalgo, E., Cabrera, E., Imperial, J., and Ruiz-
Argueso, T. (1993). Molecular analysis of a microaerobically induced operon required for
hydrogenase synthesis in Rhizobium leguminosarum biovar viciae, Mol Microbiol 8, 471-
81.
Rippka, R. (1972). Photoheterotrophy and chemoheterotrophy among blue-green algae,
Archives of Microbiology 87, 91-98.
Rousset, M., Dermoun, Z., Hatchikian, C. E., and Belaich, J. P. (1990). Cloning and
sequencing of the locus encoding the large and small subunit genes of the periplasmic
[NiFe]hydrogenase from Desulfovibrio fructosovorans, Gene 94, 95-101.
Sakamoto, T., and Bryant, D. A. (1997). Temperature-regulated mRNA accumulation
and stabilization for fatty acid desaturase genes in the cyanobacterium Synechococcus sp.
strain PCC 7002, Molecular Microbiology 23, 1281-1292.
Sakamoto, T., and Bryant, D. A. (1998). Growth at low temperature causes nitrogen
limitation in the cyanobacterium Synechococcus sp. PCC 7002, Archives of
Microbiology 169, 10-19.
Sakamoto, T., Delgazio, V. B., and Bryant, D. A. (1998). Growth on urea can trigger
death and peroxidation of the cyanobacterium Synechococcus sp. strain PCC 7002,
Applied and Environmental Microbiology 64, 2361-2366.
Sambrook, J., Fritsch, E., and Maniatis, T. (1989). Molecular Cloning: A Laboratory
Manual, 2nd ed. edn (Cold Spring Harbor, NY, Cold Spring Harbor Laboratory Press).
Sandmann, G., and Boger, P. (1981). Plastocyanin and cytochrome c- 553, two different
electron donors to photosystem I in algae. In Photosynthesis II. Electron Transport and
288
Photophosphorylation, P. Akoyunoglou, ed. (Philadelphia, Balaban International Science
Services), pp. 623-632.
Saraste, M. (1990). Structural features of cytochrome oxidase, Quarterly Review of
Biophysics 23, 331-366.
Scawen, M. D., Ramshaw, J. A., and Boulter, D. (1975). The amino acid sequence of
plastocyanin from spinach. (Spinacia oleracea L.), Biochem J 147, 343-9.
Scherer, S. (1990). Do photosynthetic and respiratory electron transport chains share
redox proteins?, Trends in Biochemical Sciences 15, 458-462.
Schiefer, W., and Happe, T. (1999). Isolation of the ndhAIGE gene cluster of Anabaena
sp. PCC 7120 (DBSOURCE embl locus APC242867, accession AJ242867.1).
Schluchter, W. M., Zhao, J., and Bryant, D. A. (1993). Isolation and characterization of
the ndhF gene of Synechococcus sp. strain PCC 7002 and initial characterization of an
interposon mutant, J Bacteriol 175, 3343-52.
Schmetterer, G. (1994). Cyanobacterial Respiration. In The Molecular Biology of
Cyanobacteria, D. A. Bryant, ed. (Dorderecht, The Netherlands, Kluwer Academic
Publishers), pp. 408-435.
Schmitz, O., Boison, G., Hilscher, R., Hundeshagen, B., Zimmer, W., Lottspeich, F., and
Bothe, H. (1995). Molecular biological analysis of a bidirectional hydrogenase from
cyanobacteria, Eur J Biochem 233, 266-76.
Schmitz-Linneweber, C., Alcaraz, J. P., Cottet, A., Maier, R. M., Herrmann, R. G., and
Mache, R. (2000). Spinach chloroplast genes.
289
Schreiber, U., Schliwa, U., and Bilger, W. (1986). Continuous recording of
photochemical and non-photochemical chlorophyll fluorescence quenching with a new
type of modulation fluorometer, Photosynthesis Research 10, 51-62.
Schubert, H., Matthijs, H. C. P., and Mur, L. R. (1995). In vivo assay of P700 redox
changes in the cyanobacterium Fremyella diplosiphon and the role of cytochrome c
oxidase in regulation of photosynthetic electron transfer, Photosynthetica 31, 517-527.
Schwartz, E., Gerischer, U., and Friedrich, B. (1998). Transcription regulation of
Alcaligenes eutrophus, Journal of Bacteriology 180, 3197-3204.
Seefeldt, L. C., and Arp, D. J. (1986). Purification to homogeneity of Azotobacter
vinelandii hydrogenase: a nickel and iron containing alpha beta dimer, Biochimie 68, 25-
34.
Shikanai, T., Endo, T., Hashimoto, T., Yamada, Y., Asada, K., and Yokota, A. (1998).
Directed disruption of the tobacco ndhB gene impairs cyclic electron flow around
photosystem I, Proc Natl Acad Sci U S A 95, 9705-9.
Shimada, H., and Sugiura, M. (1991). Fine structural features of the chloroplast genome:
comparison of the sequenced chloroplast genomes, Nucleic Acids Res 19, 983-95.
Sone, N., Tano, H., and Ishizuka, M. (1993). The genes in the thermophilic
cyanobacterium Synechococcus vulcanus encoding cytochrome-c oxidase [published
erratum appears in Biochim Biophys Acta 1994 Apr 28;1188(2):255], Biochim Biophys
Acta 1183, 130-8.
South, T. L., and Summers, M. F. (1990). Zinc fingers (Salem, Massachusetts, Elsevier
Science Publications).
290
Stanier, R., and Cohen-Bazire, G. (1977). Phototrophic prokaryotes: the cyanobacteria,
Annu Rev Microbiol 31, 225-274.
Stanier, R. Y., Kunisawa, R., Mandel, M., and Cohen-Bazire, G. (1971). Purification and
properties of unicellular blue-green algae (order Chroococcales), Bacteriology Reviews
35, 171-205.
Steigerwald, V. J., Beckler, G. S., and Reeve, J. N. (1990). Conservation of hydrogenase
and polyferredoxin structures in the hyperthermophilic archaebacterium Methanothermus
fervidus, J Bacteriol 172, 4715-8.
Stevens, J., S. E., and Porter, R. D. (1980). Transformation in Agmenellum
quadruplicatum., National Academy of Sciences Proceedings USA 10, 6052-6056.
Stevens, S. E., and van Baalen, C. (1973). Characteristics of nitrate reduction in a mutant
of the blue-green alga Agmenellum quadruplicatum, Plant Physiology 51, 350-356.
Storz, G., and Imlay, J. A. (1999). Oxidative Stress, Current Opinion in Microbiology 2,
188-194.
Studier, F. W., Rosenberg, A. H., Dunn, J. J., and Dubendorff, J. W. (1990). Use of T7
RNA Polymerase to direct expression of cloned genes, Methods in Enzymology 185, 60-
89.
Stürzl, E., Scherer, S., and Böger, P. (1982). Reconstitution of electron transport by
cytochrome c-553 in a cell-free system of Nostoc muscorum, Photosynthesis Research 3,
191-201.
Sugita, M., Lijun, L., Ohta, M., and Itadani, H. (1995). Genes encoding the group I
intron-containing tRNALeu and subunit L of NADH dehydrogenase from the
cyanobacterium Synechococcus PCC 6301, DNA Research 2, 71-76.
291
Sültemeyer, D., Price, G. D., Bryant, D. A., and Badger, M. R. (1997b). PsaE- and NdhF-
mediated electron transport affect bicarbonate transport rather than carbon dioxide uptake
in the cyanobacterium Synechococcus sp. PCC 7002, Planta 201, 36-42.
Tabor, S., and Richardson, C. C. (1987a). DNA sequence analysis with a modified
bacteriophage T7 DNA polymerase, Proceedings of the National Academy of Sciences
84, 4767-4771.
Tamagnini, P., Troshina, O., Oxelfelt, F., Salema, R., and Lindblad, P. (1997).
Hydrogenases in Nostoc sp. strain PCC 73102, a strain lacking a bidirectional enzyme,
Applied and Environmental Microbiology 63, 1801-1807.
Tano, H., Ishizuka, M., and Sone, N. (1991). The cytochrome C oxidase genes in blue-
green algae and characteristics of the deduced protein sequence for subunit II of the
thermophilic cyanobacterium Synechococcus vulcanus, Biochem Biophys Res Commun
181, 437-42.
Thiaglingam, S., and Yang, T. (1993). Purification and characterization of NADH
dehydrogenase from Bacillus megaterium, Canadian Journal of Microbiology 39.
Thomas, P. E., Ryan, D., and Levin, W. (1976). An improved staining procedure for the
detection of the peroxidase activity of cytochrome p-450 on sodium dodecyl sulfate
polyacrylamide gels, Analytical Biochemistry 75, 168-176.
Thompson, J. D., Higgins, D. G., and Gibson, T. J. (1994a). Clustal w: improving the
sensitivity of progressive multiple sequence alignment through sequence weighting,
position-specific gap penalties and weight matrix choice, Nucleic Acids Research 22,
4673-4680.
292
Thompson, J. D., Higgins, D. G., and Gibson, T. J. (1994b). CLUSTAL W: improving
the sensitivity of progressive multiple sequence alignment through sequence weighting,
position-specific gap penalties and weight matrix choice, Nucleic Acids Res 22, 4673-80.
Tran-Betcke, A., Warnecke, U., Bocker, C., Zaborosch, C., and Friedrich, B. (1990).
Cloning and nucleotide sequences of the genes for the subunits of NAD-reducing
hydrogenase of Alcaligenes eutrophus H16, Journal of Bacteriology 117276, 2920-2929.
Trinder, P. (1966). Determination of Glucose in Blood using Glucose Oxidase with an
alternative oxygen acceptor, Annals of Clinical Biochemistry 24, 24-28.
Trnka, M., and Peschek, G. A. (1986). Immunological identification of aa3-type
cytochrome oxidase in membrane preparations of the cyanobacterium Anacystis nidulans,
Biochem Biophys Res Commun 136, 235-41.
Van Baalen, C. (1962). Studies on marine blue-green algae, Botanica Marina 4, 129-139.
Volbeda, A., Charon, M. H., Piras, C., Hatchikian, E. C., Frey, M., and Fontecilla-
Camps, J. C. (1995). Crystal structure of the nickel-iron hydrogenase from Desulfovibrio
gigas, Nature 373, 580-587.
von Heijne, G. (1986). A new method for predicting signal sequence cleavage sites,
Nucleic Acids Research 14, 4683-4690.
Walker, J. E. (1992a). The NADH:ubiquinone oxidoreductase (complex I) of respiratory
chains, Q Rev Biophys 25, 253-324.
Walker, J. E. (1992b). The NADH:ubiquinone oxidoreductase (complex I) of respiratory
chains, Q Rev Biophys 25, 253-324.
Walker, J. E., Arizmendi, J. M., Dupuis, A., Fearnley, I. M., Finel, M., Medd, S. M.,
Pikington, S. J., Runswick, M. J., and Skehel, J. M. (1992). Sequences of 20 subunits of
293
NADH:ubiquinone oxidoreductase from bovine heart mitochondria, Journal of Molecular
Biology 226, 1051-1072.
Wastyn, M., Achatz, A., Trnka, M., and Peschek, G. A. (1987). Immunological and
spectral characterization of partly purified cytochrome oxidase from the cyanobacterium
Synechocystis 6714, Biochem Biophys Res Commun 149, 102-11.
Weidner, U., Geier, S., Ptock, A., Friedrich, T., Leif, H., and Weiss, H. (1993). The gene
locus of the proton-translocating NADH: ubiquinone oxidoreductase in Escherichia coli.
Organization of the 14 genes and relationship between the derived proteins and subunits
of mitochondrial complex I, Journal of Molecular Biology 233, 109-122.
Weiss, H., Friedrich, T., Hofhaus, G., and Preis, D. (1991). The respiratory-chain NADH
dehydrogenase (complex I) of mitochondria, European Journal of Biochemistry 197, 563-
576.
Winterbourn, C. C., Hawkins, R. E., Brian, M., and Carrel, R. W. (1975). The estimation
of red cell superoxide dismutase activity, Journal of Laboratory Clinical Medicine 85,
337-341.
Wolf, I., Buhrke, T., Dernedde, J., Pohlmann, A., and Friedrich, B. (1998). Duplication of
hyp genes involved in maturation of [NiFe] hydrogenases in Alcaligenes eutrophus H16,
Arch Microbiol 170, 451-9.
Wu, L.-F., and Mandrand, M. A. (1993). Microbial hydrogenases: Primary structure,
classification, signatures and phylogeny, FEMS Microbiolog Reviews 104, 243-270.
Xu, X., Koyama, N., Cui, M., Yamagishi, A., Nosoh, Y., and Oshima, T. (1991).
Nucleotide sequence of the gene encoding NADH dehydrogenase from an alkalphile,
Bacillus sp. strain YN-1, Journal of Biochemistry 109.
294
Yagi, T., Hon-nami, K., and Ohnishi, T. (1988). Purification and characterization of two
types of NADH-quinone reductases from Thermus thermophilusHB-8, Biochemistry 27.
Yagi, T., Yano, T., Di Bernardo, S., and Matsuno-Yagi, A. (1998). Procaryotic complex I
(NDH-1), an overview, Biochimica et Biophysica Acta 1364, 125-133.
Yanisch-Perron, E., Vieira, J., and Messing, J. (1985). Improved M13 phage cloning
vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors, Gene
33, 103-119.
Yu, L., Zhao, J., Mühlenhoff, U., Bryant, D. A., and Golbeck, J. H. (1993). PsaE is
Required for in Vivo Cyclic Electron Flow around Photosystem I in the Cyanobacterium
Synechococcus sp. PCC 7002, Plant Physiology 103.
Yun, C.-H., Beci, R., Crofts, A. R., Kaplan, S., and Gennis, R. B. (1990). Cloning and
DNA sequencing of the fbc operon encoding the cytochrome bc1 complex from
Rhodobacter sphaeroides: characterization of the fbc deletion mutants, and
complementation by a site-specific mutational variant, European Journal of Biochemistry
194, 399-411.
Zhang, L., McSpadden, B., Pakrasi, H. B., and Whitmarsh, J. (1992). Copper-mediated
Regulation of Cytochrome c553 and Plastocyanin in the Cyanobacterium Synechocystis
6803, The Journal of Biological Chemistry 267, 19054-19059.
Zhang, L., Pakrasi, H. B., and Whitmarsh, J. (1994). Photoautotrophic growth of the
cyanobacterium Synechocystis sp. PCC 6803 in the absence of cytochrome c553 and
plastocyanin, Journal of Biological Chemistry 269, 5036-5042.
295
Zuber, M., Harker, A. R., Sultana, M. A., and Evans, H. J. (1986). Cloning and
expression of Bradyrhizobium japonicum uptake hydrogenase structural genes in
Escherichia coli, Proc Natl Acad Sci U S A 83, 7668-72.
296
APPENDIX A: RESTRICTION MAPS OF ELECTRON TRANSFER GENES
Restriction maps of Synechococcus sp. PCC 7002 electron transport genes
The restriction enzyme maps for all Synechococcus sp. PCC 7002 genes/clones
reported for this study are shown in this appendix. For details, please refer to RESULTS.
296
Type I NADH dehydrogenase genes from Synechococcus sp. PCC 7002
A1. Restriction map of the Synechococcus sp. PCC 7002 ndhAIGE gene cluster
200 400 600 800 1000 1200 1400 1600 1800 2000 2200 2400 2600 2800 3000 3200 3400 3600
ndhA ndhGndhI ndhEfraH (slr0298 homol.) hypo. prot. (slr1536 homol.)
NdeI
PvuI
SspIPvuI
StyI
DraI
StyI
DraI
DraI
AvaI
HindIII
SacI
SmaI
AvaI
PvuI
PvuI HincII
StyI
StyI
SspI
HincII SspI
StyI StyI
HindIII
StyI
StyI
DraI
StyI
StyI
StyI
HindIII
DraI
PvuI
StyI
EcoRI
EcoRI
297
A2. Restriction map of the Synechococcus sp. PCC 7002 ndhB gene
300 600 900 1200 1500 1800 2100 2400 2700
ndhB Ferredoxin sll1430
AflIIBglII
XbaI
SapI
AflIII
BamHI SspI
AvaII EcoRI
BglIEcoRI
298
A3. Restriction map of the Synechococcus sp. PCC 7002 ndhD1 gene
200 400 600 800 1000 1200 1400 1600 1800 2000 2200
ndhD1
P1174 (1492<-1512)
HincII BamHI
SspI
PvuISspI
HindIII
AvaIPvuI
NcoI
StyI
PvuI
BglII
A4. Restriction map of the Synechococcus sp. PCC 7002 ndhD2 gene
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300
ndhD2sigG sl r1546XbaI XbaIStyIStyI
PvuIPvuINcoI
StyI
AvaI
SmaI
AvaI
SmaI
HincII
HindIII
HincII
StyI
PvuI
HincII
StyI
NcoI
StyI
StyI
PvuI
299
A4. Restriction map of the Synechococcus sp. PCC 7002 ndhF3 ndhD3 genes. Data from (Klughammer et al., 1999).
400 800 1200 1600 2000 2400 2800 3200 3600 4000 4400 4800 5200 5600 6000 6400
ndhF3 ndhD3 sll1734 rbcR (sll1594) sll1735 XbaIEcoRI
PvuI
HincIIStyI
StyI
PstI
HincII
StyI
SspI
EcoRV
XbaI
StyI PvuIPvuI
StyI EcoRV
PvuI
HindIII
EcoRI
DraI
NdeI
BglII
SspI
DraI
HincIIStyI StyI
PvuI
StyI
EcoRI
StyI
NcoI
StyI
PvuI
EcoRV
PvuI
PvuI
KpnI
PvuI
PstI
NcoI
StyI
KpnI
HindIIIStyI
EcoRI
PvuIDraI
HincII
SspI
300
A5. Restriction map of the Synechococcus sp. PCC 7002 ndhF4 ndhD4 genes.
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600 3900 4200 4500
ndhF4 ndhD4 slr1302 XbaI HindIII
SspIStyI
AvaI
HincII
StyI
KpnI
EcoRV
PvuIPvuIKpnI
StyI
SspI
PvuIPvuI
SspI DraI
StyI
PvuI
NcoI
StyI
EcoRIHincII NcoI
StyI
PvuI
StyI
StyI
SacI
KpnI
PvuI
HincII
StyI
301
A6. Restriction map of the Synechococcus sp. PCC 7002 ndhF1 gene. Data from (Schluchter et al., 1993).
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600 3900 4200 4500
ndhF1 sll0175 ssl3451 petHEcoRI
DraINcoI
StyI
StyI PvuI
SspI
StyI
HincII
HincII
StyI
SmaI
AvaI
NcoI
StyI
HincII
PstI PvuIAvaIPvuI
PvuI
StyI
NcoI
StyI
StyI
NcoI
StyI
PvuI
StyI
NcoI
StyI
DraINcoI
StyI
BamHI KpnI
EcoRV
NcoI
StyI
HindIII
302
A7. Restriction map of the Synechococcus sp. PCC 7002 ndhF5 and ndhF2 genes
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600 3900 4200 4500 4800
ndhF5 ndhF2Na+/H+ antiporter (sll0689) slr2010
slr2006
ssr3410
AvaI
SmaI
StyI
StyI
DraI
StyI SspI
HincII
AvaI
SmaI
DraI
BamHI
EcoRI NcoI
StyI
StuI
NcoI
StyI
SspI
PvuI
StuI
StuI
StyI
PvuI
PvuIXbaI
SspI
SspI
NcoI
StyI
PvuI
StyI
PvuI
NcoI
StyI
StyI
PvuI
NcoI
StyI
StyI StyI
PvuI
NcoI
StyI
DraI
AvaI
SmaI
SspI PvuI
StyI
HindIII
BglII
HindIII
AvaI
SmaI
HincII
303
A8. Restriction map of the Synechococcus sp. PCC 7002 ndhCKJ gene cluster
500 1000 1500 2000 2500 3000 3500
HP, sll0606 homol.ndhK ndhJndhC
HP Vng2274c (Halobacterium sp. NRC-1)
sll0997 BamHI
BamHIPvuI XbaI
HindIII
StyI
PvuI
PvuI
StyI
StyI
StyI
NcoI
StyI
KpnIDraI
PvuI
HincII
HincII
StuISspI
SspI
DraIPvuI StyI
KpnI
BglIISspI
StyI
StyI
BglII
StyI PvuI
NcoI
StyI
NcoI
StyI
304
A9. Restriction map of the Synechococcus sp. PCC 7002 ndhH gene
200 400 600 800 1000 1200 1400 1600 1800 2000 2200
ndhH serine esterase
sl l0355BamHI
BamHIStyI
PvuI
EcoRV
PvuI
EcoRI
PvuIStyI
SspI
PvuI
HincII
305
A10. Restriction map of the Synechococcus sp. PCC 7002 ndhL gene
100 200 300 400 500 600 700 800 900 1000 1100 1200 1300 1400 1500 1600 1700
ndhL
TrpA (slr0966)slr0815 HindIII HindIII
NcoI
StyI
PvuI
StyI
StyI
PvuIBamHIEcoRV StyI
PvuI
PstI NcoI
StyI
PvuI
306
Type II NADH dehydrogenase genes from Synechococcus sp. PCC 7002
A11. Restriction maps of the Synechococcus sp. PCC 7002 ndbA and ndbB genes
307
200 400 600 800 1000 1200 1400 1600 1800
ndbA (slr0851 homolog)
HindIII - HindIII construct (pUC19), 2111bp
EcoNI
NcoI
StyI
AvaII XmnI
Afl I I I
BstXI
PvuI
200 400 600 800 1000 1200 1400 1600 1800
ndbB (slr1743 homolog)
HindIII - BamHI construct (pBluescript II SK+)
HindIII BamHIBglIAclI
XcmI
BglII
HaeII
NcoI
RsaI
HincII
SpeI
308
Hydrogenase gene cluster genes from Synechococcus sp. PCC 7002
A12. Restriction map of the hydrogenase gene cluster from Synechococcus sp. PCC 7002.
1K 2K 3K 4K 5K 6K 7K 8K 9K 10K 11K 12K 13K 14K
hypFUreC hoxF hoxH hypDhypE hypBhoxU hyp3hoxYhoxE hypA hypC
hoxWXbaI DraI
SmaIPvuI
AvaIDraI
NcoI
StyI
PvuI
StyI
HincII
SspI
StyI
DraI
NcoI
StyI PvuI
StyIPvuI
PvuINcoI
StyI
PvuI PvuI
BglII
DraI
PvuI
NcoI
StyI
PvuI
DraI
KpnI
SspIPvuI
HindIII
PvuI
StyI
PvuI
HincII
SspI
SspI
HindIII
StyI
PvuI
StyI
NcoI
StyI
NcoI
StyI
DraI
SspISspI
PvuI PvuI
BglII
NcoI
StyI
PvuI
StyI
EcoRV
SspI
PvuI
EcoRV
DraI
NcoI
StyI
PvuI StyI
HindIII
EcoRI
PvuI
StyI
HincII
PvuI
PvuI
NcoI
StyI
SspI
DraI
DraI
PvuI
AvaI
SmaI
HincII
EcoRV
StyI
SspI
PvuI
DraI
EcoRI
BglII
BamHI
PvuI
DraI
HincII
PvuI
PvuI
DraI
PvuI
HindIII
StyI
DraI
BglII
SspI
HindIII
SspI
HindIII
PvuI
SspI
DraI
StyI
DraI
PvuI
HincII
PvuI
StyI
EcoRI
EcoRI
XbaI
EcoRV
PvuI
HincII
EcoRV
SspI
EcoRV
StyI
309
Mobile electron transport genes from genes from Synechococcus sp. PCC 7002
A13. Restriction map of petJ1 gene from Synechococcus sp. PCC 7002
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300
sll0185 homolog sll0415 ABC transporterpetJ1AvaI
PvuI
PvuI
PvuISacI
HindIIIEcoRV
BamHIStyI
KpnISspI
StyIPvuI
PvuI
HindIII
KpnI StyI
SspI
XbaI
PvuI
AvaI
HincII
310
A14. Restriction map of the petJ2 gene from Synechococcus sp. PCC 7002
300 600 900 1200 1500 1800 2100 2400 2700 3000
algI (alginate-o-acetyltransferase)slr0436 ccmK-1 petJ2sl l0102
PvuI
BamHI
HincII
BglII
BglII
NcoI
StyI
PvuI
SspIStyI
PstI SacI
XbaI
SmaI
AvaI
StyI
StyI
PvuI
PvuI
HindIII SspI
DraI
311
A15. Restriction map of the cytM gene from Synechococcus sp. PCC 7002
300 600 900 1200 1500 1800 2100 2400 2700
sll1704 csGA (cell-cell signaling factor C)
s l r1215
sll1434-penicillin binding pr… cyt M
NcoI
StyI
PvuI
StyIPvuI
PvuI
PvuIDraI
PvuI
BglII
XbaI
StyI
PvuI
PvuI
StyI
PvuI
StyI PvuI
StyI
StyI
312
A16. Restriction map of the bcpA gene from Synechococcus sp. PCC 7002
800 1600 2400 3200 4000 4800 5600 6400 7200 8000 8800 9600 10400 11200 12000 12800 13600
DNA polymerase I-slr0707 hypothetical-sll1858trigger factor-sll0533
bioF gene-slr0917
mfd gene-sll0377ilvC gene-sll1363
hypothetical protein
hypothetical proteinpepM-slr0918
blue-copper protein
hypothetical-slr0204glnB gene SspI
DraI
PvuI
StyI
StuI
PvuI
PvuI
HincII
XbaI
BamHI
HincII
HincII
EcoRI
StyI
StyI
HincII
PvuI
StyI
DraI
NcoI
StyI
StyI
KpnI
NcoI
StyI
StyI
PvuI
NcoI
StyI
StyI
PvuI
DraINcoI
StyI
BamHI
HincII
PvuI
EcoRI
EcoRV
HincII
PstI
BglII
EcoRV
PvuI
StuI
HincII
StyI
SspI
SspI
HincII
PvuI
BamHI
PvuI
PvuI
XbaI
XbaI
DraI
DraI
StuI
HincII
HincII
DraI
HindIII
PvuI
SspI
XbaI
StyI
PvuI
KpnI
AvaI
SmaI
EcoRV
PvuI
BamHIStyI
PvuI
DraI
SspI
SspI
SspI
StyI
XbaI
PvuI
NcoI
StyI
StyI
HincII
AvaI
SmaI
SspI
StyI
PvuI
DraI
HincII
PvuI
XbaI
PvuINcoI
StyI
HindIII
EcoRI
DraI
KpnI
EcoRI
SspI
StuI PvuI
SmaI
AvaI
313
Heme-copper oxidase genes from Synechococcus sp. PCC 7002
A17. Restriction map of the ctaCIDIEI operon from Synechococcus sp. PCC 7002
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600 3900 4200 4500 4800
ctaDIctaCI ctaEIsll1898 slr1542EcoRI
SacI
AvaI
KpnI
SphI
SmaI
BamHI
PstI
HincII
XbaI
BamHI
NcoI
StyI
StyI
PvuI
SspI
NcoI
StyI
PvuI
DraIDraI PvuI
HincII
SspI
NcoI
StyI
PvuI
StyI
PvuI
StyI
NcoI
StyI
NcoI
StyI
HincII
NcoI
StyI
PvuI
NcoI
StyI
HincII
StuI PvuI EcoRVBglII
PvuI
314
A18. Restriction map of the ctaCIIDIIEII operon from Synechococcus sp. PCC 7002
300 600 900 1200 1500 1800 2100 2400 2700 3000 3300 3600 3900 4200 4500 4800 5100
ctaDIIctaCIIsll1486 ctaEIIsll1485 sldI
AvaI
SmaI
BamHI
XbaI
HincII
PvuI
EcoRI DraI
PvuI
PvuI
PvuI
DraI PvuI PvuI
PvuI
StyI
PvuI HincII
PvuI PvuI
EcoRV
SspI
NdeI
NcoI
StyI
NdeI
PvuI
NcoI
StyI
SspIPvuI
315
APPENDIX B: CLUSTALW ALIGNMENTS OF ELECTRON TRANSPORTPROTEINS
The ClustalW amino acid alignments for the Synechococcus sp. PCC 7002
electron transport proteins are presented in this appendix. For details, please see
RESULTS.
Figure B1. ClustalW alignment of NdhA amino acid sequences. NdhA amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
316
NdhA ClustalW Amino Acid Alignment
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
1 0 2 0 3 0 4 0 5 0M N S G I D L Q G S F I E T L Q S F G L S H E I A K T I W L P L P L L L M I I G A T V G V L V V VM T S G I D L Q N S F L Q S L Q G F G L P P G L A K L F W I P L P S I L M I I G A T V G V L V V VM N S G I D L Q G T F I K S L I D L G I P P G T A K A I W M P L P M I L M L I G A T V G V L V C V
M I I D T T T T K V Q A I N S F S R L E F L K E V Y E T I W M L F P I L I L V L G I T I G V L V I VM S W I S P E L I E I L L T I L K A V V I L L V V V T C G A F M S
M S G I D L Q . . F I S L G . P E . A K . I W P L P . . L M . I G A T V G V L V . V
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
6 0 7 0 8 0 9 0 100W L E R K I S A A A Q Q R V G P E Y A G P L G V L Q P V A D G L K L V F K E D V V P A K T D P W L FW L E R K I S A A A Q Q R I G P E Y A G P L G V L Q P V A D G I K L V F K E D V V P A K A D P W L FW L E R K I S A A A Q Q R I G P E Y I G P L G L L A P V A D G L K L V F K E D I V P A Q A D P W L FW L E R E I S A S I Q Q R I G P E Y A G P L G I L Q A L A D G T K L L F K E N L L P S R G D T Y L FF G E R R L L G L F Q N R Y G P N R V G W G G S L Q L V A D M I K M F F K E D W I P K F S D R V I FW L E R K I S A A A Q Q R I G P E Y A G P L G . L Q P V A D G . K L V F K E D . V P A . . D P W L F
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
110 120 130 140 150T L G P A L V V I P V F L S Y L I V P F G Q N L V I T D L N V G I F L W I S L S S I A P I G L L M ST L G P V L V V L P V F L S Y L I V P F G Q N L V I T D I N V G I F L W I A L S S I A P I G L L M ST L G P I L V V L P V F L S Y L I V P F G Q N I V I T N V G T G I F L W I A L S S I Q P I G L L M AS I G P S I A V I S I L L G Y L I I P F G S R L V L A D L S I G V F L W I A V S S I A P I G L L M ST L A P M I A F T S L L L A F A I V P V S P G W V V A D L N I G I L F F L M M A G L A V Y A V L F AT L G P . L V V . P V F L S Y L I V P F G Q N L V I T D L N . G I F L W I A L S S I A P I G L L M S
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
160 170 180 190 200G Y S S N N K Y A L L G G L R A A A Q S I S Y E I P L A L A V L A I A M M S N S L S T I D I V E Q QG Y A S N N K Y S L L G G L R A A A Q S I S Y E I P L S L A V L A I V M M S N S L S T I D I V D Q QG Y S S N N K Y S L L G G L R A A A Q S I S Y E I P L A L S V L A I V M M S N S L S T V D I V N Q QG Y G S N N K Y S F L G G L R A A A Q S I S Y E I P L T L C V L S I S L L S N S S S T V D I V E A QG W S S N N K Y S L L G A M R A S A Q T L S Y E V F L G L S L M G V V A Q A G S F N M T D I V N S QG Y S S N N K Y S L L G G L R A A A Q S I S Y E I P L . L V L A I V M M S N S L S T . D I V . Q Q
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
210 220 230 240 250S G Y G I L G W N I W R Q P V G F L I F W I A A L A E C E R L P F D L P E A E E E L V A G Y Q T E YS G Y G I L G W N I W R Q P V G F L I F W I A A L A E C E R L P F D L P E A E E E L V A G Y Q T E YS G Y G I L G W N I W R Q P L G F M I F W I A A L A E C E R L P F D L P E A E E E L V A G Y Q T E YS K Y G F W G W N L W R Q P I G F I V F I I S S L A E C E R L P F D L P E A E E E L V A G Y Q T E YA - - - - H V W N V I P Q F F G F I T F A I A G V A V C H R H P F D Q P E A E Q E L A D G Y H I E YS G Y G I L G W N I W R Q P . G F . I F W I A A L A E C E R L P F D L P E A E E E L V A G Y Q T E Y
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
260 270 280 290 300A G M K F G L F Y V G S Y V N L V L S A L I V S I L Y L G G W E F P I P L D K L A G W L N V A P S TA G M K F A L F Y L G S Y V N L V L S A L V F S V L Y L G G W D F P I P L E N V A N W L G V A P T TS G M K F A L F Y L S S Y V N L I L S A L L V A V L Y L G G W D F P I P I N V L A N L V G V S E A NS G I K F G L F Y V A S Y L N L L I S S L F V T V L Y L G G W N L S I P Y I F I S - - - E F F E I NS G M K F G L F F V G E Y I G I V T I S A L M V T L F F G G W Q G P L L P P F I W - - - - - - - - -S G M K F G L F Y V G S Y V N L V L S A L . V . V L Y L G G W . F P I P . . A . V
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
310 320 330 340 350P W L Q V I T A S L G I I M T L V K T Y A L V F I A V L L R W T L P R V R I D Q L L N F G W K F L LS W L Q V L M A A L G I T M T V L K S Y F L I F I A I L L R W T V P R V R I D Q L L N L G W K F L LP V L Q V V S A A L G I T M T L V K A Y F L V F I A I L L R W T V P R V R I D Q L L D L G W K F C YK I D G V F G T T I G I F I T L A K T F L F L F I P I T T R W T L P R L R M D Q L L N L G W K F L L- - - - - - - - - - - - - - F A L K T A F F M M M F I L I R A S L P R P R Y D Q V M S F G W K I C L
L Q V . A L G I M T L . K T Y F L . F I A I L L R W T L P R V R I D Q L L N L G W K F L L
NdhA-7002NdhA-6803NdhA-7120NdhA-Spinacia oleraceaNuoH-E. coli
360 370 380 390 400P V A L V N L L L T A A L K L A F P I A F G GP V A L A N L L I T A A L K L T F P M A F G GQ L V S E P T F N R Q P L K L A F P R P S A GP I S L G N L L L T T S S Q L F S LP L T L I N L L V T A A V I L W Q A QP . . L . N L L . T A A L K L F P . G
317
Figure B2. ClustalW alignment of NdhI amino acid sequences. NdhI amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
318
NdhI ClustalW Amino Acid Alignment
NdhI-7002NdhI-6803NdhI-7120NdhI-Spinacia oleraceaNuoI-E. coli
1 0 2 0 3 0 4 0 5 0M F K I L K Q V G D Y A K D A A Q A A K - - - - - - - - - - - Y I G Q G L S V
M F N N I L K Q V G D Y A K E S L Q A A K - - - - - - - - - - - Y I G Q G L A VM L K F L K Q V G D Y A K E A V Q A G R - - - - - - - - - - - Y I G Q G L S VM F P M V T G F I N Y G Q Q T I R A A R - - - - - - - - - - - Y I G Q S F M I
P G N V L R L E N L P A A D A D Q L A G N G G C H S L A G A I R G N K T M T L K E L L V G F G T Q VM . L K Q V G D Y A K . . . Q A A . Y I G Q G L V
NdhI-7002NdhI-6803NdhI-7120NdhI-Spinacia oleraceaNuoI-E. coli
6 0 7 0 8 0 9 0 100- - - - - T F D H M R R R P V T V Q Y P Y E K L I P S E R F R G R I H F E F D K - - - - - C I A C E- - - - - T F D H M S R R P I T V Q Y P Y E K L I P S E R F R G R I H F E F D K - - - - - C I A C E- - - - - T F D H M R R R P V T V Q Y P Y E K L I P G E R F R G R I H Y E F D K - - - - - C I A C E- - - - - T L S H A N R L P V T I Q Y P Y E K L I T S E R F R G R I H F E F D K - - - - - C I A C ER S I W M I G L H A F A K R E T R M Y P E E P V Y L P P R Y R G R I V L T R D P D G E E R C V A C N
T F D H M R R P V T V Q Y P Y E K L I P S E R F R G R I H F E F D K C I A C E
NdhI-7002NdhI-6803NdhI-7120NdhI-Spinacia oleraceaNuoI-E. coli
110 120 130 140 150V C V R V C P I N L P V V D W E F D K S I K K K T L K H Y S I D F G V C I F C G N C V E Y C P T N CV C V R V C P I N L P V V D W E F N K A V K K K E L K H Y S I D F G V C I F C G N C V E Y C P T N CV C V R V C P I N L P V V D W E F D K A T K K K K L N H Y S I D F G V C I F C G N C V E Y C P T N CV C V R A C P I D L P V V D W K L E T D I R K K R L L N Y S I D F G I C I F C G N C V E Y C P T N CL C A V A C P V G C I S L Q K A E T K D G R - W Y P E F F R I N F S R C I F C G L C E E A C P T T AV C V R V C P I N L P V V D W E F . K . K K K L H Y S I D F G V C I F C G N C V E Y C P T N C
NdhI-7002NdhI-6803NdhI-7120NdhI-Spinacia oleraceaNuoI-E. coli
160 170 180 190 200L S M T E E Y E L A T Y D R H E L N Y D N - V A L G R L P Y K V T Q D P M V T P L R E L A Y L P Q GL S M T E E Y E L A A Y D R H D L N Y D N - V A L G R L P Y K V T E D P M V T P L R E L G Y L P K GL S M T E E Y E L A T Y D R H E L N Y D S - V A L G R L P Y K V T D D P M V T P L R E L V Y L P K GL S M T E E Y E L S T Y D R H E L N Y N Q - I A L G R L P I S I T D D - - - - - - - - - - Y T I R TI Q L T P D F E M G E Y K R Q D L V Y E K E D L L I S G P G K Y P E Y N - - - - - - - F Y R M A G ML S M T E E Y E L A T Y D R H E L N Y D . V A L G R L P Y K V T . D P M V T P L R E L . Y L P . G
NdhI-7002NdhI-6803NdhI-7120NdhI-Spinacia oleraceaNuoI-E. coli
210 220 230 240 250V I D P H D L P Q G S Q R A G E H P E D I L E R L Q A E K Q Q E T A D QV I E P H N L P K G S Q R A G Q H P E D L V K A EV L D P H D L P A N A P R P G A R P E D L V E K T E AI L N - - - S P Q T K E K A C DA I D - - G K D K G E A E N E A K P I D V K S L L PV I D P H L P G R A G . P E D . .
319
Figure B3. ClustalW alignment of NdhG amino acid sequences. NdhG amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
320
NdhG ClustalW Amino Acid Alignment
NdhG-7002NdhG-6803NdhG-7120NdhG-Spinacia oleraceaNuoJ-E. coli
10 20 30 40 50V N L A E G V Q I V S F V I L T V M M L G S A L G V V L F E N I V Y S A F L L G G V F I S I S G F YM N L A E G V Q Y I S F L I L A F L V I G A A L G V V L L S N I V Y S A F L L G G V F L S I S G I YM N L A E G V Q V V P F G I L A T M L I G P A L G V V L A T S I V Y S A F L L G G V F I S I A G M YM D L P G P I H D F L L V F L G S G L I L G A L G V V L F T N P I F S A F S L G L V L V C I S L F Y
M E F A F Y I C G L I A I L A T L R V I T H T N P V H A L L Y L I I S L L A I S G V FM N L A E G V Q . F . I L . . . I G . A L G V V L T N I V Y S A F L L G G V F . S I S G Y
NdhG-7002NdhG-6803NdhG-7120NdhG-Spinacia oleraceaNuoJ-E. coli
60 70 80 90 100I L L N A D F V A A A Q V L I Y V G A I N V L I L F A I M L V N K K E D F S Q M P G R L L R Q G A TI L L N A D F V A A A Q V L V Y V G A V S V L I L F A I M L V N K R E D F S K I P G R WL R N V S TL L L N G D F V A A A Q V L V Y V G A V N V L I L F A I M L V N K R Q D F T P Y P S A G I R K V L TI L A N S H F V A S A Q L L I Y V G A I N V L I I F S V M F M S G P E Y D K K F Q L WT V G D G V TF S L G A Y F A G A L E I I V Y A G A I M V L F V F V V M M L N - - - - L G G S E I E Q E R Q WL KI L L N A D F V A A A Q V L V Y V G A I N V L I L F A I M L V N K . E D F . P . . R . . . T
NdhG-7002NdhG-6803NdhG-7120NdhG-Spinacia oleraceaNuoJ-E. coli
110 120 130 140 150A L V C L G L F A L L G T M V L V T P WE - - - - - - - L S T I S P A A V G N S V V A I A K H F F SA L V C T G I F A L L S T M V L I T P WQ - - - - - - - I N E T G P F V E N - T L V T I G K H F F SA I V S V G L F A L L S T M V L A T P WA - - - - - - - Y S T T P K V G D G - S I I V I G E H F F SS L V C I S L F V S L I S T I L N T S WY G I I WT T K S N Q I L E Q D L I N A S Q Q I G I H L S TP Q V WI G P A I L S A I M L V V I V Y A I L G - - - - V N D Q G I D G T P I S A K A V G I T L F GA L V C . G L F A L L . T M V L . T P W . N . . . S . . . I G H F F S
NdhG-7002NdhG-6803NdhG-7120NdhG-Spinacia oleraceaNuoJ-E. coli
160 170 180 190 200D F L L P F E L A S V L L L I A M V G A I I L A R R D I I P E I T T T E E G S T G L Q L P E R P R ED Y L L P F E L A S V L L L M A M V G A I I L A R R D L I P E L S E E N K T A T A L T L P E R P R ED F L L P F E L A S V L L L M A M V G A I I L A R R E Y L P E V T P I WL T S T V L T L P E R P R ED F F L P F E L I S I I L L V S L I G A I A V A R QP Y V L A V E L A S M L L L A G L V V A F H V G R E E R A G E V L S N R K D D S A K R K T E E H AD F L L P F E L A S V L L L . A M V G A I I L A R R . . P E . . T . L L P E R P R E
NdhG-7002NdhG-6803NdhG-7120NdhG-Spinacia oleraceaNuoJ-E. coli
210 220 230 240 250L A G A N S P K S K Q NL T S A S KL V G A G S E T Q E
L A
321
Figure B4. ClustalW alignment of NdhE amino acid sequences. NdhE amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
NdhE ClustalW Amino Acid Alignment
NdhE-7002NdhE-6803NdhE-7120NdhE-Spinacia oleraceaNuoK-E. coli
10 20 30 40 50M Q I Q L E Y I L V L A A A L F C I G I Y G L V T S R N A V R V L M S I E L LM H L Q L Q Y C L I L A A A L F C I G I Y G L I T S R N A V R V L M S I E L L
M Q L R Y F L L L A A A L F C I G I Y G L I T S R N A V R V L M S I E L LM I L E H V L V L S A F L F S I G I Y G L V T S R N L V R A L M C L E L I
R R Q R E K K N G G A R M I P L Q H G L I L A A I L F V L G L T G L V I R R N L L F M L I G L E I M. Q L Y . L . L A A A L F C I G I Y G L V T S R N A V R V L M S I E L L
NdhE-7002NdhE-6803NdhE-7120NdhE-Spinacia oleraceaNuoK-E. coli
60 70 80 90 100L N S V N L N F M G F S N F L D P G E I K G Q V F T V F V L T V A A A E A A V G L A I I L A I Y R NL N A V N L N L M G F S N Y L D P S N I R G Q V F A I F V I T I A A A E A A V G L A I V L A I Y R NL N A V N L N L M A F S N Y L D S T L I K G Q V F T V F V I T V A A A E A A V G L A I V L A I Y R NL N A V N L N F V T F S D F F D S R Q L K G N I F S I F V I A I A A A E A A I G P A I V S S I Y R NI N A S A L A F V V A G S Y WG - - Q T D G Q V M Y I L A I S L A A A E A S I G L A L L L Q L H R RL N A V N L N F M . F S N Y L D . I K G Q V F . I F V I T . A A A E A A V G L A I V L A I Y R N
NdhE-7002NdhE-6803NdhE-7120NdhE-Spinacia oleraceaNuoK-E. coli
110 120 130 140 150R N T I D M E Q F N L L K NR E T T D M E Q F N L L K WR D T V D M E Q F N L L K WR K S I R I N Q S N L L N KR Q N L N I D S V S E M R GR T . D M E Q F N L L K
322
Figure B4. ClustalW alignment of NdhB amino acid sequences. NdhB amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 7942, Anabaena sp. PCC Spinacia oleracea, and E. coli.
The consensus amino acid sequence is underneath. Gray high-lighted regions are
identical amino acids while lighter shading indicates a conserved amino acid. A dash is
equivalent to a gap in the sequence. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
323
NdhB ClustalW Amino Acid Alignment
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
10 20 30 40 50M D F S S T V A S Q L Y T G A I L P E S I V I L T L I V V L V G D LM D F S S N V A A Q L N A G T I L P E G I V I V T L L L V L I V D LM D F L T - L A G Q L N A G V I L P E G I V I V T L L T V L V T D LM D F A N - L A A Q L N A G T I L P E G I V I V T L M G V L I V D L
M I WH V Q N E N F I L D S T R I F M K A F H L L L F D G S L I F P E C I L I F G L I L L L M I D S
M D F . A . Q L . G I L P E I V I . T L . . V L . . D L
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
60 70 80 90 100I V G R A R S G WI P Y A A I A G L L G S V F A L Y L G WD N P H P V A F L G A F N S D N L S I L FI G G R K V A L A L P Y L A I A G L L V S V G L L V T S WS M A D P I G F I G A F N G D N L S I I FI L G R Q S L R L T P A L A I T G L S A A I A V L T L Q WN T S Q N L A F L G G F N G D N L S I V FI L G R T S S R WI G Y L A I A G L L A A I V A L Y F Q WD A T N P I S F T G A F I G D D L S I I FT S D Q K D I P WL Y F I S S T S L V M S I T A L L F R WR E E P M I S F S G N F Q T N N F N E I F
M D V T P L M R V D G F A M L YI . G R . . . A I G L . . . . L W . F G . F . . D N L S I . F
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
110 120 130 140 150R G I I V L S T A F T I M M S V R Y V E R S G T A L S E F I C I L L T A T L G G M F L S G A N E L VR A I I A L S T V V T I L M S V R Y V Q Q T G T S L A E F I A I L L T A T L G G M F L S A A N E L VR G I V L L S A A V T I L L S I R Y V E Q S G T S L G E F I T I L L T A S L G G M F L S G A N E L VR G I I A L S A V V T I L M S I R Y V E Q S G T A L A E F I A I L L T A T L G G M F V S G A S E L VQ F L I L L C S T L C I P L S V E Y I E C T E M A L T E F L L F I L T A T L G G M F L C G A N D L IT G L V L L A S L A T C T F A Y P WL E G Y N D N K D E F Y L L V L I A A L G G I L L A N A N H L AR G I I . L S . . . T I S . R Y V E . G T L E F I . I L L T A T L G G M F L S G A N E L V
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
160 170 180 190 200M I F V S L E M L S I S S Y L L T G Y M K R D P R S N E A A L K Y L L I G A A S S A I F L Y G V S LM V F I S L E M L S I S S Y L M T G Y M K R D P R S N E A A L K Y L L I G A S S S A I F L Y G L S LT I F V S L E T L S I S S Y L L T G Y M K R D P R S N E A A L K Y L L I G A A S S A I F L Y G V S LM I F I S L E T L S I S S Y L L T G Y T K R D P R S N E A A L K Y L L I G A S S T A V F L Y G V S LT I F V A P E C F S L C S Y L L S G Y T K K D V R S N E A T T K Y L L M G G A S S S I L V H G F S WS L F L G I E L I S L P L F G L V G Y A F R Q K R S L E A S I K Y T I L S A A A S S F L L F G M A L
I F . S L E L S I S S Y L L T G Y K R D P R S N E A A L K Y L L I G A A S S A I F L Y G . S L
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
210 220 230 240 250L Y G L S G G K T I L S E I A L G F T D P Q - G G Q S L A L A I A L V F A I A G I A F K I S A V P FL Y G L S G G E T Q L V L I A E K L V N A D T V G Q S L G L A I A L V F V I A G I A F K I S A V P FL Y G L A G G E T Q L P A I A E K L G E A Q - - - - P L A L L I S L I F V I A G I A F K I S A V P FL Y G L S G G Q T E L N A I A N G I I T A N - V G Q S L G A V I A L V F V I A G I G F K I S A A P FL Y G S S G G E I E L Q E I V N G L I N T Q M Y N - S P G I S I A L I F I T V G I G F K L S P A P SV Y A Q S G - - - D L S F V A L G K N L G D G M L N E P L L L A G F G L M I V G L G F K L S L V P FL Y G L S G G T L I A G . . . . . . S L . L . I A L . F . I A G I . F K I S A V P F
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
260 270 280 290 300H Q WT P D V Y E G S P T P V V A F L S V G S K A A G F A L A I R L L V T V F N P V S E E WH F I FH Q WT P D V Y E G S P T P V V A F L S V G S K A A G F A V A I R L L V T A F G G I T D E WH V I FH Q WT P D V Y E G S P T P I V A F L S V G S K A A G F A L A I R L L V T A Y P A L T E Q WH F V FH Q WT P D V Y E G A P T P V I A F L S V G S K A A G F A L A I R L L T T V F P F V A E E WK F V FH Q WT P D V Y E G S P T P V V A F L S V T S K V A A S A S A T R I F D I P F Y F S S N E WH L L LH L WT P D V Y Q G A P A P V S T F L A T A S K I A I F G V V M R L F L Y A P V G D S E A I R V V LH Q WT P D V Y E G S P T P V V A F L S V G S K A A G F A . A I R L L . T . F . . E E WH . F
324
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
310 320 330 340 350T A L A I L S M V L G N V V A L A Q T S M K R M L A Y S S I A Q A G F V M I G L V A G - - T D A G YT A L A V L S M V L G N V V A L A Q T S M K R M L A Y S S I G Q A G F V M I G L V A G - - S E D G YT A L A I L S L V L G N V V A L A Q T S M K R L L A Y S S I A Q A G F V M I G L I A G - - T E A G YT A L A V L S M I L G N V V A L A Q T S M K R M L A Y S S I A Q A G F V M I G L I A G - - T D A G YE I L A I L S M I L G N L I A I T Q T S M K R M L A Y S S I G Q I G Y V I I G I I V G D - S N D G YA I I A F A S I I F G N L M A L S Q T N I K R L L G Y S S I S H L G Y L L V A L I A L Q T G E M S MT A L A . L S M . L G N V V A L A Q T S M K R M L A Y S S I . Q A G F V M I G L I A G . . G Y
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
360 370 380 390 400S S V I F Y L L V Y L F M N L G A F T C V I L F S L R T - G - - T D Q I A E Y A G L Y Q K D P L L TA S M V F Y M L I Y L F M N L G A F S C I I L F T L R T - G - - S D Q I S D Y A G L Y H K D P L L TS S M V Y Y L L I Y L F M N L G G F A C V I L F S L R T - G - - T D Q I S E Y A G L Y Q K D P L V TA S M I F Y L L V Y L F M N L C G F T C I I L F S L R T - G - - T D Q I A E Y S G L Y Q K D P L L TA S M I T Y M L F Y I S M N L G T F A C I V L F G L R T - G - - T D N I R D Y A G L Y T K D P F L AE A V G V Y L A G Y L F S S L G A F G V V S L M S S P Y R G P D A D S L F S Y R G L F WH R P I L A
S M . . Y L L . Y L F M N L G . F C . I L F S L R T G T D Q I . Y A G L Y K D P L L T
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
410 420 430 440 450L G L S V C L L S L G G I P P L A G F F G K I Y L F WA G WQ A G L Y G L V L L G L I T S V I S I YL G L S I C L L S L G G I P P L A G F F G K I Y I F WA G WQ S G L Y G L V L L G L V T S V V S I YL G L S L C L L S L G G I P P L A G F F G K L Y L F WA G WQ A G L Y G L V L L A L I T S V I S I YL G L S I S L L S L G G I P P L A G F F G K I Y L F WA G WQ A G L Y WL V L L G L V T S V I S I YL S L A L C L L S L G G L P P L A G F F G K I Y L F WC G WQ A G L Y F L V L I G L L T S V L S I YA V M T V M M L S L A G I P M T L G F I G K F Y V L A V G V Q A H L WWL V G A V V V G S A I G L YL G L S . C L L S L G G I P P L A G F F G K I Y L F WA G WQ A G L Y L V L L G L . T S V I S I Y
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
460 470 480 490 500Y Y I R V I K M M V V K E P Q E M S D S V R N Y P A V T WT A V G M K P L Q V G L V L S V I I T S LY Y I R V V K M M V V K E P Q E M S E V I K N Y P A I K WN L P G M R P L Q V G I V A T L V A T S LY Y I R V I K M M V V K E P Q E M S E S V R N Y P E T N WN L P G M Q P L R A G L V V C V I A T A VY Y I R V V K M M V V K E P Q E M S D V V K N Y P E I R WN L P G F R P L Q V G L V L T L I A T S VY Y L K I I K L L M T G R N Q E I T P H V R N Y R R S - - P L R S K N S I E F S M I V C V I A S T IY Y L R V A V S L Y L H A P E - - - Q P G R D A P S N - WQ Y S A G G - - - I V V L I S A L L V L VY Y I R V . K M M V V K E P Q E M S . V R N Y P W. L G P L . G . V . . . I A T . .
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
510 520 530 540 550A G I L S N P L F V I A D Q S V T S - - - T P M L Q V A N H P T E Q V V A Q V D S E L V G V A I A DA G I L A N P L F N L A T D S V V S - - - T K M L Q T A L Q Q T G E T P A I A I S H D L PA G I L S N P L F N L A S A S V S G S S F L G L A P A A E V V T T T A T P V A L S E P P A A SA G I L S N P L F T L A N N S V A N - - - T A I L Q A T K V V S T Q V S A I P A E K P D G L NP G I S M N P I I T I A Q D T L FL G V WP Q P L I S I V R L A M P L MA G I L N P L F . A S V . .
NdhB-7002NdhB-6803NdhB-7942NdhB-7120NdhB-Spinacia oleraceaNuoN-E. coli
560 570 580 590 600H
325
Figure B5. ClustalW alignment of NdhD1 amino acid sequences. NdhD1 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
326
NdhD1 ClustalW Amino Acid Alignment
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
10 20 30 40 50M N F A N F P WL S T I I L F P I I A A L F L P L I P D K D G K T V R WY A L T I G L I D F V I I VM N T A N F P WL T T I I L L P I A A S L L I P I I P D K D G K T I R WY A L T V G L I D F A L I V
M N T F P WL T T I I L L P I V A A L F I P I I P D K D G K T V R WY S L A V G L V D F A L I VM I E L L L M T
M L L P WL - - - I L I P F I G G F L C WQ T E R F G V K V P R WI A L I T M G L T L A L S LF P WL . T I I L . P I . A . L . P . I P D K D G K T . R WY A L . G L I D F A L I V
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
60 70 80 90 100T A F Y T G - Y D F G N - - - - P N L Q L V E S Y T WV E A I D L R WS V G A D G L S M P L I L L TY A F Y T S - Y D F A N - - - - P D L Q L V E S Y P WV P Q L D L N WS V G A D G L S M P L I I L TY A F Y S G - F D L S E - - - - P G L Q L V E S Y T WL P Q I D L K WS V G A D G L S M P L I I L TY V F F Y H - F Q P D D - - - - P L I Q L V E D Y K WI N F F D F H WR L G I D G L S I G P I L L TQ L WL Q G G Y S L T Q S A G I P Q WQ S E F D M P WI P R F G I S I H L A I D G L S L L M V V L TY A F Y . G Y D . P L Q L V E S Y W. P . D L . WS V G A D G L S M P L I . L T
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
110 120 130 140 150G F I T T L A I L A A WP V S F K P K - L F Y F L M L L M Y G G Q I A V F A V Q D M L L F F F T WEG F I T T L A T L A A WP V T L K P R - L F Y F L L L A M Y G G Q I A V F A V Q D M L L F F L V WEG F I T T L A T M A A WP V T L K P K - L F Y F L M L L M Y G G Q I A V F A V Q D I L L F F L V WEG F I T T L A T L A A WP V T R N S Q - L F H F L M L A M Y S A Q I G L F S S R D L L L F F I M WEG L L G V L A V L C S WK E I E K Y Q G F F H L N L M WI L G G V I G V F L A I D M F L F F F F WEG F I T T L A T L A A WP V T K P . L F Y F L M L . M Y G G Q I A V F A V Q D M L L F F . WE
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
160 170 180 190 200L E L V P V Y L I L S I WG - - - - - G K K R L Y A A T K F I L Y T A G G S L F I L I A A L T M A FL E L I P V Y L L L A I WG - - - - - G K K R Q Y A A T K F I L Y T A G G S L F I L L A S L T M A FL E L V P V Y L I L S I WG - - - - - G K K R L Y A A T K F I L Y T A G G S L F I L L A G L T L A FL E L I P V Y L L L S M WG - - - - - G K K R L Y S A T K F I L Y T A G G S I F L L M G V L G V G LM M L V P M Y F L I A L WG H K A S D G K T R I T A A T K F F I Y T Q A S G L V M L I A I L A L V FL E L V P V Y L L L S I WG G K K R L Y A A T K F I L Y T A G G S L F I L . A . L T . A F
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
210 220 230 240 250Y G - - - - D T V T F D M T A I A Q K D F G I N L Q L L L Y G G L L I A Y G V K L P I F P L H T WLY G - - - - D T V T F D M R S L A L K D Y A L N F Q L L L Y A G F L I A Y A I K L P I I P L H T WLY G - - - - D V N T F D M S A I A A K D I P V N L Q L L L Y A G F L I A Y G V K L P I F P L H T WLY G S - - - N E P T L N L E T L V N Q S Y P V A L E I I F Y I G F F I A F A V K L P I I P L H T WLV H Y N A T G V WT F N Y E E L L N T P M S S G V E Y L L M L G F F I A F A V K M P V V P L H G WLY G D T F D M L A . K D . . N L Q L L L Y . G F L I A Y A V K L P I . P L H T WL
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
260 270 280 290 300P D A H G E A T A P A H M L L A G I L L K M G G Y A L L R M N A G M L P D A H A L F G P V L V I L GP D A H G E A T A P A H M L L A G I L L K M G G Y A L I R M N A G I L P D A H A Y F A P V L V V L GP D A H G E A T A P A H M L L A G I L L K M G G Y A L L R M N V G M L P D A H A V F A P V L V I L GP D T H G E A H Y S T C M L L A G I L L K M G A Y G L V R I N M E L L P H A H S I F S P WL M I I GP D A H S Q A P T A G S V D L A G I L L K T A A Y G L L R F S L P L F P N A S A E F A P I A M WL GP D A H G E A T A P A H M L L A G I L L K M G G Y A L L R M N . G . L P D A H A . F A P V L V I L G
327
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
310 320 330 340 350V V N I V Y A A L T S F A Q R N L K R K I A Y S S I S H M G F V L I G M A S F T D L G T S G A M L QV V N I I Y A A L T S F A Q R N L K R K I A Y S S I S H M G F V I I G F A S F T D L G L S G A V L QV V N I I Y A A F T S F A Q R N L K R K I A Y S S I S H M G F V L I G L A S F T D L G M S G A M L QT M Q I I Y A A S T S P G Q R N L K K R I A Y S S V S H M G F I I I G I S S I T D T G L N G A I L QV I G I F Y G A WM A F A Q T D I K R L I A Y T S V S H M G F V L I A I Y T G S Q L A Y Q G A V I QV V N I I Y A A T S F A Q R N L K R K I A Y S S I S H M G F V L I G . A S F T D L G S G A . L Q
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
360 370 380 390 400M I S H G L I G A S L F F M V G A T Y D R T H T L M L D E M G G V G K K M K K I F A M WT T C S M AM V S H G L I G A S L F F L V G A T Y D R T H T L M L D E M G G V G K R M P K I F A M F T A C S M AM I S H G L I G A S L F F M V G A T Y D R T H T L M L D E M G G I G Q K M K K G F A M WT A C S L AI I S H G F I G A A L F F L A G T S Y D R I R L V Y L D E M G G I A I P M P K I F T L F S S F S M AM I A H G L S A A G L F I L C G Q L Y E R I H T R D M R M M G G L WS K M K WL P A L S L F F A V AM I S H G L I G A S L F F L V G A T Y D R T H T L M L D E M G G . G K M K K I F A M . T C S M A
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
410 420 430 440 450S L A L P G M S G F V A E L M V F V G F A T S D A Y S P T F R V I I V F L A A V G V I L T P I Y L LS L A L P G M S G F V A E L M V F V G F A T S D A Y S S T F K V I V V F L M A V G V I L T P I Y L LS L A L P G M S G F V A E L M V F V G F A T S D A Y N L V F R T I V V V L M G V G V I L T P I Y L LS L A L P G M S G F I A E L I V F F G L I T S Q K Y L L I P K L L I T F G M A I G M I L T P I Y L LT L G M P G T G N F V G E F M I L F G - - - - - - - S F Q V V P V I T V I S T F G L V F A S V Y S LS L A L P G M S G F V A E L M V F V G F A T S D A Y S F . . I I V F L M A V G V I L T P I Y L L
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
460 470 480 490 500S M L R E I L Y G P E N K E L V A H E K L I D A E P R E V F V I A C L L I P I I G I G L Y P K A V TS M L R E I F Y G K E N E E L V S H Q Q L I D A E P R E V F V I A C L L V P I I G I G F Y P K L L TS M L R E M L Y G P E N E E L V N H T N L V D V E P R E V F I I G C L L V P I I G I G F Y P K L I TS M S R Q M F Y G - Y K L F N I S N S S F F D S G P R E L F V S T S I F L P V I G I G V Y P D L V LA M L H R A Y F G - K A K S Q I A S Q E L P G M S L R D V F M I L L L V V L L V L L G F Y P Q P I LS M L R E . . Y G E N E L V H L . D . E P R E V F V I . C L L V P I I G I G F Y P K L . T
NdhD1-7002(slr0331)NdhD1-7120(slr0331)NdhD1-6803(slr0331)NdhD-Spinacia oleraceaNuoM-E. coli
510 520 530 540 550Q I Y A S T T E N L T A I L R Q S V P S L Q Q T A Q A P S L D V A V L R A P E I RQ M Y D A T T V Q L T A R L R D S V P T L A Q E K Q E - - V A R V S L S A P V I G NQ I Y D P T I N Q L V Q T A R R S V P S L V Q Q A N L S P L E V T A L R P P T I G FS L S V E K V E A I L S N Y F Y RD T S H S A I G N I Q Q WF V N S V T T T R PQ . Y T . . L . R S V P . L Q . . L P I
328
Figure B6. ClustalW alignment of NdhD2 amino acid sequences. NdhD2 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
329
NdhD2 ClustalW Amino Acid Alignment
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
10 20 30 40 50M D S L Q I P WL T T A I A F P L L A A L V I P L I P D K E G K T I R WY T L WR C P H R F C L L V
M L E H F P WL T T M I A L P L V A A L F I P L I P D K D G K Q V R WY A L G V G L A D F V L M SM A D G F P WL T A I I L L P L V A S A F I P L L P D K E G K L V R WY A L G V G I A D F V L M C
M I E L L L M TM L L P WL - - - I L I P F I G G F L C WQ T E R F G V K V P R WI A L I T - M G L T L A L S
P WL T I . . P L . A . . I P L . P D K . G K . R WY A L . . . F . L M .
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
60 70 80 90 100T A F WQ N - - Y D F G R - - - - T E F Q L T K N F A WI P Q L G L N WS L G V D G L S M P L I I LY V F WT N - - Y D I S S - - - - T G F Q L Q E K F S WI P Q F G L S WS V S V D G I S M P L V L LY T F WH H - - Y D T S S - - - - A T F Q L V E K Y D WL P Q I G F S WA V S V D G I S M P L V L LY V F F Y H - - F Q P D D - - - - P L I Q L V E D Y K WI N F F D F H WR L G I D G L S I G P I L LL Q L WL Q G G Y S L T Q S A G I P Q WQ S E F D M P WI P R F G I S I H L A I D G L S L L M V V LY . F W . Y D . F Q L E . WI P Q F G . S W L . V D G L S M P L V L L
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
110 120 130 140 150A T L I T T L A T L A A WN V T K K P K - L F A G L I L V M L S A Q I G V F A V Q D L L L F F I M WA G L V T T L S I F A A WQ V D H K P R - L F Y F L M L V L Y A A Q I G V F V A Q D M L L L F I M WA G F V T T L S M L A A WQ V N L K P R - L F Y F L M L V L Y S A Q I G V F V A Q D L L L F F I M WT G F I T T L A T L A A WP V T R N S Q - L F H F L M L A M Y S A Q I G L F S S R D L L L F F I M WT G L L G V L A V L C S WK E I E K Y Q G F F H L N L M WI L G G V I G V F L A I D M F L F F F F WA G L . T T L A L A A W. V . K P . L F F L M L V . Y S A Q I G V F . A Q D L L L F F I M W
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
160 170 180 190 200E L E L V P V Y L L I S I WG - - - - - G K K R L Y A A T K F I L Y T A L G S V F I L A F T L A L AE L E L V P V Y L L V C I WG - - - - - G Q K R Q Y A A M K F L L Y T A A A S V F I L V A A L G L AE L E L V P V Y L L V S I WG - - - - - G Q K R R Y A A T K F L L Y T A A A S I F I L I A G L A M AE L E L I P V Y L L L S M WG - - - - - G K K R L Y S A T K F I L Y T A G G S I F L L M G V L G V GE M M L V P M Y F L I A L WG H K A S D G K T R I T A A T K F F I Y T Q A S G L V M L I A I L A L VE L E L V P V Y L L . S I WG G K K R . Y A A T K F . L Y T A A . S . F I L . A . L A L A
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
210 220 230 240 250F Y G - G D V - - - T F D M Q A L G L K D Y P L A L E L L A Y A G F L I G F G V K L P I F P L H S WF Y G - D V T - - - T F D I A E L G L K D Y P I A L E L F L Y A G L L I A F G V K L A I F P F H T WL Y G - D N T - - - T F D I V E L G A K N Y P L A L E L L L Y A G L L I A F G V K L A I F P L H T WL Y G S N E P - - - T L N L E T L V N Q S Y P V A L E I I F Y I G F F I A F A V K L P I I P L H T WF V H Y N A T G V WT F N Y E E L L N T P M S S G V E Y L L M L G F F I A F A V K M P V V P L H G WF Y G T T F D . E L G . K Y P . A L E L L L Y A G F L I A F G V K L P I F P L H T W
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
260 270 280 290 300L P D A H S E A S A P V S M I L A G V L L K M G G Y G L I R L N M E M L P D A H I R F A P L L I V LL P D A H G E A S A P V S M I L A G V L L K M G G Y G L I R L N L G L L E D A H V Y F A P I L V I LL P D A H G E A S A P V S M I L A G V L L K M G G Y G L I R L N L E L L P D A H I Y F A P V L A T LL P D T H G E A H Y S T C M L L A G I L L K M G A Y G L V R I N M E L L P H A H S I F S P WL M I IL P D A H S Q A P T A G S V D L A G I L L K T A A Y G L L R F S L P L F P N A S A E F A P I A M WLL P D A H G E A S A P V S M I L A G V L L K M G G Y G L I R L N L E L L P D A H . F A P . L . . L
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
310 320 330 340 350G I V N I V Y G A L T A F G Q T N L K R R L A S S S I F P H G L S S L G L L S F T D L G M N G A V LG V V N I I Y G G F S S F A Q D N M K R R L A Y S S V S H M G F V L L G I A S F T D L G I S G A M LG V I N I I Y G G L N S F A Q T H M K R R L A Y S S V S H M G F V L L G I A S F T D V G V S G A M LG T M Q I I Y A A S T S P G Q R N L K K R I A Y S S V S H M G F I I I G I S S I T D T G L N G A I LG V I G I F Y G A WM A F A Q T D I K R L I A Y T S V S H M G F V L I A I Y T G S Q L A Y Q G A V IG V . N I I Y G A . S F A Q T N . K R R L A Y S S V S H M G F V L L G I . S F T D L G . . G A . L
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
360 370 380 390 400Q M L S H G F I A A A L F F L S G V T Y E R T H T L M M D E M S G I A R L M P K T F A M F T A A A MQ M L S H G L I A A V L F F L A G V T Y D R T H T L S L A Q M G N I G K V M P T V F A L F T M G A MQ M L S H G L I A A V L F F L A G V T Y D R T H T M A M D N L G G I G Q A M P K V F A L F T A G T MQ I I S H G F I G A A L F F L A G T S Y D R I R L V Y L D E M G G I A I P M P K I F T L F S S F S MQ M I A H G L S A A G L F I L C G Q L Y E R I H T R D M R M M G G L WS K M K WL P A L S L F F A VQ M L S H G L I A A . L F F L A G V T Y D R T H T . M D M G G I . . M P K . F A L F T . A M
330
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
A S L A L P G M S G F V S E L T V F L G L S N S D A Y S Y G F K P I A I F L T A V G V I L T P I T CA S L A L P G M S G F V S E L A V F V G V S S S D I Y S T P F K T V T V F L A A V G L V L T P I Y LA S L A L P G M S G F V S E L K V F I G V T T S D I Y S P T F C T V M V F L A A V G V I L T P I Y LA S L A L P G M S G F I A E L I V F F G L I T S Q K Y L L I P K L L I T F G M A I G M I L T P I Y LA T L G M P G T G N F V G E F M I L F G S - - - - - - - F Q V V P V I T V I S T F G L V F A S V Y SA S L A L P G M S G F V S E L V F . G . . . S D . Y S F K V . . F L A V G . I L T P I Y L
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
460 470 480 490 500F Q C C G V F Y G K G S Q A P P R C G G E - - - - - - - - - - - - - - - - - - - - - - - - - - - - DL S M L R Q L F Y - G N N I P P S C N L E Q D N L S A N S D Q E A V C F G T S C V L P G N A I Y D DL S M L R Q V F Y - G T G A E L S C N I N N G A Y Q N Q E D E G T A C F G T D C L L P G E A V Y R DL S M S R Q M F Y - G Y K L F N I S N S S - - - - - - - - - - - - - - - - - - - - - - - - - - F F DL A M L H R A Y F - G K A K S Q I A S Q E L P - - - - - - - - - - - - - - - - - - - - - - - - - - GL S M L R Q . F Y G . C N . E . D
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
510 520 530 540 550A K P R E I F V A V C L L A P I I A I G L Y P K L A T T T Y D L K T V E V A S K V R A A L P L Y A EA R P R E V F I A A C F L L P I I A V G L Y P K L A T Q T Y D A T T V A V N S Q V R Q S Y V Q I A EA S V R E V F I A V S F L V L I I G V G V Y P K I A T Q L Y D V K T V A V N T Q V R Q S Y T Q I A QS G P R E L F V S T S I F L P V I G I G V Y P D L V L S L S V E K V E A I L S N Y F Y RM S L R D V F M I L L L V V L L V L L G F Y P Q P I L D T S H S A I G N I Q Q WF V N S - - - V T TA P R E V F . A . . L . P I I . . G . Y P K L A T T Y D . K T V A V . S . V R . S . A
NdhD2-7002 (slr1291)NdhD2-6803(slr1291)NdhD2-7120(slr1291)NdhD-Spinacia oleraceaNuoM-E. coli
560 570 580 590 600Q L P - - - Q N G D R Q A Q M G L S S Q M P A L I A P R FT N P R V Y A E A L T A P H I P T T D F A T V K V Q PS N P Q I Y A K G F F T P Q I V E P E V M A V S G V I K
T R P. P . . .
331
Figure B7. ClustalW alignment of NdhD3 amino acid sequences. NdhD3 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b). Data for NdhD3 is taken from Klughammer et al.
(Klughammer et al., 1999).
332
NdhD3 ClustalW Amino Acid Alignment
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
10 20 30 40 50M L S F L L F L P L V G I G A I A L F P - - - - - R P L T R I V A T V F T V V T L A I S S G L L IM L S V L I W I P I L S A I V I G F WP S N P N Q S S R I R L V A L T V A A I V L I W N L F I L FM L S L L L I L P V I G A L I I G F F P G N I - P A K Q L R Q I T E V F A V L T L V W S L L V L F
MI E L L L M T Y V F F Y H FML L P WL I L I P F I G G F L C WQ T E R F G - - V K V P R WI A L I T MG L T L A L S L Q L WL
M L S L . . P . . G . . . I . . P R . A . . L T L . . S L . L F
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
60 70 80 90 100N - - L N L Q D - - - - A G M Q Y T E F H N WL S I L G L N Y N L G V D G L S L P L I V L N S L L TK - - F D I S N - - - - P G M Q F Q E Y L P WN E T L G L S Y Q L G V D G L S I L M L V L N S L L TK - - F D V T D - - - - P Q F Q F Q E Y L P WI P Q L G L N Y S L A I D G L S L P L V I L N N L L TQ - - P - - D D - - - - P L I Q L V E D Y K WI N F F D F H W R L G I D G L S I G P I L L T G F I TQ G G Y S L T Q S A G I P Q W Q S E F D M P WI P R F G I S I H L A I D G L S L L M V V L T G L L G. . . . D P . Q . E . P WI L G L Y L G I D G L S L . . V L N L L T
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
110 120 130 140 150L V A I Y S I G E S N H R P K - L Y Y S L I L L I N S G I T G A L I A N N L L L F F L F Y E I E L IWI A I Y S S S N K T E R P R - L F Y S M I L L V S G G V A G A F L S E N L L L F F L F Y E L E L IG V A I Y S I G P N V N R S R - L Y Y G L I L L I N A G I S G A L L A Q N L L L F I V F Y E L E L IT L A T L A A W P V T R N S Q - L F H F L M L A MY S A Q I G L F S S R D L L L F F I M WE L E L IV L A V L C S W K E I E K Y Q G F F H L N L MW I L G G V I G V F L A I D MF L F F F F WE MM L V. . A I Y S . R . L F Y L I L L I . G . . G A F L A N L L L F F . F Y E L E L I
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
160 170 180 190 200P F Y L L I A I WG - - - - - G E K K G Y A S T K F L I Y T A I S G L C V L A A F L G I V W L S Q SP F Y L L I S I WG - - - - - G E K R A Y A G I K F L I Y T A V S G A F I L A T F L G M V W L T G SP F Y L MI A I WG - - - - - G E K R G Y A S M K F L L Y T A F S G L L V L A A F L G M S L L S G SP V Y L L L S M WG - - - - - G K K R L Y S A T K F I L Y T A G G S I F L L M G V L G V G L Y G S NP M Y F L I A L WG H K A S D G K T R I T A A T K F F I Y T Q A S G L V M L I A I L A L V F V H Y NP F Y L L I A I WG G E K R . Y A . T K F L I Y T A . S G L . L A A F L G . V . L . S
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
210 220 230 240 250S N - - - - F D F E N L T L E N L E F N T K V I L L T I L L I G F G I K I P L V P L H T WL P D A YT S - - - - F A F D A I S T Q G L S A G M Q F I L L V G I I L G F G I K I P L V P F H T WL P D A YH S - - - - F D Y N P E I T Q T F T E S A Q T I L L I L I L L G F G I K I P L V P L H T WL P D A YE P - - - T L N L E T L V N Q S Y P V A L E I I F Y I G F F I A F A V K L P I I P L H T WL P D T HA T G V WT F N Y E E L L N T P M S S G V E Y L L M L G F F I A F A V K M P V V P L H G WL P D A H
. F . E L . Q . . . I L L . G . . I G F G I K I P L V P L H T WL P D A Y
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
260 270 280 290 300V E A N P A V T V L L G G V F A K L G T Y G L V R F G L Q L F P D V W S T V S P A L A V I G T V S VV E A S A P I A I L L G G V L A K L G T Y G L L R F G L G M F P Q T W S T V A P T L A I WG A V S AT E A S P A T A I L L G G I L A K L G T Y G I I R F G L Q L F P Q T W A Q F A P V L A I I G T V T VG E A H Y S T C ML L A G I L L K MG A Y G L V R I N M E L L P H A H S I F S P W L MI I G T M Q IS Q A P T A G S V D L A G I L L K T A A Y G L L R F S L P L F P N A S A E F A P I A MW L G V I G I. E A A . . L L G G I L A K L G T Y G L . R F G L L F P . . W S F A P . L A I I G T V . .
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
310 320 330 340 350MY G S L A A I A Q R D L K R MV A Y S S I G H MG Y I L V S T A A G T E L S L L G A V A Q MI S HI Y G A V I A I A Q K D I K R MV A Y S S I G H MG Y V L L A S A A S T S L A L V G A V S Q MF S HL Y G A L S A I A Q K D I K R MV A Y S S I G H MG Y I L V A A A A G T E L S V L G A V A Q MV S HI Y A A S T S P G Q R N L K K R I A Y S S V S H MG F I I I G I S S I T D T G L N G A I L Q I I S HF Y G A WM A F A Q T D I K R L I A Y T S V S H MG F V L I A I Y T G S Q L A Y Q G A V I Q MI A H. Y G A . A I A Q . D I K R MV A Y S S I G H MG Y I L . A . A A G T . L . L . G A V . Q MI S H
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
360 370 380 390 400S L I L A L L F H L V G I I E R K V G T R D L D V L N G L M N P V R G L P L T S S L L I L A G M A SG L I L A I L F H L V G I I E E K V G T R E L D K L N G L M S P I R G L P L I S A L L V L S G M A SG L I L A L L F H L V G I V E R K A G T R D L D V L N G L M N P I R G L P L T S A L L I T G G M A SG F I G A A L F F L A G T S Y D R I R L V Y L D E M G G I A I P M P K I F - - - T L F S S F S M A SG L S A A G L F I L C G Q L Y E R I H T R D MR MM G G L W S K M K W L P - - - A L S L F F A V A TG L I L A . L F H L V G I . E . K . G T R D L D L N G L M P . R G L P L S A L L . G M A S
333
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
410 420 430 440 450A G I P G L V G F V A E F L V F Q G - - - - - S F S R F P - - I P T L F C I I A S G L T A V Y F V IA G I P G L T G F I A E F I V F Q G - - - - - S F T A F P - - L S T L L C V A S S G L T A V Y F V IA G I P G L V G F A A E F I V F Q G - - - - - S F P T F P - - I P T L L C I L A S G L T A V Y F V IL A L P G M S G F I A E L I V F F G L I T S Q K Y L L I P K L L I T F G M A I G M I L T P I Y L L SL G MP G T G N F V G E F MI L F G - - - - - S F Q V V P - - V I T V I S T F G L V F A S V Y S L AA G I P G L . G F . A E F I V F Q G S F . F P . T L . C . . . S G L T A V Y F V I
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
460 470 480 490 500L L N R T C F G R L D S H T A Y Y P K V F A S E K I P A I A L - - T V I I L F L G L Q P A W L T R WL L N R T C F G R L D N N Q A Y Y A K V L W S E K T P A L I L - - A A L I I F L G V Q P T W L V R WL L N R T C F G K L D N Q R A Y Y P K V L A S E MI P A L V L - - T A I I F F L G V Q P N Y L V H WMS R Q MF Y G Y K L F N I S - - - N S S F F D S G P R E L F - - V S T S I F L P V I G - - I G V YML H R A Y F G K A K S Q - - - - - - - I A S Q E L P G MS L R D V F MI L L L V V L L V L L G F YL L N R T C F G . L D . A Y Y K V . A S E . P A . . L . . . I . F L G V Q P . L . . W
NdhD3-7002(sll1733)NdhD3-7120(sll1733)NdhD3-6803(sll1733)NdhD-Spinacia oleraceaNuoM-E. coli
510 520 530 540 550I E P T T S Q F I A A I P T V Q T I A L T P A E L S K A PS E P T T T A M V A A I P P V E K T V I S Q V V L KT Q T T T N E M I A Q L P H E T S A I E Q L A Q G V T L PP D L V L S L S V E K V E A I L S N Y F Y RP Q P I L D T S H S A I G N I Q Q WF V N S V T T T R P
. P T T . . A A I P . . . . . .
334
Figure B8. ClustalW alignment of NdhD4 amino acid sequences. NdhD4 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
335
NdhD4 ClustalW Amino Acid Alignment
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
10 20 30 40 50M L S A L I WL P L A G A L L V A I L P Q G E K N Q F S R T M A L G A A A L V F V WT A WL G F HM L S V L I F A P L L G A L L I G L L P S G I N G R S S R N V A L I F A S V T F L WS V I L A S QM L S A L I WG P I F G A I L I A I I P N P D H D C Y S R K I A L G I M V A M A G L S V L L A G Q
M I E L L L M T Y V F F Y HM L L P WL I L I P F I G G F L C WQ T E R - F G V K V P R WI A L I T M G L T L A L S L Q L WL Q
M L S . L I . . P . . G A . L . . . . P S R . A L . M . . . . S . . L . Q
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
60 70 80 90 100Y D V - - - - - - A I A G L Q F V E H Y L WI E WL G L N Y D L G V D G L S L P L L A L N A L L T LF Q P - - - - - - G E V N Q Q F S E F L P WI D T L G L S Y N L G V D G L S L P L L V L N G L L T GF N I - - - - - - S D P Q M Q F V E Y Y P WL P S L G L N Y H L G V D G L S L P L L L L N S A L V VF Q P - - - - - - D D P L I Q L V E D Y K WI N F F D F H WR L G I D G L S I G P I L L T G F I T TG G Y S L T Q S A G I P Q WQ S E F D M P WI P R F G I S I H L A I D G L S L L M V V L T G L L G VF . . . P . Q F V E Y P WI L G L Y . L G V D G L S L P L L . L N G L L T .
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
110 120 130 140 150V A L WI S P K D L H R P R - F Y Y A L F L L L Q A S V N G A F L A Q D V L L F F L F Y E I E I I PI A I Y S S D E S L Q R P K - F Y Y S L I L V L S A G V S G A F L A Q D L L L F F L F Y E L E L I PI A I F S T N T E I E R P R - F Y Y A L L L L L S G G V A G A F L A Q D L L L F F L F F E L E I I PL A T L A A WP V T R N S Q - L F H F L M L A M Y S A Q I G L F S S R D L L L F F I M WE L E L I PL A V L C S WK E I E K Y Q G F F H L N L M WI L G G V I G V F L A I D M F L F F F F WE M M L V P. A . . S . . R P . F Y Y . L . L . L . G V . G A F L A Q D L L L F F L F . E L E L I P
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
160 170 180 190 200L Y F L I A I WG - - - - - G K K R G Y A A I K F L L Y T A V S G I L I L A S F L G L A F L T E S NL Y L L I A I WG - - - - - G A R R G Y A A T K F L I Y T A F S G I L I L A S F L G L V WL S G S GL Y F L I A I WG - - - - - G Q R R G Y A A M K F L L Y T A L S G F L V L V S F L G WF WL T K A PV Y L L L S M WG - - - - - G K K R L Y S A T K F I L Y T A G G S I F L L M G V L G V G L Y G S N EM Y F L I A L WG H K A S D G K T R I T A A T K F F I Y T Q A S G L V M L I A I L A L V F V H Y N AL Y F L I A I WG G K . R G Y A A T K F L L Y T A . S G I L . L . S F L G L . . L .
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
210 220 230 240 250T - - - - F A Y S A L H S D L L P L T T Q L I L L G G I L V G F G I K I P F L P F H T WL P D A H VS - - - - F A L S T L N A Q S L P L A T Q L L L L A G I L V G F G I K M P L V P F H T WL P D A H VN - - - - F D Y N P S L A D A L P V K T Q M L L L L P L L L G L G I K I P I F P F H T WL P D A H VP - - - T L N L E T L V N Q S Y P V A L E I I F Y I G F F I A F A V K L P I I P L H T WL P D T H GT G V WT F N Y E E L L N T P M S S G V E Y L L M L G F F I A F A V K M P V V P L H G WL P D A H S. F Y L . L P . . T Q . L L L . G . L . G F G I K . P . . P F H T WL P D A H V
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
260 270 280 290 300E A S T P V S V I L A G V L L K V G T Y G L L K F G I G L F P L A WA V V A P WL A I WA A I S A LE A S T P I S V L L A G V L L K L G T Y G L L R F G M N L L P A A WN Y L A P WL A A WA V V S V LE A S T P V S V L L A G V L L K L G T Y G L L R F G L G L Y L E A WV E F A P Y L A T L A A I S A LE A H Y S T C M L L A G I L L K M G A Y G L V R I N M E L L P H A H S I F S P WL M I I G T M Q I IQ A P T A G S V D L A G I L L K T A A Y G L L R F S L P L F P N A S A E F A P I A M WL G V I G I FE A S T P . S V L L A G V L L K . G T Y G L L R F G . L . P A W. F A P WL A . . A . I S . L
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
310 320 330 340 350Y G A S C A I A Q K D M K K V V A Y S S I A H M A F I L L A A A A A T P L S L A A A E I Q M V S H GY G S S C A I A Q T D M K K M V A Y S S I G H M G Y V L L A A A A A T P L S T L G A V M Q M I S H GY G A S C A I A Q K D M K K V V A Y S S I A H M G Y I L L A A A A A T R L S V T A A S A Q M V S H GY A A S T S P G Q R N L K K R I A Y S S V S H M G F I I I G I S S I T D T G L N G A I L Q I I S H GY G A WM A F A Q T D I K R L I A Y T S V S H M G F V L I A I Y T G S Q L A Y Q G A V I Q M I A H GY G A S C A I A Q . D M K K . V A Y S S I . H M G F I L L A A A A A T L S . G A . . Q M I S H G
336
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
360 370 380 390 400L I S G L L F L L V G I V Y K K T G S R D V D Y L R G L L T P E R G L P L T G S L M I L G V M A S AL I S A L L F L L V G V V Y K K A G S R D L D V I R G L L N P E R G M P V I G T L M I V G V M A S AI I S A L L F L L V G V V Y K K T G S R D V D K L Q G L L T P E R G L P I T G S L M I L G V M A S AF I G A A L F F L A G T S Y D R I R L V Y L D E M G G I A I P - - - M P K I F T L F S S F S M A S LL S A A G L F I L C G Q L Y E R I H T R D M R M M G G L WS K M K WL P - - - A L S L F F A V A T LL I S A L L F L L V G . V Y K K . G S R D . D . G L L . P E R G L P . G . L M I . G V M A S A
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
410 420 430 440 450G L P G M A G F I A E F L I F R G - - - - - S F P V Y P - - V A T L L C M V G T G L T A V Y F L L MG T P G M V G F I S E F I I F R G - - - - - S F A V F P - - V Q T L L S M I G T G L T A V Y F L I LG I P G M V G F I A E F L V F R G - - - - - S F P I F P - - T Q T L L C L I G S G L T A V Y F L L MA L P G M S G F I A E L I V F F G L I T S Q K Y L L I P K L L I T F G M A I G M I L T P I Y L L S MG M P G T G N F V G E F M I L F G - - - - - S F Q V V P - - V I T V I S T F G L V F A S V Y S L A MG . P G M . G F I A E F . I F R G S F V . P V . T L L . I G . G L T A V Y F L . M
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
460 470 480 490 500I N K V F F G R L T P E L I N - - M S P V N WA D Q F P A V M L V I L L F V F G L Q P Q WL V R WSV N K A F F G R L S A Q V M N - - L P R I Y WS D R A P A F I L A V L I V I F G I Q P S WL A R WTI N R V F F G R L T M E L S H - - L P K V R WQ E Q I P A I A L A V V I I A L G I Q P H WL T Q WSS R Q M F Y G Y K L F N I S N S S F F D S G P R E L F V S T S I F L P V I G I G V Y P D L V L S L SL H R A Y F G K A K S Q I A S Q E L P G M S L R D V F M I L L L V V L L V L L G F Y P Q P I L D T S. N . . F F G R L . . . N L P . W D F P A . . L . V L . . . . G . Q P WL . WS
NdhD4-7002(sll0027)NdhD4-7120(sll0027)NdhD4-6803(sll0027)NdhD-Spinacia oleraceaNuoM-E. coli
510 520 530 540 550E I D T A A L V A - - - - - - - - - S P T A I E I S L K NE P T I T A M F S - - - - - - - - V E S T V A T V S L N K V K E K SE P Q T A V L L T G H P D L V S V P A P P R I E I E V K E L G E VV E K V E A I L S - - - - - - - - - - N Y F Y RH S A I G N I Q Q - - - - - - - - - - WF V N S V T T T R PE . . A . . . . . . . .
337
Figure B9. ClustalW alignment of NdhD amino acid sequences from Synechococcus sp.
PCC 7002 against each other. The Synechococcus sp. PCC 7002 NdhD amino acid
sequences were aligned using the ClustalW alignment program from MacVector v. 6.5.
The consensus amino acid sequence is underneath the alignments. Gray high-lighted
regions indicate identical amino acids while lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids determined with the
ClustalW program (Thompson et al., 1994b).
338
NdhD ClustalW Amino Acid Alignment
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
10 20 30 40M N F A N F P W L S T I I L F P I I A A L F L P L I P D K D G K T V R W Y A L TM D S L Q I P W L T T A I A F P L L A A L V I P L I P D K E G K T I R W Y T L W
M L S F L L F L P L V G I G A I A L F P R - - - - P L T R I V A TM L S A L I W L P L A G A L L V A I L P Q G E K N Q F S R T M A L
L S . I P L . . A L . . L . P . . .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
50 60 70 80I G L I D F V I I V T A F Y T G Y D F G N P N L Q L V E S Y T W V E A I D L R WR C P H R F C L L V T A F W Q N Y D F G R T E F Q L T K N F A W I P Q L G L N WV F T V V T L A I S S G L L I N L N L Q D A G M Q Y T E F H N W L S I L G L N YG A A A L V F V W T A W L G F H Y D V A I A G L Q F V E H Y L W I E W L G L N Y. . . . . . Y D . Q E . W . L G L N .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
90 100 110 120S V G A D G L S M P L I L L T G F I T T L A I L A A W P V S F K P K L F Y F L MS L G V D G L S M P L I I L A T L I T T L A T L A A W N V T K K P K L F A G L IN L G V D G L S L P L I V L N S L L T L V A I Y S I G E S N H R P K L Y Y S L ID L G V D G L S L P L L A L N A L L T L V A L W I S P K D L H R P R F Y Y A L F
L G V D G L S P L I . L L . T . A . . . . . P K L . Y L
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
130 140 150 160L L M Y G G Q I A V F A V Q D M L L F F F T W E L E L V P V Y L I L S I W G G KL V M L S A Q I G V F A V Q D L L L F F I M W E L E L V P V Y L L I S I W G G KL L I N S G I T G A L I A N N L L L F F L F Y E I E L I P F Y L L I A I W G G EL L L Q A S V N G A F L A Q D V L L F F L F Y E I E I I P L Y F L I A I W G G KL L . G . F . . Q D . L L F F . . E . E L . P . Y L L I I W G G K
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
170 180 190 200K R L Y A A T K F I L Y T A G G S L F I L I A A L T M A F Y G D T V T F D M T AK R L Y A A T K F I L Y T A L G S V F I L A F T L A L A F Y G G D V T F D M Q AK K G Y A S T K F L I Y T A I S G L C V L A A F L G I V W L S Q S S N F D F E NK R G Y A A I K F L L Y T A V S G I L I L A S F L G L A F L T E S N T F A Y S AK R . Y A A T K F . L Y T A . . I L A L . . A F . T F D A
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
210 220 230 240I A Q K D F G I N L Q L L L Y G G L L I A Y G V K L P I F P L H T W L P D A H GL G L K D Y P L A L E L L A Y A G F L I G F G V K L P I F P L H S W L P D A H SL T L E N L E F N T K V I L L T I L L I G F G I K I P L V P L H T W L P D A Y VL H S D L L P L T T Q L I L L G G I L V G F G I K I P F L P F H T W L P D A H VL . L . L . G . L I G F G . K . P . P L H T W L P D A H .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
250 260 270 280E A T A P A H M L L A G I L L K M G G Y A L L R M N A G M L P D A H A L F G P VE A S A P V S M I L A G V L L K M G G Y G L I R L N M E M L P D A H I R F A P LE A N P A V T V L L G G V F A K L G T Y G L V R F G L Q L F P D V W S T V S P AE A S T P V S V I L A G V L L K V G T Y G L L K F G I G L F P L A W A V V A P WE A . P V . . L A G V L L K G Y G L . R . P D A . . P .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
290 300 310 320L V I L G V V N I V Y A A L T S F A Q R N L K R K I A Y S S I S H M G F V L I GL I V L G I V N I V Y G A L T A F G Q T N L K R R L A S S S I F P H G L S S L GL A V I G T V S V M Y G S L A A I A Q R D L K R M V A Y S S I G H M G Y I L V SL A I W A A I S A L Y G A S C A I A Q K D M K K V V A Y S S I A H M A F I L L AL . . . G . V . . Y G A L A A Q . L K R . A Y S S I H M G . . L . .
339
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
330 340 350 360M A S F T D L G T S G A M L Q M I S H G L I G A S L F F M V G A T Y D R T H T LL L S F T D L G M N G A V L Q M L S H G F I A A A L F F L S G V T Y E R T H T LT A A G T E L S L L G A V A Q M I S H S L I L A L L F H L V G I I E R K V G T RA A A A T P L S L A A A E I Q M V S H G L I S G L L F L L V G I V Y K K T G S R
A T . L G A . Q M . S H G L I . A . L F L V G . Y . T T
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
370 380 390 400M L D E M G G V G - - - K K M K K I F A M W T T C S M A S L A L P G M S G F V AM M D E M S G I A - - - R L M P K T F A M F T A A A M A S L A L P G M S G F V SD L D V L N G L M N P V R G L P L T S S L L I L A G M A S A G I P G L V G F V AD V D Y L R G L L T P E R G L P L T G S L M I L G V M A S A G L P G M A G F I A
. D G . . R . P T . . . M A S . . L P G M G F V A
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
410 420 430 440E L M V F V G F A T S D A Y S P T F R V I I V F L A A V G V I L T P I Y L L S ME L T V F L G L S N S D A Y S Y G F K P I A I F L T A V G V I L T P I T C F Q CE F L V F Q G - - - - - - - S F S R F P I P T L F C I I A S G L T A V Y F V I LE F L I F R G - - - - - - - S F P V Y P V A T L L C M V G T G L T A V Y F L L ME V F G S . P I . L . V G . L T . Y . .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
450 460 470 480L R E I L Y G P E N K E L V A H E K L I D A E P R E V F V I A C L L I P I I G IC G V F Y G - - K G S Q A P P R C G G E D A K P R E I F V A V C L L A P I I A IL N R T C F G R L D S H T A Y Y P K V F A S - - - E K I P A I A L T V I I L F LI N K V F F G R L T P E L I N M S P V N W A - - - D Q F P A V M L V I L L F V F. . G . . . A E F A . . L . . I . . .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
490 500 510 520G L Y P K A V T Q I Y A S T T E N L T A I L R Q S V P S L Q Q T A Q A P - - - -G L Y P K L A T T T Y D L K T V E V A S K V R A A L P L Y A E Q L P Q N G D R QG L Q P A W L T R W I E P T T S Q F I A A I P T V Q T I A L T P A E L S - - - -G L Q P Q W L V R W S E I D T A A L V A S P T A I E I S L K NG L P . T . T . . A . . . .
NdhD1-7002(slr0331)NdhD2-7002 (slr1291)NdhD3-7002(sll1773)NdhD4-7002(sll0027)
530 540 550 560- - - - - S L D V A V L R A P E I R NA Q M G L S S Q M P A L I A P R F- - - - - - - - - - - - K A P
A P
340
Figure B10. ClustalW alignment of NdhF1 amino acid sequences. NdhF1 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b). Data for NdhF1 was acquired from Schluchter et al.
(Schluchter et al., 1993).
341
NdhF1 ClustalW Amino Acid Alignment
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
10 20 30 40 50ME P L Y Q Y A WL I P V L P L L G A MV I G I G L I S L N K F T N K L R Q L Y A V F V L S L I G TME L L Y Q L A WL I P V L P L F G A T V V G I G L I S F N Q A T N K L R Q I N A V F I I S C L G AME V I Y Q Y A WL I P V L P L L G A ML V G L G L I S F N Q T T N R L R Q L N A V L I I S L MG AME H I Y Q Y A WI I P F L P L P V P L L I G A G L L F F P T A T K N L R R I WA F S S I S L L S I
MN ML A L T I I L P L I G F V L L A F S R G R WS E N V S A I V G V G S V A L A A L V T AME . Y Q Y A WL I P V L P L . G A L . G . G L I S F N T N . L R Q . A V . I S L . G A
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
60 70 80 90 100S MA L S F G L L WS Q I Q G H E A F T Y T L E WA A A G D F H L Q MG Y T V D H L S A L MS V I VA L V MS G A L L WD Q I Q G H A S Y A Q MI E WA S A G S F H L E MG Y V I D H L S A L ML V I VA L G L S S A L L WS Q L Q G H P T Y L R T L E WA A A G N F H L T MG Y T I D N L T S L ML V I AV MI F S MK L A I Q Q I N S N S I Y Q Y L WS WT I N N D F S L E F G Y L MD P L T S I MS ML IL S A L I S S L T A S K T Y S - - - - Q P L WT WMS V G D F N I G F N L V L D G L S L T ML S V V. . L S . L L WS Q I Q G H Y . E WA . A G D F H L MG Y . . D L S . L ML V I V
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
110 120 130 140 150T T V A L L V MI Y T D G Y MA H D P G Y V R F Y A Y L S I F S S S ML G L V F S P N L V Q V Y I FT S V A L L V MI Y T D G Y MA H D P G Y V R F Y A Y L S L F A S S ML G L V I S P N L V Q V Y I FT S V A V L V MV Y T D G Y MA H D P G Y V R F Y A Y L S L F G S S ML G L V V S P N L V Q I Y I FT T V A I L V L I Y S D N Y MS H D Q G Y L R F F A Y MS F F N T S ML G L V T S S N L I Q I Y I FT G V G F L I H MY A S WY MR G E E G Y S R F F A Y T N L F I A S MV V L V L A D N L L L MY L GT . V A . L V MI Y T D G Y MA H D P G Y V R F Y A Y L S L F . S S ML G L V . S P N L V Q . Y I F
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
160 170 180 190 200WE L V G MC S Y L L I G F WY D R K A A A D A C Q K A F V T N R V G D F G L L L G ML G L Y WA TWE L V G MC S Y L L I G F WY D R K A A A D A C Q K A F V T N R V G D F G L L L G I L G L Y WA TWE L V G MC S Y L L V G F WY D R K S A A D A A Q K A F V T N R V G D F G L L L G I L G L F WA TWE L V G MC S Y L L I G F WF T R P I A A N A C Q K A F V T N R V G D F G L L L G I L G L Y WI TWE G V G L C S Y L L I G F Y Y T D P K N G A A A MK A F V V T R V G D V F L R F A L F I L Y N E LWE L V G MC S Y L L I G F WY D R K . A A D A C Q K A F V T N R V G D F G L L L G I L G L Y WA T
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
210 220 230 240 250G S F E F D L MG D R L MD L V S T G Q I S S L L A I V F A V L V F L G P V A K S A Q F P L H V WLG S F D F G T I G E R L E G L V S S G V L S G A I A A I L A I L V F L G P V A K S A Q F P L H V WLG S F D F Q I MG D R L A E L V Q T G S I S N F L A V L F A I L V F L G P V A K S A Q F P L H V WLG S F E F R D L F E I F N N L I K N N E V N S L F C I L C A F L L F A G A V A K S A Q F P L H V WLG T L N F R E MV E L A P A H L A D G - - - N N ML MWA T L ML L G G A V G K S A Q L P L Q T WLG S F . F MG E R L L V . G . S . . A . . A . L V F L G P V A K S A Q F P L H V WL
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
260 270 280 290 300P D A ME G P T P I S A L I H A A T MV A A G V F L V A R MY P V F E P I P E A MN V I A WT G A TP D A ME G P T P I S A L I H A A T MV A A G V F L V A R MY P V F E P I P V V MN T I A F T G C FP D A ME V P T P I S A L I H A A T MV A A G V F L V A R MY P V F E H V P A A MN V I A F T G A FP D A ME G P T P I S A L I H A A T MV A A G I F L V A R L L P L F V V I P Y I MY V I S F I G I IA D A MA G P T P V S A L I H A A T MV T A G V Y L I A R T H G L F L MT P E V L H L V G I V G A VP D A ME G P T P I S A L I H A A T MV A A G V F L V A R MY P V F E I P . MN V I A F T G A
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
310 320 330 340 350T A F L G A T I A L T Q N D I K K G L A Y S T MS Q L G Y MV MA MG I G G Y T A G L F H L MT H AT A F L G A T I A L T Q N D I K K G L A Y S T I S Q L G Y MV MA MG I G A Y S A G L F H L MT H AT A F L G A T I A I T Q N D I K K G L A Y S T I S Q L G Y MV MA MG V G A Y S A G L F H L MT H AT V L L G A T L A L A Q K D I K R S L A Y Y T MS Q L G Y MML A L G MG S Y R T A L F H L I T H AT L L L A G F A A L V Q T D I K R V L A Y S T MS Q I G Y MF L A L G V Q A WD A A I F H L MT H AT A F L G A T I A L T Q N D I K K G L A Y S T MS Q L G Y MV MA MG . G A Y . A G L F H L MT H A
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
360 370 380 390 400Y F K A ML F L G S G S V I H G ME E V V G H N A V L A Q D MR L MG G L R K Y MP I T A T T F L IY F K A ML F L C S G S V I H G ME G V V G H D P I L A Q D MR I MG G L R K Y MP I T A T C F L IY F K A ML F L G S G S V I H G ME A V V G H D P A L A Q D MR L MG G L R K Y MP A T G L T F L IY S K A L L F L A S G S L I H S MG T I V G Y S P D K S Q N MV L MG G L T K H V P I T K T S F L IF F K A L L F L A S G S V I L A C H - - - - H - - - - E Q N I F K MG G L R K S I P L V Y L C F L VY F K A ML F L . S G S V I H G ME V V G H P . L A Q D MR L MG G L R K Y MP I T . T . F L I
342
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
410 420 430 440 450G T L A I C G I P P F - A G F WS K D E I L G L A F E A N P V - L WF I G WA T A G MT A F Y MF RG T L A I C G I P P F - A G F WS K D E I L G L A F Q A N P L - L WF V G WA T A G MT A F Y MF RG C L A I S G I P P F - A G F WS K D E I L G A A Y A S N P L - L WF I G WMT A G I T A F Y MF RG T L S L C G I P P L - A C F WS K D E I L N D S WV Y S P I - F A I I A Y F T A G L T A F Y MF RG G A A L S A L P L V T A G F F S K D E I L A G A MA N G H I N L MV A G L V G A F MT S L Y T F RG T L A I C G I P P F A G F WS K D E I L G . A . . N P . L WF I G W. T A G MT A F Y MF R
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
460 470 480 490 500MY F L T F E G - - - - - - - - - - - - - - - - - - - - - - - - - - E F R G T D Q Q L Q E K L L T AMY F MT F E G - - - - - - - - - - - - - - - - - - - - - - - - - - G F R G N D Q E A K D G V L Q FMY F S T F E G - - - - - - - - - - - - - - - - - - - - - - - - - - K F R G N D E K I K D K L L K AI Y L L T F E G H L N F F C K N Y S G K K S S S F Y S I S L WG K K E L K T I N Q K I S L L N L L TMI F I V F H G - - - - - - - - - - - - - - - - - - - - - - - - - - - K - - - E Q - - - - - - I H AMY F . T F E G F R G D Q . . . L A
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
510 520 530 540 550A G Q A P - - - - - - - - - - - - - - - - - - - - - - - - - - - - - E E G H H G S K P H E S P L T MY G L L P N F G P G A MN V K E L D H E A G H - - - - - - - - - - - D D H G H S S E P H E S P L T MK T I L L E L E S A E P T P V F G P G A MK K G E L A A T G G H H D G H G H H S S S P H E S P WT MMN N K E R A S F F S K K P Y E I N V K L T K L L R S F I T I T Y F E N K N I S L Y P Y E S D N T MH A V K G - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - V T H
. . . . . . H S S P H E S P . T M
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
560 570 580 590 600T F P L MA L A V P S V L I G L L G V P WG N - - - - - R F E A F V F S P N E A A E A A E H G - - -T F P L MA L A V P S V L I G L L G R P WA N - - - - - Q F E A F I H A P G E V V E H A A E - - - -T L P L L I L A I P S ML I G L V G T P Y N N - - - - - Y F E R F I F S P T E S L A E V L E K A A EL F P L I I L I MF T L F V G F I G I P F N Q E G MD L D I L T K WL T P S I N L L H S N S E N - FS L P L I V L L I L S T F V G A L I V P P L Q G - - - - - - - - - V L P Q T T E L A H G - - - - - -T F P L . . L A . P S . L I G L L G . P . . N F E F . . P . E L . H .
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
610 620 630 640 650F E L T E F L I MG G N S V G I A L I G I T I A S L MY L Q Q R I D P A R L A E K - - - - - - - - -F E WG E F Y V MA G N S I G I A L I G I T V A S L MY WQ H K F D P K V L A E K - - - - - - - - -F D P N E F Y I MA G G S V G V S L I G I T L A S L MY L Q R K I D P A A I A A K - - - - - - - - -V D WY E F V I N A I F S I S I A F F G I F I A F F F Y K P I Y S S L K N F D L I N S F D K R G Q KS - - - - ML T L E I T S G V V A V V G I L L A A WL WL G K R T L V T S I A N S A P - - - - - - -F . E F . I MA G S . G I A L I G I T . A S L MY L Q . . D P . A K
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
660 670 680 690 700- - - - - - - F P V L Y Q L S L N K WY F D D I Y N N V F V MG T R R L A R Q I L E V D Y R V V D G- - - - - - - F P S L Y Q L S L N K WY F D D L Y D K L F V Q G S R R V A R Q I ME V D Y K V I D G- - - - - - - I K P L Y E L S L N K WY F D D I Y H R V F V L G L R R L A R Q V ME V D F R V V D GR I L G D N I I T I I Y N WS A N R G Y I D A F Y S T F L I K G I R S L S E L V S F F D R R I I D G- - - - - - - G R L L G T WWY N A WG F D WL Y D K V F V K P F L G I A WL L K - - - R D P L N S
. . L Y . L S L N K WY F D D . Y . V F V G R R L A R Q . E V D . R V . D G
NdhF1-7002(slr0844)NdhF1-6803(slr0844)NdhF1-7120(slr0844)NdhF-Spinacia oleraceaNuoL-E. coli
710 720 730 740 750A V N L T G I A T L L S G E G L K Y I E N G R V Q F Y A L - I V F G A V L G F V I F F S V AA V N L T G L V T L V S G E G L K Y L E N G R A Q F Y A L - I V F G A V L G F V I V F S L TA V N L T G F F T L V S G E G L K Y L E N G R V Q F Y A L - I V F G A V L G L V I V F G V TI P N G F G V T S F F V G E G I K Y V G G G R I S S Y L F - WY L L Y V S I F L F I F T F TMMN I P A V L S R F A G K G L L L S E N G Y L R WY V A S MS I G A V V V L A L L MV L RA V N L T G . . T L . S G E G L K Y . E N G R . Q F Y A L I V F G A V L G F V I . F . . T
343
Figure B11. ClustalW alignment of NdhF2 amino acid sequences. NdhF2 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
344
NdhF2 ClustalW Amino Acid Alignment
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
10 20 30 40 50MN E L T I G WV I F P F V V G F S I Y L L P K I D R - - - - - - - - Y L A I
MN F P T A I A P N N L L I A I V A L L V L A L MG A F G G Y L F R P L V R - - - - - - - - P S A LME H I Y Q Y A WI I P F L P L P V P L L I G A G L L F F P T A T K N L R R I WA F S S I S
MN ML A L T I I L P L I G F V L L A F S R G R WS E N V S A I V G V G S V A L A A. . . . . . L . . . . . . F . . . . A .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
60 70 80 90 100F V S I C S L I F G F V Q I F Q P E P - - - - - - - Y S L K L L G MY G V D L - - L V D D Q S G Y FL MT L G T I L L A A V G F A L P N A - - - - - - - Q Q WQ L MD R F G I L L - - Q L D N L G S Y FL L S I V MI F S MK L A I Q Q I N S N S I Y Q Y L WS WT I N N D F S L E F G Y L MD P L T S I ML V T A L S A L I S S L T A S K T Y S - - - - Q P L WT WMS V G D F N I G F N L V L D G L S L T ML . . . . . . . . . . . W . F . . . D L .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
110 120 130 140 150I L T N A A V A I A V T V Y - - C WK S A K S - - A F F F T Q L V V L Q G A L N A V F V C A D L I SL L T N G L V T L A V L L Y - - C WA S P R T - - T F F Y V Q L MV L H V S L N A A F L S T D L I SS ML I T T V A I L V L I Y S D N Y MS H D Q G Y L R F F A Y MS F F N T S ML G L V T S S N L I QL S V V T G V G F L I H MY A S WY MR G E E G Y S R F F A Y T N L F I A S MV V L V L A D N L L L. . V . . . V . Y . S F F . . . S . . . L I
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
160 170 180 190 200L Y V A L E A I S I A A F L L MT Y Q R T D R S I WI G L R Y L F L S N T A ML F - Y L I G A V L VL Y V C L E V V G L S S F L L I I Y P R Q A A S S WI G L R Y L F V T N T A L L F - Y L I G V ML VI Y I F WE L V G MC S Y L L I G F WF T R P I A A N A C Q K A F V T N R V G D F G L L L G I L G LMY L G WE G V G L C S Y L L I G F Y Y T D P K N G A A A MK A F V V T R V G D V F L R F A L F I L. Y . E . V G . S . L L I . . T . . . . F V . N . . F L . G . . .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
210 220 230 240 250Y Q A T K S F A F V G L A E A P S D A I A - - - - - - - - - - - - - - L I F L G L L T K G G V F V SY Q A T N S L D F Q G L A T A P Y E A I A - - - - - - - - - - - - - - L I F L G L L I K G E I F L SY WI T G S F E F R D L F E I F N N L I K N N E V N S L F C I L C A F L L F A G A V A K S A Q F P LY N E L G T L N F R E MV E L A P A H L A D G - - - N N ML MWA T L ML L G G A V G K S A Q L P LY . . T S F L . E . . I A L . F . G . . . K . F
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
260 270 280 290 300G L WL P L T H S E A E T P V S A ML S G - V V V K A G I F P L L R C G - - - I L V P D L D L WL RG L WS P Q T S S I A S A P V A A L L S G - I V V K A G I L P L L R F A - - - S L S E R L A MMV WH V WL P D A M- E G P T P I S A L I H A A T MV A A G I F L V A R L L P L F V V I P Y I MY V I SQ T WL A D A M- A G P T P V S A L I H A A T MV T A G V Y L I A R T H G L F L MT P E V L H L V G
. WL P . T P V S A L . . V A G I . . . R . . . P . .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
310 320 330 340 350L F G L A T A L L G I I F A I L E T D A K R L L A F S T I S K L G L L L S A P A V A G - - - - - L AG L A I A T A L L G MG L G MF A R D S R R I L A Y S T I S Q MG F I L V A P A V G G - - - - - L YF I G I I T V L L G A T L A L A Q K D I K R S L A Y Y T MS Q L G Y MML A L G MG S Y R T A L F HI V G A V T L L L A G F A A L V Q T D I K R V L A Y S T MS Q I G Y MF L A L G V Q A WD A A I F H. . G . . T . L L G . . A . . D . K R . L A Y S T S Q . G . . A . V . .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
360 370 380 390 400A L S H G L V K S S L F L MA G Q L P T - - - - - R N F Q E L R Q T - - - - - - - - - - - - K I A SA L T H G L A K A C L F L L V G S L P E - - - - - R D L D K L Q A Q - - - - - - - - - - - - P I S YL I T H A Y S K A L L F L A S G S L I H S MG T I V G Y S P D K S Q N MV L MG G L T K H V P I T KL MT H A F F K A L L F L A S G S V I L A C H H E Q N I F K MG G L R - - - - - - - - K S I P L V Y. . T H . K A L F L . G S L P I
345
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
410 420 430 440 450S - - - - - - - - - - - - - - - - - - L WL P - - - - - - - - - - - - - - L A I A C L S MV G MP LK - - - - - - - - - - - - - - - - - - L WL P - - - - - - - - - - - - - - MV L A S S S I I G L P IT S F L I G T L S L C G I P P L A C - F WS K D E I L N D S WV Y S P I - F A I I A Y F T A G L T AL C F L V G G A A L S A L P L V T A G F F S K D E I L A G A MA N G H I N L MV A G L V G A F MT S
W . . A . G .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
460 470 480 490 500L V G F S S K A L L L K - - - - - - - - - - - - - - - - - - - N I A P WQ A - - - - - - - - - - - -L A G F E A K T L T L E T L S - - - - - - - - - - - - - - - L N E L P WT G - - - - - - - - - - - -F Y MF R I Y L L T F E G H L N F F C K N Y S G K K S S S F Y S I S L WG K K E L K T I N Q K I S LL Y T F R MI F I V F H G K E Q - - - - - - - - - - - - - - I H A H A V K G - - - - - - - - - - - -L F L . W .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
510 520 530 540 550- - - MG L N - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -- - - I L MN - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -L N L L T MN N K E R A S F F S K K P Y E I N V K L T K L L R S F I T I T Y F E N K N I S L Y P Y E- - - V T H S - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -
. N
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
560 570 580 590 600- - - - - - - - I A A V G T A L A F A K F L F I P - - - - - - - H D A A T K F T A K S T T F WG - -- - - - - - - - L A G V G T A I I L A K F I F L I P S - - - - - F D K N V D L D K S P WG L F L - -S D N T ML F P L I I L I MF T L F V G F I G I P F N Q E G MD L D I L T K WL T P S I N L L H S N- - - - - - L P L I V L L I L S T F V G A L I V P P L Q G - - V L P Q T T E L A H G S ML T L E I T
L . . . . . . F . F . . P D T S
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
610 620 630 640 650- - - - - - - - - - - - - - - - A I A F L F S G V I L G N G F Y L E A Y Q L D N I P K A L I K I A -- - - - - - - - - - - - - - - - A V L L L L G A L T L G N V I Y P E A F S ME N G I K A T A S F L -S E N F V D WY E F V I N A I F S I S I A F F G I F I A F F F Y K P I Y S S L K N F D L I N S F D K- - - - - - - - - - - - - - - - S G V V A V V G I L L A A WL WL G K R T L V T S I A N S A P G R -
. . . . G . L . Y . . . . .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
660 670 680 690 700- - - - - - - - - - I G WA L Y - - - - - - WL I MK - - - - - - - - - - - - - - - - - - - - - - -- - - - - - - - - - L G S A I Y - - - - - - WWG L R - - - - - - - - - - - - - - - - - - - - - - -R G Q K R I L G D N I I T I I Y N - - - - - WS A N R G Y I D A F Y S T F L I K G I R S L S E L V S- - - - - - - - - L L G T WWY N A WG F D WL Y D K V F V K P F L G I A WL L K - - - - - - - - -
. G . . . Y W . .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
710 720 730 740 750- - - - R I E F K L P R I F E A F E Q L I G A MS V V L T G - - - - - - L F WMV T L- - - - K I P WQ P P D WG E R L D H L I G T MA I ML ML - - - - - - L F S S V L IF F D R R I I D G I P N G F G V T S F F V G E G I K Y V G G G R I S S Y L F WY L L Y V S I F L F I- - - R D P L N S MMN I P A V L S R F A G K G L L L S E N G - - - - Y L R WY V A S MS I G A V V
. I P . . . G . . . . L F W V .
NdhF2-7002(slr2009)NdhF2-6803(slr2009)NdhF-Spinacia oleraceaNuoL-E. coli
760 770 780 790 800
F T F TV L A L L MV L R
346
Figure B12. ClustalW alignment of NdhF3 amino acid sequences. NdhF3 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
347
NdhF3 ClustalW Amino Acid Alignment
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
10 20 30 40 50MN T F F S Q S V WL V P C Y P L - - - L G MG L S A L WMP S I T R K T G P R P A G Y V N ML L TML E S L S R I I WL V P C Y A L - - - L G A L L A V P WS P G L T R Q T G P R P A G Y I S T L MTMA Q F L L E T V WL V P L Y A L - - - I G G L L A I P WS P G I I R K T G P R P A G Y V N L I MT
ME H I Y Q Y A WI I P F L P L P V P L L I G A G L L F F P T A T K N L R R I WA F S S I S L L SMN ML A L T I I L P L I G F - - V L L A F S R G R WS E N V S A I V G - V G S V A L A A L V T
M . . WL V P Y . L L G . . L . . WS P . T R T G P R P A G Y . L . T
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
60 70 80 90 100F MA L V H S C L A F I E R WE Q P A L K P S L T WL Q A A D L T L S I D L D I S S I T I G A L I LF V A F L H S L L V L I H I WQ Q P A I D L S F P WL H A A D L E I N F D L K I S T V N I A A L V LF L A L V H S A I A L Q A T WS H P P Q E V F L P WL S T A G L D L T I A I E I S S I S V G A MV VI V MI F S MK L A I Q Q I N S N S I Y Q Y L WS WT I N N D F S L E F G Y L MD P L T S I MS MLA L S A L I S S L T A S K T Y S Q P - - - - L WT WMS V G D F N I G F N L V L D G L S L T ML S VF . A . . H S L A . WS Q P . . . WL . A D L L F L I S . . . . . A L . L
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
110 120 130 140 150I A G I N L L A Q L Y A V A Y L E MD WG WA R F F A T MS L F E A G MC A L V L C N S L F F S Y VI T G L N L G A Q I Y A I G Y L E R D WG WA R F F S L MA L F E A G L C T L V L C N S L F F S Y VI T G L N F L A Q I F A I G Y ME MD WG L G R F Y S L L G L F E A G L C A L V L C N N L F F S Y VI T T V A I L V L I Y S D N Y MS H D Q G Y L R F F A Y MS F F N T S ML G L V T S S N L I Q I Y IV T G V G F L I H MY A S WY MR G E E G Y S R F F A Y T N L F I A S MV V L V L A D N L L L MY LI T G . N . L A Q I Y A . . Y ME D WG . . R F F A M L F E A G MC . L V L C N N L F F S Y V
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
160 170 180 190 200V L E I L T L G T Y L L I G Y WF N Q S L V V T G A R D A F L T K R V G D L F L L MG V V A L L P LV L E I L T L G T Y L L I G Y WF N Q S L V V T G A R D A F L T K R V G D L I L L MG V V A L L P LI L E I L T L G T Y L L V G L WF S Q P L V V S G A R D A F L T K R V G D L F L L MG V L G L WP LF WE L V G MC S Y L L I G F WF T R P I A A N A C Q K A F V T N R V G D F G L L L G I L G L Y WIG WE G V G L C S Y L L I G F Y Y T D P K N G A A A MK A F V V T R V G D V F L R F A L F I L Y N E. L E I L T L G T Y L L I G . WF . Q P L V V . G A R D A F L T K R V G D L F L L MG V . . L . P L
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
210 220 230 240 250A G T WN F D G L A E WA A T A E L D P T L A T L L C L A - - - - L I A G P L G K C A Q F P L H L WA G S WN Y D D L A Q WA A S A D L N P T A A T L L C L A - - - - L I A G P L A K C A Q F P L H L WA G T WN Y P E L A Q WA Q T A N V D P T I I T L V G L A - - - - L V A G P MG K C A Q F P L H L WT G S F E F R D L F E I F N N L I K N N E V N S L F C I L C A F L L F A G A V A K S A Q F P L H V WL G T L N F R E MV E L A P A H L A D G N N ML MWA T L - - - ML L G G A V G K S A Q L P L Q T WA G T WN F . L A E WA . A . D P T . . T L . C L A L . A G P . G K C A Q F P L H L W
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
260 270 280 290 300L D E A ME S P V P A - T V V R N S L V V G T G A WV L I K L Q P I F A L S D F A S T F MI A I G AL D E A ME G P I P A - T I L R N T L V V A T G A WV L V K V Q P I L A L S P V A L T V MI A I G SL D E A ME G P MP S - T I L R N S V V V A S G A WV L I K L Q P V L T L S P V V S S F I V A I G AL P D A ME G P T P I S A L I H A A T MV A A G I F L V A R L L P L F V V I P Y I MY V I S F I G IL A D A MA G P T P V S A L I H A A T MV T A G V Y L I A R T H G L F L MT P E V L H L V G I V G AL D E A ME G P P . T . . R N . . V V A . G A WV L . K L Q P . F . L S P . . . . . A I G A
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
310 320 330 340 350T T A L G A A MV A I A Q I D I K R S L S Y S V S A Y MG MV F MA V G S Q Q D Q T T L V L L L T YV T A I G A S L I A I A Q I D I K R F L S Y V V S A Y MG L V F I A V G T G Q G E T A L Q L I F T YV T A I G G S L I A I A Q I D V K R C L S Y S V S A Y MG L V F I A V G A R Q D E A A L L L V L T HI T V L L G A T L A L A Q K D I K R S L A Y Y T MS Q L G Y MML A L G MG S Y R T A L F H L I T HV T L L L A G F A A L V Q T D I K R V L A Y S T MS Q I G Y MF L A L G V Q A WD A A I F H L MT HV T A L G A . . A I A Q I D I K R L S Y S V S A Y MG V F . A V G Q . T A L L L . T H
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
360 370 380 390 400G V A MA I L V MA I G G V V L V N - - - - - - - - - I S Q D L T Q Y G G L WS R R P I T G I C Y LT F A MA I L V MC V G G I I L N N - - - - - - - - - V T Q D L T Q Y G G L WS R R P I S G L S Y LA V S A A L L V MS T G G I I WN S - - - - - - - - - I T Q D V T Q L G G L WS R R P I S G L A F IA Y S K A L L F L A S G S L I H S MG T I V G Y S P D K S Q N MV L MG G L T K H V P I T K T S F LA F F K A L L F L A S G S V I L A C H - - - - - - - - H E Q N I F K MG G L R K S I P L V Y L C F LA . A L L V MA . G G . I L . . Q D . T Q G G L WS R R P I . G L F L
348
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
410 420 430 440 450V G A A S L V A L P P F G G - F WS L A Q L T T N F WK T S P I L A V I L I T V - N A L T S F S I MV G V A S L I A L P P F G T - F WA WL K L A E N L S A T S P L L V G V L L V V - N L L T A F N V TV G T L G L I G F P P L G G - F WA L L K L A D G L WG T H P WL V G I V I A V - N A L T A F S L VI G T L S L C G I P P L A C - F WS K D E I L N D S WV Y S P I F A I I A Y F T - A G L T A F Y MFV G G A A L S A L P L V T A G F F S K D E I L A G A MA N G H I N L MV A G L V G A F MT S L Y T FV G . A S L . A L P P . G . F WS . L . . W. T S P I L . . I . . . V N . L T A F .
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
460 470 480 490 500R E F G L I F G G K P - - - - - - - - - K Q MT V R S P E G L WA - - - - - - - - - - - - - - - - -R G F C L I F G G E A - - - - - - - - - K P MT V R S P E G L WA - - - - - - - - - - - - - - - - -R E F G L I F G G K A - - - - - - - - - K Q MT E R S P E V H WP - - - - - - - - - - - - - - - - -R I Y L L T F E G H L N F F C K N Y S G K K S S S F Y S I S L WG K K E L K T I N Q K I S L L N L LR MI F I V F H G K E - - - - - - - - - - Q I H A H A V K G V T H - - - - - - - - - - - - - - - - -R F . L I F G G K . K Q MT . R S P E G L W.
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
510 520 530 540 550- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -T MN N K E R A S F F S K K P Y E I N V K L T K L L R S F I T I T Y F E N K N I S L Y P Y E S D N T- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
560 570 580 590 600L V L P MV I L A G F A L H S P - F I L A K L N - - - - - - - F L P D WH Q L N L P L A A - - - - -L V L P MV V T V G F A L H L S - L I L K Q G N - - - - - - - L L P D F A D I N WG L S S - - - - -MV L P MMI L F G F V L H L P - L I L Q A L S - - - - - - - L L P D WA I L N K D V V L - - - - -ML F P L I I L I MF T L F V G - F I G I P F N Q E G MD L D I L T K WL T P S I N L L H S N S E N- S L P L I V L L I L S T F V G A L I V P P L Q G V L P Q T T E L A H G S ML T L E I T S G - - - -
V L P M. I L . G F . L H . L I L L N . L P D W. L N . L .
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
610 620 630 640 650- - - - - V L I I S T MV G G G T A MY - L Y L N E K I S K P I H I F S D P V R E F F A K D - - - -- - - - - V L I A S S L L G V G S S A F - I Y L N P K I T K P I D L P L P V V Q N F F A Y D - - - -- - - - - L L I WS T I F G C S I S S V - I Y L S N I I P K P I R L P WK G L Q D L L A Y D - - - -F V D WY E F V I N A I F S I S I A F F G I F I A F F F Y K P I Y S S L K N F D L I N S F D K R G Q- - - - V V A V V G I L L A A WL WL G K R T L V T S I A N S A P G R L L G T WWY N A WG - - - -
V L I . S . . . G . . . I Y L I K P I . L . . . A . D
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
660 670 680 690 700- - - - - - - - - - - - - - - - - - - L Y T A E L Y K N T V I F A V A L I S K I I D WL D R Y F V D- - - - - - - - - - - - - - - - - - - L Y T D K F Y K L T I V A V I D S I S R L I N WF D K T F V D- - - - - - - - - - - - - - - - - - - F Y T P N L Y R I T I I F S V A Q L S K F A D MI D R F V V DK R I L G D N I I T I I Y N WS A N R G Y I D A F Y S T F L I K G I R S L S E L V S F F D R R I I D- - - - - - - - - - - - - - - - - - - - - F D WL Y D K V F V K P F L G I A WL L K - - - R D P L N
. Y T D L Y . T . I . . . I S . L . . D R V D
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
710 720 730 740 750G V I N F L G L A T L F G G Q S L K Y N N S G Q S Q S Y A L S I V A G I L L F I A A L S Y P L L K HG V I N L I G I V T I F S G Q S L K Y N V S G Q T Q F Y V L S I V L G L T L I G A F L S Y S L L G QG I V N F V G L F S L L G G E G L K Y S T S G Q T Q F Y A L T V L L G V G V L G A WV T WP F WG FG I P N G F G V T S F F V G E G I K Y V G G G R I S S Y L F WY L L Y V S I F L F I F T F TS MMN I P A V L S R F A G K G L L L S E N G Y L R WY V A S MS I G A V V V L A L L MV L RG . . N . . G . . S . F . G G L K Y S G Q . Q . Y . L S . . L G . . . . . A . L . .
NdhF3-7002(sll1732)NdhF3-6803(sll1732)NdhF3-7120(sll1732)NdhF-Spinacia oleraceaNuoL-E. coli
760 770 780 790 800WQ FA FQ F L D L MF
349
Figure B13. ClustalW alignment of NdhF4 amino acid sequences. NdhF4 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
350
NdhF4 ClustalW Amino Acid Alignment
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
10 20 30 40 50MS E F L L Q S V WL V P V Y G I T G A L L T - L P WS L G L I R R T G P R P A A Y L N L I MT F LMS D F L L Q S S WF I P F Y G L I G S I L S - L P WS F R L I K Q T G P R P A A Y F N V F MT L VMN Q F L F A T S WC V P F Y S L L G A L L T - L P WG I G I V R R T G P R P A A Y F N L L T T I V
ME H I Y Q Y A WI I P F L P L P V P L L I - G A G L L F F P T A T K N L R R I WA F S S I S L LMN ML A L T I I L P L I G F V L L A F S R G R WS E N V S A I V G V G S V A L A A L V T A L S
M . F L L Q . . W. . P F Y G L . G . L L . L P WS . . . . T G P R P A A Y . N L . T L .
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
60 70 80 90 100G L L H G S F A F A S L WN MP P Q - - - Q L S L E WL Q V A D L N L S L V I E I S P V N L G A MES A I H G MV A L S A I WQ T P S E - - - Q I V F H WL Q V A D L D L T L A V E I S P V S L G A L SG F A H S L WV F K D I WS R E Q E - - - N L V I T WF Q A A D L N L S F A L E L S P V S MG A T VS I V MI F S MK L A I Q Q I N S N S I Y Q Y L WS WT I N N D F S L E F G Y L MD P L T S I MSA L I S S L T A S K T Y S - - - - - - - - Q P L WT WMS V G D F N I G F N L V L D G L S L T ML S. . . H . A I W. Q . . . . W Q V A D L N L . F . . E . S P V S L G A
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
110 120 130 140 150L V T G I C F MA Q L Y G L G Y L E K D WS I A R F Y G L MG F F E A A L S G L A I S D S L L L S YV V T G I S F L V Q I F G L G Y ME K D WS L A R F Y G L L G F F E A A L G G I A L S D S L F L S YL I T G L S L L A Q I Y A L G Y ME K D WS L A R F F G L L G F F E A A L S G L A I S D S L F L S YL I T T V A I L V L I Y S D N Y MS H D Q G Y L R F F A Y MS F F N T S ML G L V T S S N L I Q I YV V T G V G F L I H MY A S WY MR G E E G Y S R F F A Y T N L F I A S MV V L V L A D N L L L MYL V T G . F L . Q I Y . L G Y ME K D WS . A R F F G L G F F E A A L . G L A . S D S L . L S Y
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
160 170 180 190 200G L L E V L T L S T Y L L V G F WY A Q P L V V T A A R D A F L T K R V G D I L L L MG I V A L S SG L L E ML T L S T Y L L V G F WY A Q P L V V T A A R D A F L T K R V G D I I L L MG V V A L S SG L L E I L T L S T Y L L V G F WY A Q P L V V T A A R D A F WT K R V G D L L L L MA V V T L S TI F WE L V G MC S Y L L I G F WF T R P I A A N A C Q K A F V T N R V G D F G L L L G I L G L YL G WE G V G L C S Y L L I G F Y Y T D P K N G A A A MK A F V V T R V G D V F L R F A L F I L Y NG L L E . L T L S T Y L L V G F WY A Q P L V V T A A R D A F . T K R V G D . . L L MG . V . L S .
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
210 220 230 240 250Y G T G L T F S E L E T WA A N P P - - - - L P P WE A S L V G L A L I S G P I G K C A Q F P L N LY G Q G L T F S Q L D N WA S T V P - - - - V T G I T A T L L G L S L I A G P T G K C A Q F P L N LL A G S L N F S D L Y E WV Q T A N - - - - L D P V T A T L L C L G L I A G P A G K C A Q F P L H LI T G S F E F R D L F E I F N N L I K N N E V N S L F C I L C A F L L F A G A V A K S A Q F P L H VE L G T L N F R E MV E L A P A H L A D - - - G N N ML MWA T L ML L G G A V G K S A Q L P L Q T
. G . L F S . L E WA . . . A . L . . L . L I A G P . G K C A Q F P L . L
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
260 270 280 290 300WL D E A ME G P N P - A G I I R N S V V V S A G A Y V L L K ME P V F T I T P I T S D A L I I I GWL D E A ME G P N P - A G I MR N S V V V S A G A Y V L I K L Q P V F T L S P I A S K T L I V L GWL D E A ME G P N P - A S V MR N S L V V A G G A Y L L Y K L Q P I L I L S P V A L N V L I I I GWL P D A ME G P T P I S A L I H A A T MV A A G I F L V A R L L P L F V V I P Y I MY V I S F I GWL A D A MA G P T P V S A L I H A A T MV T A G V Y L I A R T H G L F L MT P E V L H L V G I V GWL D E A ME G P N P A . . I R N S . V V . A G A Y L L . K L P . F . . . P . . . L I I I G
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
310 320 330 340 350T V T T V G A S L V A L A Q I D I K R A L S H S T S A Y L G L V F I A V G L N Q V D I A L L L L L TT L T V V MT S L I A I A Q I D I K R T L S H S T S V Y L G L V F I A V G L G Q V D I A F L L L F AG V T A I G A S L V S I A Q T D I K R A L S H S T S A Y MG L V F L A V G L E Q G G V A L ML L L TI I T V L L G A T L A L A Q K D I K R S L A Y Y T MS Q L G Y MML A L G MG S Y R T A L F H L I TA V T L L L A G F A A L V Q T D I K R V L A Y S T MS Q I G Y MF L A L G V Q A WD A A I F H L MT. V T . . . A S L . A L A Q D I K R . L S H S T S . Y L G L V F L A V G L Q . D . A L L L . T
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
360 370 380 390 400H A I A K A L L F MS I G A V I L N T H - - - - - - - - - G Q N I T E MG G L WS R MP A T T S A FH A I A K A L L F MS I G S I I F T T S - - - - - - - - - G Q N I T E MG G L WN R MP V T T T S FH A I A K A L L F MS S G S V I F T T H - - - - - - - - - S Q D L T E MG G L WS R MP A T T T A FH A Y S K A L L F L A S G S L I H S MG T I V G Y S P D K S Q N MV L MG G L T K H V P I T K T S FH A F F K A L L F L A S G S V I L A C H - - - - - - - - H E Q N I F K MG G L R K S I P L V Y L C FH A I A K A L L F MS S G S V I . T H Q N I T E MG G L W R MP . T T T F
351
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
410 420 430 440 450V V G S A G L V C L F P L G T - F WT MR R WV D G F WD T P P WL V L L L V G V - N F C S S F N LV V G S A G L L A V F P L G M- F WT WQ K WF S G D WL V S WP L L A L L I F V - N L F S A L N LV V G S A G MV T L L P L G S - F WA ML S WA D G L V R V S P WV I G V L I L V - N G L T A L N LL I G T L S L C G I P P L A C - F WS K D E I L N D S WV Y S P I F A I I A Y F T - A G L T A F Y ML V G G A A L S A L P L V T A G F F S K D E I L A G A MA N G H I N L MV A G L V G A F MT S L Y TV V G S A G L . . L P L G F W. W. G W. S P . . . . L . . V N . T A L N L
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
460 470 480 490 500T R V F R S V F L G A P - - K P K T R R S P E - - - - - - - V V W- - - - - - - - - - - - - - - - -T R V F R L V F L G K P - - Q P K T R R A P E - - - - - - - V P W- - - - - - - - - - - - - - - - -T R V F R L A F WG Q P - - Q Q K T R R A P E - - - - - - - V G W- - - - - - - - - - - - - - - - -F R I Y L L T F E G H L N F F C K N Y S G K K S S S F Y S I S L WG K K E L K T I N Q K I S L L N LF R MI F I V F H G K E - - Q I H A H A V K G - - - - - - - V T H - - - - - - - - - - - - - - - - -T R V F R L V F G . P Q K T R R . P E V . W
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
510 520 530 540 550- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -L T MN N K E R A S F F S K K P Y E I N V K L T K L L R S F I T I T Y F E N K N I S L Y P Y E S D N- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
560 570 580 590 600Q MA V P MV S L I L MT L MV P F F - - - - - - - - - - - - - L H Q WQ L L F N P S L P T L V E RP MA V P MV S L I I V T L L V P I A - - - - - - - - - - - - - P L Q WS F WL S A T Y P L G L T ST MA F P MV T L I I L T L L L P L M- - - - - - - - - - - - - L Q Q WY L L P - - - - - - A WE ST ML F P L I I L I MF T L F V G F I G I P F N Q E G MD L D I L T K WL T P S I N L L H S N S E N- - S L P L I V L L I L S T F V G A L - - - - - - - - - - - - - I - - - V P P L Q G V L P Q T T E L
MA . P MV . L I I . T L V P . . L Q W L P . E
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
610 620 630 640 650P L I V T L A I P A L MI T G G L G L V A G L T I T - - - L N P S L S - - - - - - - - - - R P R Q LP - V T Q WA MP L L MV A G I T G I L L G S L MP - - - L R R N L S - - - - - - - - - - R S S R LI - - D WY V V L V L V S S T V A G V V I G S T I H - - - L H K A WS - - - - - - - - - - R S T V LF - V D WY E F V I N A I F S I S I A F F G I F I A F F F Y K P I Y S S L K N F D L I N S F D K R GA - - - H G S ML T L E I T S G V V A V V G I L L A A WL WL G K R T - - - - - - - - - - L V T S I
. . . . . L I . . . . G . V . G . I L . S R . L
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
660 670 680 690 700Y L R F L Q D L L A Y D F Y - - - - - - - - I D R I Y N V T V V WL V T T L S K L A A WF D R Y V VP V R F L Q D L F A Y D V Y - - - - - - - - L D K I Y G A T V V A A V A A I A K I S T WF D R Y V IA WR F I Q D L L G Y D F Y - - - - - - - - I D R I Y R L T V V S A V A L L S R I S A WS D R Y L VQ K R I L G D N I I T I I Y N WS A N R G Y I D A F Y S T F L I K G I R S L S E L V S F F D R R I IA N S A P G R L L G T WWY N A WG - - - - F D WL Y D K V F V K P F L G I A WL L K - - - R D P L
R F L Q D L L . Y D . Y I D . I Y . T V V . V . . L S . L . WF D R Y . .
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
710 720 730 740 750D G F V N L T G L A T L F S G S A L R Y N V S G Q S Q F Y V L T I V L G MI L G L V WF MA T G Q WD G I V N L V S L V T I F S G S A L K Y N V T G Q S Q F Y L L T I L V G V A L - L I WF S L S G Q WD G L V N L V G F A T I F G G Q G L K Y S I S G Q S Q G Y ML T I L A V V G A L G F F I S WS L G LD G I P N G F G V T S F F V G E G I K Y V G G G R I S S Y L F WY L L Y V S I F L F I F T F TN S MMN I P A V L S R F A G K G L L L S E N G Y L R WY V A S MS I G A V V V L A L L MV L RD G . V N L G . . T . F . G G L K Y . . G Q S Q . Y . L T I L . G V . . . L . . F . . . .
NdhF4-7002(sll0026)NdhF4-6803(sll0026)NdhF4-7120(sll0026)NdhF-Spinacia oleraceaNuoL-E. coli
760 770 780 790 800T MI T D F WS N Q L AMA I R Q F WS S WL S L I L PL D K L P F
F
352
Figure B14. ClustalW alignment of NdhF5 amino acid sequences. NdhF5 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Spinacia oleracea, and E. coli. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
353
NdhF5 ClustalW Amino Acid Alignment
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
10 20 30 40 50M I D D I T I I W I L L P F V V G F S I Y L L P R W N R - - - - - - - Y F A L A
M T L F A V S A E F N A A P W A T V V I C L A L M A G F T G Y L L P A T I R - - - - - - - F L T L AM T T I T L T W I T L P F L L G F I I Y L V P K L D K - - - - - - - Y L A L G
M E H I Y Q Y A W I I P F L P L P V P L L I G A G L L F F P T A T K N L R R I W A F S S I SM N M L A L T I I L P L I G F V L L A F S R G R W S E N V S A I V G - - - - - - - V G S V A
. . I . . . I L P F . . G F . Y L . P . . . . . L A
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
60 70 80 90 100I A A L S V V Y S I G L L W S L E P - - - - - - - - F T L E L L D S F G V T L - - M F D E L S G Y FV C F G T G F L A Y L G F S L P E A - - - - - - - - Q S W Y L L D S F G V V F - - Q L D A L S G Y FA A L A S A G Y A A Q L F V A Q S P - - - - - - - - L E L R L L D N F G V T L - - T L D E L T G Y FL L S I V M I F S M K L A I Q Q I N S N S I Y Q Y L W S W T I N N D F S L E F G Y L M D P L T S I ML A A L V T A L S A L I S S L T A S - K T Y S Q P L W T W M S V G D F N I G F N L V L D G L S L T M. A . . . . . . S . . L . . . W L L D F G V F L D L S G Y F
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
110 120 130 140 150I L M N G L V T G A V L L Y C F D K Q K S P F F Y T Q L V I L H G A V N A T F C - - - - C A D L I SL L T N A L V T L A V L V Y C W N T G R S A F F Y A Q L I I L H A S L N S A F L - - - - C A D F M SI L T N A L V T I A V I L Y C W Q S D K T A F F Y V Q T M M L H G S V N A A F A - - - - C T D F I SS M L I T T V A I L V L I Y S D N Y M S H D Q G Y L R F F A Y M S F F N T S M L G L V T S S N L I QL S V V T G V G F L I H M Y A S W Y M R G E E G Y S R F F A Y T N L F I A S M V V L V L A D N L L L. L N . L V T . A V L . Y C . . . . F F Y . Q . L H . . N A . F . C D L I S
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
160 170 180 190 200L Y V A L E C I G I A A F L L I T Y S R S D R S L W V G L R Y L F I S N T A M L F - Y L I G A V L VL Y V A L E V V A I A A F C L M T Y P R E P R I I W L G L R Y L L L S N T A M L F - Y L I G V A L VL Y V A L E V S G I A A F L L I A Y P R T D R S I W V G L R Y L F I S N V A M L F - Y L V G A V L AI Y I F W E L V G M C S Y L L I G F W F T R P I A A N A C Q K A F V T N R V G D F G L L L G I L G LM Y L G W E G V G L C S Y L L I G F Y Y T D P K N G A A A M K A F V V T R V G D V F L R F A L F I LL Y V A L E . V G I A A F L L I . Y R T D R . W . G L R Y L F . S N A M L F Y L . G . . L .
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
210 220 230 240 250Y Q A S N S F A F S G L A V A P K E A I A - - - - - - - - - - - - - - L I F L G L L T K G G I F V SY K T N Q S F A F S G L T Q A P P E A I A - - - - - - - - - - - - - - L I F L G L L T K G G I F L AY Q T N H S F A F S S L R G A P P E A L A - - - - - - - - - - - - - - L I F L G L L V K G G V F V SY W I T G S F E F R D L F E I F N N L I K N N E V N S L F C I L C A F L L F A G A V A K S A Q F P LY N E L G T L N F R E M V E L A P A H L A D G - - - N N M L M W A T L M L L G G A V G K S A Q L P LY . S F A F S L A P P E A I A L I F L G L L . K G G . F . .
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
260 270 280 290 300G L W L P L T H G E S E T P V S A L L S G - V V V K A G V F P L A R C A - - - L L V P E L D P V V RG L W L P Q T H G E A A T P V S A M L S G - A V V K A G A L P L L R C A - - - L L S D Q L L L L V QG L W L P L T H S E S D T P V S A L L S G - V V V K T G V Y P L V R C A - - - L I L D E V D P V I RH V W L P D A M - E G P T P I S A L I H A A T M V A A G I F L V A R L L P L F V V I P Y I M Y V I SQ T W L A D A M - A G P T P V S A L I H A A T M V T A G V Y L I A R T H G L F L M T P E V L H L V GG L W L P T H E . T P V S A L L S G . V V K A G V . P L A R C A L . . P E . V V
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
310 320 330 340 350L F G V G T A L L G V G Y A V F E K D T K R M L A F H T V S Q L G F V L A A P A V G G - - - - - F YI L G V A T A L F G V V Y A M L A K D S K R M L A F H T V S Q M G F V L A A P I A G G - - - - - F YI L G A G T A L L G V S Y A I F E K D T K R M L A W S T I S Q L G W I M S A P E V A G - - - - - F YF I G I I T V L L G A T L A L A Q K D I K R S L A Y Y T M S Q L G Y M M L A L G M G S Y R T A L F HI V G A V T L L L A G F A A L V Q T D I K R V L A Y S T M S Q I G Y M F L A L G V Q A W D A A I F HI . G . . T A L L G V Y A . . K D . K R M L A . T . S Q L G . . . A P . V G G F Y
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
360 370 380 390 400A L T H G L V K G A L F L T A G Q L S - - - - - - - - - S R N - - - F K V L R E - - - - - Q S I P RA L S H G L V K S S L F L L A G N L P - - - - - - - - - S R D - - - F K V L K K - - - - - T P I A AA L T H G L V K S V L F L I A G S L P - - - - - - - - - S R N - - - F K E L K N - - - - - K P I N TL I T H A Y S K A L L F L A S G S L I H S M G T I V G Y S P D K S Q N M V L M G G L T K H V P I T KL M T H A F F K A L L F L A S G S V I L A C H - - - - H E Q N I F K M G G L R K S - - - - I P L V YA L T H G L V K . . L F L . A G S L S R N F K V L . P I
354
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
410 420 430 440 450A - - - - - - - - - - - - - - - - - - Y W - - - - - - - - - - - - - - - - - - W V L V L A - - - - -G - - - - - - - - - - - - - - - - - - F W - - - - - - - - - - - - - - - - - - V P L L L A - - - - -S - - - - - - - - - - - - - - - - - - V W - - - - - - - - - - - - - - - - - - I A L V I G - - - - -T S F L I G T L S L C G I P P L A C - F W S K D E I L N D S W V Y S P I F A I I A Y F T A G L T A FL C F L V G G A A L S A L P L V T A G F F S K D E - - - - - - - - - - - - - I L A G A M A N - - - -. F W . A L . . A
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
460 470 480 490 500- - - - - - - - - - - - - - - - - C A S I S G - - - - - L P F L A G Y S S K - - - - - - - - - - - -- - - - - - - - - - - - - - - - - S S S I A G - - - - - F P L L A G F E A K - - - - - - - - - - - -- - - - - - - - - - - - - - - - - S L S I S G - - - - - F P L F S G F G A K - - - - - - - - - - - -Y M F R I Y L L T F E G H L N F F C K N Y S G K K S S S F Y S I S L W G K K E L K T I N Q K I S L L- - - - - - - - - - - G H I N L M V A G L V G - - - - - A F M T S L Y T F R - - - - - - - - - - - M
. S I S G F P . S G . K
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
510 520 530 540 550- I L T M K N - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - I L P W Q S- T L T L K G - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - L P P W L A- L L T M K N - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - L L P W Q VN L L T M N N K E R A S F F S K K P Y E I N V K L T K L L R S F I T I T Y F E N K N I S L Y P Y E SI F I V F H G K E Q - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - I H A H A V K
. L T M K N L . P W
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
560 570 580 590 600F A L N V A A V G T A I S F A K F I - F L P K K T D P S - - - - L K I L P N F Y G A I V L L L G - -I A L N I A A V G T A I S F S K F V - F L K P T F V G - - - - - K T Y P L G L G L A L V V L L G - -I V M N V A A L G T A I T F A K F I - F L P H G G K - - - - - - R E V K Q G L W P G V I L L I S - -D N T M L F P L I I L I M F T L F V G F I G I P F N Q E G M D L D I L T K W L T P S I N L L H S N SG V T H S L P L I V L L I L S T F V G A L I V P P L Q G - - - V L P Q T T E L A H G S M L T L E I T. . N . A A L G T A I . F . K F V F L . L . . . L L L
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
610 620 630 640 650- - - - - - - - - - - - - - - - - - - - - - G L F V T N S F Y L E A Y Q - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - G L A V G N V V Y W Q A F T - - - - - - - - - - - - - -- - - - - - - - - - - - - - - - - - - - - - A L I V A N I V Y Y D A Y T - - - - - - - - - - - - - -E N F V D W Y E F V I N A I F S I S I A F F G I F I A F F F Y K P I Y S S L K N F D L I N S F D K RS - - - - - - - - - - - - - - - - - - - - - G V V A V V G I L L A A W L W L G - - - - - - - - - - -
G L . V . N . . Y A Y .
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
660 670 680 690 700- - P S N I L K A L V T I G I G W L A - - - - - - - - - - - - - - - - - - - - - - - - - Y G L I F Q- - P S N L I K A T L T C V V G A G L - - - - - - - - - - - - - - - - - - - - - - - - - Y W V V V K- - V D N I I K A L A I I G I G W L V - - - - - - - - - - - - - - - - - - - - - - - - - Y L F I V QG Q K R I L G D N I I T I I Y N W S A N R G Y I D A F Y S T F L I K G I R S L S E L - - V S F F D R- - K R T L V T S I A N S A P G R L L G T W W Y N A W G F D W L Y D K V F V K P F L G I A W L L K R
N L . K A . . T I . . G W L . Y . . .
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
710 720 730 740 750R I T V K L P R - - - - - - - - V V E Q F E H L V - G - - - - - - - V M S L V L T G L F W L V L A NR L T L K L P D - - - - - - - - E G E Q V D H L L - G - - - - - - - M M S I S L T I L F A W I L VR L T I K L P R - - - - - - - - V L E Q F E H L V - G - - - - - - - F M S L M L V L L F W M V L A NR I I D G I P N G F G V T S F F V G E G I K Y V G - G G R I S S Y L F W Y L L Y V S I F L F I F T FD P L N S M M N I P A V L S R F A G K G L L L S E N G Y L R W Y V A S M S I G A V V V L A L L M V LR . T . K L P V G E Q . . H L . G M S L . L V . L F . . L .
NdhF5-7002(slr2007)NdhF5-6803(slr2007)NdhF5-7120(slr2007)NdhF-Spinacia oleraceaNuoL-E. coli
760 770 780 790 800
TR
356
Figure B15. ClustalW alignment of NdhF amino acid sequences from Synechococcus sp.
PCC 7002 against each other. The Synechococcus sp. PCC 7002 NdhD amino acid
sequences were aligned using the ClustalW alignment program from MacVector v. 6.5.
The consensus amino acid sequence is underneath the alignments. Gray high-lighted
regions indicate identical amino acids while lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids determined with the
ClustalW program (Thompson et al., 1994b).
357
7002 NdhF ClustlW Amino Acid Alignments
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
10 20 30 40 50M E P L Y Q Y A WL I P V L P L L G A M V I G I G L I S L N K F T N K L R Q L Y A V F V L S L I G
M N E L T I G WV I - - - - - - - - - - - F P F V V G F S I Y L L P K I D R Y L A I F V S I C SM N T F F S Q S V WL V P C Y P L L G M G L S A L WM P S I T R K T G P R P A G Y V N M L L T F M AM S E F L L Q S V WL V P V Y G I T G A L L T L P WS L G L I R R T G P R P A A Y L N L I M T F L G
M I D D I T I I WI L - - - - - - - - - - - L P F V V G F S I Y L L P R WN R Y F A L A I A A L S. Q . WL . P . G . . P . . . G . . . . T . P R . Y . . . . . . . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
60 70 80 90 100T S M A L S - F G L L WS Q I Q G H E A F T Y T L E WA A A G D F H L Q M G Y T V D H L S A L M S VL I F G - - - F V Q I F Q P E P - - - - - - Y S L K L L G M Y G V D L L V D - D Q S G Y F I L T N AL V H S C L A F I E R WE Q P A L - - - - K P S L T WL Q A A D L T L S I D L D I S S I T I G A L IL L H G S F A F A S L WN M P P Q - - - - Q L S L E WL Q V A D L N L S L V I E I S P V N L G A M EV V Y S - - - I G L L WS L E P - - - - - - F T L E L L D S F G V T L M F D - E L S G Y F I L M N GL . . F . L W P . S L E WL . . D . L . D . . S . I L .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
110 120 130 140 150I V T T V A L L V M I Y T D G Y M A H D P G Y V R F Y A Y L S I F S S S M L G L V F S P N L V Q V YA V A I A - - - - - - - V T V Y C WK S A K S A F F F T Q L V V L Q G A L N A V F V C A D L I S L YL I A G I N L L A Q L Y A V A Y L E M D WG WA R F F A T M S L F E A G M C A L V L C N S L F F S YL V T G I C F M A Q L Y G L G Y L E K D WS I A R F Y G L M G F F E A A L S G L A I S D S L L L S YL V T G A - - - - - - - V L L Y C F D K Q K S P F F Y T Q L V I L H G A V N A T F C C A D L I S L YL V T G . . . Y . . . Y . . D A R F Y . L . . F . A . A L . . C L . . Y
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
160 170 180 190 200I F WE L V G M C S Y L L I G F WY D R K A A A D A C Q K A F V T N R V G D F G L L L G M L G L Y WV A L E A I S I A A F L L M T Y Q R T D R S I WI G L R Y L F L S N - T A M L F Y L I G A V L V Y QV V L E I L T L G T Y L L I G Y WF N Q S L V V T G A R D A F L T K R V G D L F L L M G V V A L L PG L L E V L T L S T Y L L V G F WY A Q P L V V T A A R D A F L T K R V G D I L L L M G I V A L S SV A L E C I G I A A F L L I T Y S R S D R S L WV G L R Y L F I S N - T A M L F Y L I G A V L V Y QV . L E . . . . . . Y L L I G Y W. . . . . G . R A F L T N R V G D L F L L . G . V . L Y
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
210 220 230 240 250A T G S F E F D L M G D R L M D L V S T G Q I S S L L A I V F A V L V F L G P V A K S A Q F P L H VA T K S F A F V G L - - - - - - - - A E A P S D A I A L I F L G L L T K G G - - - - - - V F V S G LL A G T WN F D G L A E - - - - WA A T A E L D P T L A T L L C L A L I A G P L G K C A Q F P L H LY G T G L T F S E L E T - - - - WA A N P P L P P WE A S L V G L A L I S G P I G K C A Q F P L N LA S N S F A F S G L - - - - - - - - A V A P K E A I A L I F L G L L T K G G - - - - - - I F V S G LA . S F F G L . A A P . . . . A I . L G L L . . G P . . K A Q F P L L
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
260 270 280 290 300WL P D A M - E G P T P I S A L I H A A T M V A A G V F L V A R M Y P V F E P I P E A M N V I A WTWL P L T H S E A E T P V S A M L - S G V V V K A G I F P L L R C G - - - I L V P D L D L WL R L FWL D E A M - E S P V P A T V V R - N S L V V G T G A WV L I K L Q P I F A L S D F A S T F M I A IWL D E A M - E G P N P A G I I R - N S V V V S A G A Y V L L K M E P V F T I T P I T S D A L I I IWL P L T H G E S E T P V S A L L - S G V V V K A G V F P L A R C A - - - L L V P E L D P V V R L FWL P . A M E . P T P . S A . . . V V V A G . F . L . R . P . F . L . P . . . . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
310 320 330 340 350G A T T A F L G A T I A L T Q N D I K K G L A Y S T M S Q L G Y M V M A M G I G G Y T A G L F H L MG L A T A L L G I I F A I L E T D A K R L L A F S T I S K L G L L L S A P - - - - - A V A G L A A LG A T T A L G A A M V A I A Q I D I K R S L S Y S V S A Y M G M V F M A V G S Q Q D Q T T L V L L LG T V T T V G A S L V A L A Q I D I K R A L S H S T S A Y L G L V F I A V G L N Q V D I A L L L L LG V G T A L L G V G Y A V F E K D T K R M L A F H T V S Q L G F V L A A P - - - - - A V G G F Y A LG . . T A L L G . . . A . . Q D I K R . L A . S T S L G V . A G . . L . . L L
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
360 370 380 390 400T H A Y F K A M L F L G S G S V I H G M E E V V G H N A V L A Q D M R L M G G L R K Y M P I T A T TS H G L V K S S L F L M A G - - - - - - - - - - - - - - - - - - - - - Q L P - - - T R - - - - - - -T Y G V A M A I L V M A I G G V V L - - - - - - - - - V N I S Q D L T Q Y G G L WS R R P I T G I CT H A I A K A L L F M S I G A V I L - - - - - - - - - N T H G Q N I T E M G G L WS R M P A T T S AT H G L V K G A L F L T A G - - - - - - - - - - - - - - - - - - - - - Q L S - - - S R - - - - - - -T H G . . K A . L F L . G V . Q Q G G L S R P . T
358
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
410 420 430 440 450F L I G T L A I C G I P P F A G F WS K D E - - I L G L A F E A N P V L WF I G WA T A G M T A F Y- - - - - - - - - - - - - - - N F Q E L R Q T K I A S S L WL P L A I A C L S M V G M P L L V G F SY L V G A A S L V A L P P F G G F WS L A Q - - L T T N F WK T S P I L A V I L I T V N A L T S F SF V V G S A G L V C L F P L G T F WT M R R - - WV D G F WD T P P WL V L L L V G V N F C S S F N- - - - - - - - - - - - - - - N F K V L R E Q S I P R A Y WWV L V L A C A S I S G L P F L A G Y S. . . G . . . P . F W. L R I . . . W P . L . . . . G . . L . . F S
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
460 470 480 490 500M F R M Y F L T F E G E F R G T D Q Q L Q E K L L T A A G Q A P E E G H H G S K P H E S P L T M T FS K A L L L K N I A - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - P - - - - - - - - -I M R E F G L I F G G K - - - - - - - - - - - - - - - - - - - - - - - - - - - - P K Q M T V R S P EL T R V F R S V F L G A - - - - - - - - - - - - - - - - - - - - - - - - - - - - P K P K T R R S P ES K I L T M K N I L - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - P - - - - - - - - -
R . . F . G P .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
510 520 530 540 550P L M A L A V P S V L I G L L G V P WG N R F E A F V F S P N E - - - - A A E A A E H G F E L T E F- - WQ A M G L N I A A V G T A L A - - - - F A K F L F I P - - - - - - - - - - - - - - H D A A T KG L WA L V L P M V I L A G F A L H S P F I L A K L N F L P - - - - - - - - D WH Q L N L P L A A VV V WQ M A V P M V S L I L M T L M V P F F L H Q WQ L L F N P S L P T L V E R P L I V T L A I P A- - WQ S F A L N V A A V G T A I S - - - - F A K F I F L P - - - - - - - - - - - - - - K K T D P S
. WQ . . . P V . . . G A L F A K F . F L P . . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
560 570 580 590 600L I M G G N S V G I A L I G I T I A S L M Y L Q Q R I D P A R L A E K F P V L Y Q L S L N K WY F DF T A K S T T F WG A I A F L F S G V I L G N - - - - - - - - - - - - - - - - - - - - - - G F Y L EL I I S T M V G G G T A M Y L Y L N E K I S K P I H I F S D P V R E F F - - - - - - - A K D L Y T AL M I T G G L G L V A G L T I T L N P S L S R P R Q L Y L R F L Q D L L - - - - - - - A Y D F Y I DL K I L P N - F Y G A I V L L L G G L F V T N - - - - - - - - - - - - - - - - - - - - - - S F Y L EL I . . G A . . L . . . . . . . . F Y . .
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
610 620 630 640 650D I Y N N V F V M G T R R L A R Q I L E V D Y R V V D G A V N L T G I A T L L S G E G L K Y I E N GA Y Q L D N I P K A L I K I A - - I G WA L Y WL I M K R I E F K - L P R I F - - - - - - - - - - -E L Y K N T V I F A V A L I S K I I D WL D R Y F V D G V I N F L G L A T L F G G Q S L K Y N N S GR I Y N V T V V WL V T T L S K L A A WF D R Y V V D G F V N L T G L A T L F S G S A L R Y N V S GA Y Q P S N I L K A L V T I G - - I G WL A Y G L I F Q R I T V K - L P R V V - - - - - - - - - - -
. Y . . A . . I . . I . W. D Y . . V D G I N . G L A T L F G L . Y G
NdhF1-7002(slr0844)NdhF2-7002(slr2009)NdhF3-7002(sll1732)NdhF4-7002(sll0026)NdhF5-7002(slr2007)
660 670 680 690 700R V Q F Y A L I V F G A V L G - - - - F V I F F S V A- E A F E Q L I G A M S V V L - - - - - T G L F WM V T L NQ S Q S Y A L S I V A G I L L - - - - F I A A L S Y P L L K H WQ FQ S Q F Y V L T I V L G M I L G L V WF M A T G Q WT M I T D F WS N Q L A- E Q F E H L V G V M S L V L - - - - - T G L F WL V L A N
Q F Y . L . . V . . . . L F . . F . .
359
Figure B16. ClustalW alignment of NdhC amino acid sequences. NdhC amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, E. coli, and Spinacia oleracea . The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
NdhC ClustalW Amino Acid Alignment
NdhC-7002NdhC-6803NdhC-7120NdhC-Spinacia oleraceaNuoA-E. coli
10 20 30 40 50V F V L S G Y E Y F L G F L I V S S L V P I L A L T A S K L L R P K G G G P E R K T T YM F V L T G Y E Y F L G F L F I C S L V P V L A L T A S K L L R P R D G G P E R Q T T YM F V L S G Y E Y L L G F L I I C S L V P A L A L S A S K L L R P T G N S L E R R T T YM F L L Y E Y D I F WA F L I I S S V I P I L A F L F S G I L A P I S K G P E K L S S Y
M S M S T S T E V I A H H WA F A I F L I V A I G L C C L M L V G G WF L G G R A R A R S K N V P FM F V L . G Y E Y F L G F L I I S L V P . L A L . A S K L L R P . . G P E R T T Y
NdhC-7002NdhC-6803NdhC-7120NdhC-Spinacia oleraceaNuoA-E. coli
60 70 80 90 100E S G M E P I G G A WI Q F N I R Y Y M F A L V F V V F D V E T V F L Y P WA V A F N Q L G L L A FE S G M E P I G G A WI Q F N I R Y Y M F A L V F V V F D V E T V F L Y P WA V A F N Q L G L L A FE S G M E P I G G A WI Q F N I R Y Y M F A L V F V V F D V E T V F L Y P WA V A F H R L G L L A FE S G I E P M G D A WL Q F R I R Y Y M F A L V F V V F D V E T V F L Y P WA M S F D I L G V S V FE S G I D S V G S A R L R L S A K F Y L V A M F F V I F D V E A L Y L F A WS T S I R E S G WV G FE S G M E P I G G A WI Q F N I R Y Y M F A L V F V V F D V E T V F L Y P WA V A F L G L L A F
NdhC-7002NdhC-6803NdhC-7120NdhC-Spinacia oleraceaNuoA-E. coli
110 120 130 140 150V E A L I F I A I L V I A L V Y A WR K G A L E WSV E A L I F I A I L V V A L V Y A WR K G A L E WSI E A L I F I A I L V V A L V Y A WR K G A L E WSI E A L I F V L I L I V G L V Y A WR K G A L E WSV E A A I F I F V L L A G L V Y L V R I G A L D WT P A R S R R E R M N P E T N S I A N R Q RV E A L I F I A I L V V A L V Y A WR K G A L E WS
360
Figure B17. ClustalW alignment of NdhK amino acid sequences. NdhK amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, E. coli, and Spinacia oleracea. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
361
NdhK ClustalW Amino Acid Alignment
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
10 20 30 40 50M N P T S F E Q Q Q -
M S P N P A N P T D L E R V A -M V L N S D L T T Q D -
MG N E F R Y I G C I R I Y R S F N F R A Y P N C W F S L C M A K R S I G MV L I P E D S D K K K KMD Y T L T R I D P N G E N D R -
. . P . D . E
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
60 70 80 90 100T E K L L N P M G R S Q V T Q D L S E N V I L T T V D D L Y N WA R L S S L W P L L Y G T A C C F IT A K I L N P A S R S Q V T Q D L S E N V I L T T V D D L Y N WA K L S S L W P L L Y G T A C C F IK E R I I N P I E R P T I T Q D L S E N V I L T T V D D L Y N WA R L S S L W P L L F G T A C C F II E T V MN S I E F P L L D Q I A Q N S V I S T T S N D L S N WS R L S S L W P L L Y G T S C C F IY P L Q K Q E I V T D P L E Q E V N K N V F MG K L N D MV N WG R K N S I W P Y N F G L S C C Y V
E . . . N P I R . T Q D L S E N V I L T T V D D L Y N WA R L S S L W P L L Y G T A C C F I
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
110 120 130 140 150E F A A L L G S R F D F D R F G - L V P R S S P R Q A D L I L V A G T V T MK M A P A L V R L Y E EE F A A L I G S R F D F D R F G - L V P R S S P R Q A D L I I T A G T I T MK M A P A L V R L Y E EE F A A L I G S R F D F D R F G - L I P R S S P R Q A D L I I T A G T I T MK M A P Q L V R L Y E QE F A S L I G S R F D F D R Y G - L V P R A S P R Q A D L I L T A G T V T MK M A P S L L R L Y E QE M V T L F T A V H D V A R F G A E V L R A S P R Q A D L M V V A G T C F T K M A P V I Q R L Y D QE F A A L I G S R F D F D R F G L V P R S S P R Q A D L I . T A G T . T MK M A P . L V R L Y E Q
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
160 170 180 190 200MP E P K Y V I A M G A C T I T G G M F S S D S T T A V R G V D K L I P V D L Y I P G C P P R P E AMP E P K Y V I A M G A C T I T G G M F S S D S T T A V R G V D K L I P V D V Y I P G C P P R P E AMP E P K Y V I A M G A C T I T G G M F S V D S P T A V R G V D K L I P V D V Y L P G C P P R P E AMP E P K Y V I A M G A C T I T G G M F S T D S Y S T V R G V D K L I P V D V Y L P G C P P K P E AML E P K W V I S M G A C A N S G G M Y - - D I Y S V V Q G V D K F I P V D V Y I P G C P P R P E AMP E P K Y V I A M G A C T I T G G M F S . D S T A V R G V D K L I P V D V Y I P G C P P R P E A
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
210 220 230 240 250I I D A I I K L R K K V S N E T I Q E R S L K T E Q T H R Y Y S T A H S M K V V E P I L T G K Y L GI F D A I I K L R K K V A N E S I Q E R - A I T Q Q T H R Y Y S T S H Q M K V V A P I L D G K Y L QI I D A MI K L R K K I A N D S M Q E R - S L I R Q T H R F Y S T T H N L K P V A E I L T G K Y M QV I D A I T K L R K K I S R E I Y E D R - I K S Q P K N R C F T I N H K F R V G R S I H T G N Y D QY M Q A L M L L Q E S I G - - - - K E R - - - - - - - - - - - - - - - - - R P L S W V V G - - - D QI I D A I I K L R K K I . N E . Q E R . . Q T H R . Y S T H K V V I L T G K Y Q
NdhK-7002NdhK-6803NdhK-7120NdhK Spinacia oleraceaNuoB-E. coli
260 270 280 290 300MD T W N N P P K E L T E A M G M P V P P A L L T A K Q R E E AQ G T R S A P P R E L Q E A M G M P V P P A L T T S Q Q K E Q L N R GS E T R F N P P K E L T E A I G L P V P P A L L T S Q T Q K E E Q K R GA L L Y K Y K S P S T S E I P P E T F F K Y K N A A S S R E L V NG V Y R A N MQ S E R E R K R G E - - R I A V T N L R T P D E I
. T R N P P . E L . E A G P V P P A L T . . . E E . .
362
Figure B18. ClustalW alignment of NdhJ amino acid sequences. NdhJ amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, E. coli, and Spinacia oleracea. The consensus amino
acid sequence is shown underneath. Gray high-lighted regions indicate identical amino
acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
NdhJ ClustalW Amino Acid Alignment
7002-NdhJ6803-NdhJ7120-NdhJNdhJ-Spinacia oleraceaNuoC-E. coli
10 20 30 40 50MA E E N Q N P - - - - - T P E E A A I V E A G A V S Q L L T E N G F S H E S L E R D H S G I E IMA E E V N S P N E A V N L Q E E T A I A P V G P V S T W L T T N G F E H Q S L T A D H L G V E M
M A D E E L Q P V - - - - - P A A E A A I V P S G P T S Q W L T E N G F A H E S L A A D K N G V E IM Q G R L S A W L V K H G L V H R S L G F D Y Q G I E T
MV N N M T D L T A Q E P A WQ T R D H L D D P V I G E L R N R F G P D A F T V Q A T R T G V P VM. E E . E A I . G . S W L T N G F H S L A D . G V E .
7002-NdhJ6803-NdhJ7120-NdhJNdhJ-Spinacia oleraceaNuoC-E. coli
60 70 80 90 100I K V D A D - - - - - - L L I P L C T A L Y A F G F N Y L - - - - Q C Q G A Y D L G P G K E L V S FV Q V E A D - - - - - - L L L P L C T A L Y A Y G F N Y L - - - - Q C Q G A Y D E G P G K S L V S FI K V E A D - - - - - - F L L P I A T A L Y A Y G F N Y L - - - - Q F Q G G V D L G P G Q D L V S VL Q I K P E - - - - - - D WH S I A V I L Y V Y G Y N Y L - - - - R S Q C A Y D V A P G G L L A S VV WI K R E Q L L E V G D F L K K L P K P Y V ML F D L H G MD E R L R T H R E G L P A A D F S V F. V . A D L L P . . T A L Y A Y G F N Y L Q Q G A Y D . G P G . L V S F
7002-NdhJ6803-NdhJ7120-NdhJNdhJ-Spinacia oleraceaNuoC-E. coli
110 120 130 140 150Y H L L K V G D N V T D P E E V R V K V F L P R E N P V V P S V Y WI WK G A D WQ E R E S Y D M YY H L V K L T E D T R N P E E V R L K V F L P R E N P V V P S V Y WI WK A A D WQ E R E C Y D M FY H L V K V S D N A D K P E E I R V K V F L P R E N P V V P S V Y WI WK T A D WQ E R E S Y D M FY H L T R I E Y G V D Q P E E V C I K V F A P R R N P R I P S V F WV WK S A D F Q E R E S Y D M FY H L I S I D R N R D - - - - I ML K V A L A E N D L H V P T F T K L F P N A N WY E R E T W D L FY H L . K . . N . D P E E V R . K V F L P R E N P V V P S V Y WI WK A D WQ E R E S Y D M F
7002-NdhJ6803-NdhJ7120-NdhJNdhJ-Spinacia oleraceaNuoC-E. coli
160 170 180 190 200G I V Y E G H P N L K R I L M P E D WI G WP L R K D Y V S P D F Y E L Q D A YG I V Y E G H P N L K R I L M P E D WV G WP L R K D Y I S P D F Y E L Q D A YG I I Y E G H P N L K R I L M P E D WV G WP L R K D Y I S P D F Y E L Q D A YG I S Y D N H P R L K R I L M P E S WI G WP L R K D Y I V P N F Y E I Q D A YG I T F D G H P N L R R I MM P Q T WK G H P L R K D Y P R A R Y R I L A VG I . Y E G H P N L K R I L M P E D W. G WP L R K D Y I S P D F Y E L Q D A Y
363
Figure B19. ClustalW alignment of NdhL amino acid sequences. ClustalW alignment
of NdhL amino acid sequences. NdhL amino acid sequences of the following organisms
were aligned using the ClustalW alignment program from MacVector v. 6.5:
Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803, Anabaena sp. PCC 7120, and
Synechococcus sp. PCC 7942. The consensus amino acid sequence is shown underneath.
Gray high-lighted regions indicate identical amino acids while lighter shading indicates
conserved amino acids. Dashes represent insertions/deletions included to maximize the
sequence similarity. The percent identity and percent conserved amino acids were
determined with the ClustalW alignment tool. (Thompson et al., 1994b).
NdhL ClustalW Amino Acid Alignment
7002-NdhL6803-NdhL6301-NdhL7120-NdhL
10 20 30 40MP F D I P V E T L L I A T L Y L S L S V T Y L L V L P A G L Y F Y L N N
ME D L L G L L L S E T G L L A I I Y L G L S L A Y L L V F P A L L Y WY L Q KMT V T L I I A A L Y L A L A G A Y L L V V P A A L Y L Y L Q K
MI V P L L Y L A L A G A Y L L V V P V A L L F Y L K LT . . . A . L Y L . L . A Y L L V . P A . L Y . Y L .
7002-NdhL6803-NdhL6301-NdhL7120-NdhL
50 60 70 80R WY V A S S I E R L V MY F F V F F L F P G ML L L S P F L N F R P R R R E VR WY V A S S V E R L V MY F L V F L F F P G L L V L S P V L N L R P R - R Q AR WY V A S S WE R A F MY F L V F F F F P G L L L L A P L L N F R P R S R Q IR WY V V S S I E R T F MY F L V F L F F P G L L V L S P F V N L R P R P R K IR WY V A S S . E R . MY F L V F F F P G L L . L S P L N R P R R .
7002-NdhL6803-NdhL6301-NdhL7120-NdhL
90 100 110 120
AP AE V
364
Figure B20. ClustalW alignment of NdbA amino acid sequences. sequences of the
following organisms were aligned using the ClustalW alignment program from
MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803, Calothrix
vigueri, Corneybacterium glutamicum, and Mycobacterium smegmatis. The consensus
amino acid sequence is shown underneath. Gray high-lighted regions indicate identical
amino acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. The percent identity
and percent conserved amino acids were determined with the ClustalW alignment tool.
(Thompson et al., 1994b).
365
NdbA ClustalW Amino Acid Alignment
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
10 20 30 40 50M V N T Q L P N L T P Q T D K H K V V I I G G G F G G L Y A A K T L G K Y E A - A V D V T L I D K R
M N S P T S P R R P H V V I V G G G F A G L Y T A K N L R R S P - - - V D I T L I D K RL Y T A K T L A T A N - - - V S V T L I D K RL Y A A K A L A K T N - - - V N V T L I D K R
M S R P R I V I V G A G F A G Y R T A R T L S R L T R H Q A D I T L L N P T. . . V I . G . G F . G L Y T A K T L . . V D V T L I D K R
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
60 70 80 90 100N F H L F Q P L L Y Q V A T G T L S P A D I A S P L R G V L S G N K N T H V L L D E V V D I D P D SN F H L F Q P L L Y Q V A T G S L S P A D I A S P L R G V L K G Q K N I R V L M D K V I D I D P D KN F H L F Q P L L Y Q V A T G T L S P G D I S S P L R A V F S K S K N T Q V L L G E V K D I N P K AN F H L F Q P L L Y Q V A T G T L S P A D I S A P L R S V L S K S K N T K V L L G E V N D I D P N LD Y F L Y L P L L P Q V A A G I L E P R R V S V S L S G T L P - - - H V R L V L G E A D G I D L D GN F H L F Q P L L Y Q V A T G T L S P A D I S S P L R G V L S K N T . V L L G E V D I D P D .
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
110 120 130 140 150K T V V M N E G I - - - - - V N Y D S L I V A T G V S H H Y F G N D H WK P Y A P G L K T V E D A LQ K V V L E D H A P - - - - I A Y D WL V V A T G V S H H Y F G N D H WA A L A P G L K T I E D A LQ Q V I L D D K V - - - - - V P Y D T L I V A T G A N H S Y F G K D H WK D V A P G L K T V E D A IQ Q I I V G D K V - - - - - V P Y D T L I V A T G A K H S Y F G K D N WQ E L A P G L K T V E D A IR T V H Y T G P E G G E G T L A Y D R L V L A A G S V N K L L P I P G V A E H A H G F R G L P E A LQ V . . D . . V Y D . L I V A T G . H . Y F G D H W . . A P G L K T V E D A L
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
160 170 180 190 200E I R H R I F M A F E A A E K E T D P A L Q Q A WL T F V I V G G G P T G V E L A G A I A E I A Y ST I R Q R I F A A F E A A E K E S N P E R Q Q A WL T F V I V G A G P T G V E L A G A I A E I A H SE M R R R I F G A F E A A E S E T D P E K R R A WL T F V I V G G G P T G V E L A G A I A E L A Y KE M R R R I F G A F E A A E K E T D L E K R K A L L T F V I V G G G P T G V E L A G A I A E L A Y KY L R D H V T R Q V E L A A A A D D R A E C A A R C T F V V V G A G Y T G T E V A A H G A M Y T D AE . R . R I F . A F E A A E K E T D P E . A WL T F V I V G G G P T G V E L A G A I A E . A Y
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
210 220 230 240 250V L K K D F R K I D T T R A R I I L L E G M D R V L P P Y D P S L S A K A Q K S L E N L G V Q V Q TS L K D N F H R I D T R Q A K I L L I E G V D R V L P P Y K P Q L S A R A Q R D L E D L G V T V L TT L K E D F R S I D T S E T K I L L L Q G G D R I L P H I A P E L S Q V A A E S L Q K L G A I I Q TT L Q E D F R N I N T S E T R I L L L Q G G D R I L P H I A P E L S Q A A T T S L R E F G V V V Q TQ V R R H P M R T G - M R P R WM L L D V A P R V M P E M D E R L S R T A E R V L R Q R G V D V R M. L K . D F R . I D T . R I L L L . G . D R V L P P L S A . S L L G V V Q T
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
260 270 280 290 300K S L V T N I E D H L V T F K Q G D D Q C E I A A K T I V WA A G V K A S G M S K V L E D R L S A TE R M V T D I N P E Q V T V H N N G Q T E T I V T K T V L WG A G V R A S S L G K I I G D R T G A EK T R V T N I E N D I V T F K K G D E V K E I P S K T I L WA A G V K A S P M G Q V L A E R T G V EK T R V T S I E N D I V T F K Q G D E L Q T I T S K T I L WA A G V K A S P M G K V L A E R T G V EG T S V K E A T H D G V V L T D G - - - S T V D T R T L V WC V G V R P D P - - - - L V E S L G L PK T V T I E D . V T F K . G D . T I . K T I L WA A G V K A S P M G K V L . E R T G . E
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
310 320 330 340 350L D R A G R V I V E P N L S V A G Y P D V F V I G D L A N F P H Q N - - E R P L P G V A P V A M Q EL D R A G R V V V N P D L S V A S F D N I F V L G D L A N Y S H Q G - - D Q P L P G V A P V A M Q EC D H A G R V I V E P D L T I R D Y K N I F V V G D L G N F S H Q N - - G K P L P G V A P V A T Q QC D R A G R A I V E P D L S I K G H Q N I F V V G D L A N F S H Q N - - G Q P L P S V A P V A I Q EM E R - G R L L V D P H L Q V P G R P E L F A C G D V A A V P D L N Q P G Q Y T P M T A Q H A WR H. D R A G R V I V E P D L S V G . N I F V . G D L A N F S H Q N G Q P L P G V A P V A Q E
366
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
360 370 380 390 400G E Y V A K L I K Q R V N G Q E - - M A P F R Y M E L G S L A V I G Q N A A V V D L G F V K F S G FA A Y L S K L I P A R L A E K E Q I M V P F R Y I D Y G S L A V I G Q N K A V V D L G F A Q F T G LG E Y V A K L I K K R L K G Q T - - L P Q F R Y N D V G S L A M I G Q N L A V V D L G L I K L Q G FG E Y V A K L I K K R L Q G K T - - L P A F K Y N D H G S L A M I G Q N A A V V D L G L L K L K G FG K V C A H N V V A S L G R G Q - - R R A Y R H R D M G F V V D L G G A K A A A N P L G L P M S G PG E Y V A K L I K R L G F R Y D G S L A I G Q N . A V V D L G . . K . G F
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
410 420 430 440 450L A WL I - - - - - - - - - - - - - - - - - - - WI F A H V Y - - - - - - - - - - - - - - Y L I E FV A WM I - - - - - - - - - - - - - - - - - - - WV WA H V Y - - - - - - - - - - - - - - Y L I E FI A WV F - - - - - - - - - - - - - - - - - - - WL L I H I Y - - - - - - - - - - - - - - F L I E FS A WA F - - - - - - - - - - - - - - - - - - - WL L I H I Y - - - - - - - - - - - - - - F L I E FA A G A V T R G Y H L A A M P G N R V R V A A D WL L D A V L P R Q A V Q L G L V R S WS V P L E S. A W. . WL L . H V Y . L I E F
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
460 470 480 490 500D N K M V V M L - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -D N K L I V M L - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -D T K L L V V F - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -D S K L L V M I - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -S S P E V A R V P G R P E Q S G K D A G T A G S A G T A G K H A D G D E G K K Q P G G E P A K G Q PD . K L . V M .
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
510 520 530 540 550- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Q WG WN Y F T R G - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Q WG WN Y F T R G - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Q WA WN Y I T R N - - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Q WA WN Y I T R K - - - - - -G G E P G K N Q P G G E P G K N Q P G G E P G K N Q P G G E P A K N Q P G G E P A E R G P D A S A G
Q WG WN Y . T R G
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
560 570 580 590 600- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R G A R L I T G E K D L V S - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R G A R L I T D T P N P Q S - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R R S R L I T G R E A F V E - - - - -- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R G S R L I T G K E S L A F - - - - -S R S H G E R A K R S Q S G S P R A S G K P R R A V E A A G P R S A R G G S G K P G K S S R A S G T
R G A R L I T G . . S
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
610 620 630 640 650- - - - - - - - - - - - - G F K L Y Q E A A D K E T K V N I K V E A- - - - - - - - - - - - - T S T V Q K E L V R P- - - - - - - - - - - - - P K T V N Q Q N- - - - - - - - - - - - - A N N F D D S N D T N N Y S A A N N R Q P L N VG S A G K R P T A P S G P S R S A G Q P A D P G P E P P A H Q P P P G P D I A P G P V R R T D G R A
. . Q .
NdbA-7002NdbA-6803Ndb(1)-7120Ndb(2)-7120Ndb1-E. coli
660 670 680 690 700
V E G D S
367
Figure B22. ClustalW alignment of NdbB amino acid sequences. ClustalW alignment
of NdbB amino acid sequences. NdbB amino acid sequences of the following organisms
were aligned using the ClustalW alignment program from MacVector v. 6.5:
Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803, Anabaena sp. PCC 7120, and
E. coli. The consensus amino acid sequence is shown underneath. Gray high-lighted
regions indicate identical amino acids while lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. The percent identity and percent conserved amino acids were determined with
the ClustalW alignment tool. (Thompson et al., 1994b).
368
NdbB ClustalW Amino Acid Alignment
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
10 20 30 40 50M S Q P H I V I I G G G F A G L Y T A L R L L Q F P WE T S Q R P D I T L I D R Q N H F V F S P
M T D A R P R I C I L G G G F G G L Y T A L R L G Q L S WE G H T P P E I V L V D Q R D R F L F A PM T E Q T K R I V I L G G G F G G L Y T A L R V S Q L P WE T Q Q K P E I V L V D Q S D R F L F S P
M S R P R I V I V G A G F A G Y R T A R T L S R L T R H Q A - - - D I T L L N P T D Y F L Y L PP R I V I . G G G F . G L Y T A L R L Q L WE P . I L . D D . F L F P
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
60 70 80 90 100L L Y E L I T E E M Q P WE V A P T Y T E L L R H G P V K F V Q T Q V Q T V D P E Q K N V V C G D RF L Y E L V T E E M Q T WE I A P P F V E L L A E S G V I F R Q A E V T A I D F D H Q K V L L N D QL L Y E L L T G E L Q S WE I A P P F I E L L E G T G I R F Y Q A V V S G I D I D Q Q R V H L Q D GL L P Q V A A G I L E P R R V S V S L S G T L P H - - V R L V L G E A D G I D L D G R T V H Y T G PL L Y E L . T E Q WE . A P . E L L V . F Q . V . I D D V D
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
110 120 130 140 150Q - - - - - I T Y D Y L V I A A G G T T K F V N L P G I K E Y A L P F K T L N D A L H L - K E K L RD K G T E S L A F D Q L V I A L G G Q T P L P N L P G L K D Y G L G F R T L E D A Y K L - K Q K L KP E - - - - I P Y D R L V L T L G G E T P L D L V P G A I S Y A Y P F R T I A D T Y R L - E E R L RE G G E G T L A Y D R L V L A A G S V N K L L P I P G V A E H A H G F R G L P E A L Y L R D H V T R
. Y D L V . A . G G T L . P G . . Y A F R T L D A . L . L R
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
160 170 180 190 200A L E T S V A E K I R - - - - - - - - I A I V G G G Y S G V E L A C K - - - - - - - - - - - L A D RS L E Q A D A E K I R - - - - - - - - I A I V G G G Y S G V E L A A K - - - - - - - - - - - L G D RV L E E S D A E K I R - - - - - - - - V A I V G A G Y S G V E L A C K - - - - - - - - - - - L A D RQ V E L A A A A D D R A E C A A R C T F V V V G A G Y T G T E V A A H G A M Y T D A Q V R R H P M R
L E A E K I R . A I V G . G Y S G V E L A K L . D R
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
210 220 230 240 250L G D R G R L R I I D R G D E I L K N A P K F N Q L A A K E A L E A R G I WV D Y A T E V T E V T AL G E R G R I R I I E R G K E I L A M S P E F N R Q Q A Q A S L S A K G I WV D T E T T V T A I T AL G E R G R F R L V E I S D Q I L R T S P D F N R E A A K K A L D A K G V F I D L E T K V E S I G QT G M R P R WM L L D V A P R V M P E M D E R L S R T A E R V L R Q R G V D V R M G T S V K E A T HL G . R G R R . . . . I L P . F N A . L A . G . . V D T V . T
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
260 270 280 290 300D S L S L R Y K G E V D T I P A D L V L WT G G T A I A P WV K D L A L P H A G N G K L D V N A Q LT D V T L Q F R E Q E D V I P V D L V L WT V G T T V S P L I R N L A L P H N D Q G Q L R T N A Q LN T I S L E Y K N Q V D T I P V D L V I WT V G T R V T N V V K S L P F K Q N Q R G Q I T N T P T LD G V V L T D G - - - S T V D T R T L V WC V G V R P D P L V E S L G L P M E - R G R L L V D P H L
. . L . . D T I P . D L V . WT V G T . P . V . L . L P G L L
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
310 320 330 340 350Q I Q N H P N I F A L G D V A Q A E D N - - - - - - L P M T A Q V A I Q Q A D V C A WN L R G L I TQ V E G K T N I F A L G D G A E G R D A S - - G Q L I P T T A Q G A F Q Q T D Y C A WN I WA N L TQ V L D H P D I F A L G D L A D C I D A E - - G Q Q V P A T A Q A A F Q Q A D Y A A WN I WA S L TQ V P G R P E L F A C G D V A A V P D L N Q P G Q Y T P M T A Q H A WR H G K V C A H N V V A S L GQ V . P I F A L G D . A . D . G Q . P T A Q . A . Q Q . D C A WN . A L T
369
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
360 370 380 390 400N K P L L P F K F F N L G E M L T L G E N N A T L S G L G L E L E G N L A H V A R R L V Y L Y R L PG R P L L P C R Y Q P L G E M L A L G T D G A V L S G L G I K L S G P A A L L A R R L V Y L Y R F PQ R P L L P F R Y Q Q L G E M M A L G T D N A T L T G L G V K L D G S L A Y V A R R L A Y L Y R L PR G Q R R A Y R H R D M G F V V D L G G A K A A A N P L G L P M S G P A A G A V T R G Y H L A A M P
. P L L P . R . L G E M . L G A L . G L G . L G . A . A R R L . Y L Y R P
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
410 420 430 440 450- - - - - - - - - - - - - - - T WE H Q V Q V G L N - - WL V - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - T WQ H Q L T V G L N - - WL T - - - - - - - - - - - - - - - - - - -- - - - - - - - - - - - - - - T L D H Q L K V G F N - - WL V - - - - - - - - - - - - - - - - - - -G N R V R V A A D WL L D A V L P R Q A V Q L G L V R S WS V P L E S S S P E V A R V P G R P E Q S
T H Q . V G L N WL V
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
460 470 480 490 500- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - Q P L T K L L A Q- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R P L G D WL K N E P S- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - R P I I E T I Y S A V D A V N E KG K D A G T A G S A G T A G K H A D G D E G K K Q P G G E P A K G Q P G G E P G K N Q P G G E P G K
P . . . . .
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
510 520 530 540 550
G T R YN Q P G G E P G K N Q P G G E P A K N Q P G G E P A E R G P D A S A G S R S H G E R A K R S Q S G S
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
560 570 580 590 600
P R A S G K P R R A V E A A G P R S A R G G S G K P G K S S R A S G T G S A G K R P T A P S G P S R
NdbB-7002NdbB-6803NdbB-7120Ndb-E. coli
610 620 630 640 650
S A G Q P A D P G P E P P A H Q P P P G P D I A P G P V R R T D G R A V E G D S
370
Figure B23. ClustalW alignment of HypE amino acid sequences. HypE amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Anabaena sp. PCC 7120,
Synechocystis sp. PCC 6803, and E. coli. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. The percent identity and percent conserved amino
acids were determined with the ClustalW alignment tool. (Thompson et al., 1994b).
371
HypE ClustalW Amino Acid Alignment
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
10 20 30 40 50M D L T C P L L Q N - - - - T S H V L L A G G G G K L M D L I D K M F T A F G E P C - -M K Y V D E F E P E - - - - - - K A E A L R E I E K L S Q L D K H I K M E V C G G H - -M N L V C P V L D R - - - - Y P Q V L L A G G G G K L S Q L L K Q I F P A F G A S E - -
M Q L I N S L F E A F A N P W A EM S G T V K L Y Q R P L N I S G R I D M G G A G G R A A Q L I Q E L F A A F D N E W R QM . . . . . . G . G G K L Q L I . F A F .
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
60 70 80 90 100- - - - - - - H D A A A L P S T E K L A F T D S Y V V Q - - - - L F F G G - - - D I K L- - - - - - - - - H S I F K G I E E I L P N I E L I H G G C P V C V M K G R L D D A A I- - - - - - G H D A A V F T N Q S S L A F T D S Y V I N - - - - L F F G G - - - D I S LQ E D Q A R - L D L A Q L V E G D R L A F T D S Y V I D - - - - L F F G G - - - N I K LG N D - - - - - Q A A F A M A G A R M V M T D A H V V S - - - - L F F G G - - - D I S L
D A A . . . . . L A F T D S Y V . L F F G G D I L
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
110 120 130 140 150A - - - - - I G T V N D L A A - G A R P L L S V - - - G I L E E G L - L E T L WQ V V SS Q N P N V I T T F G D T M V P G S K T T L Q A K A Q G D I R M V Y S L D S L - Q I R NA - - - - - V G T V N D L A A - G A T P R I S V - - - G I L E E G L - M E T L WR V Q SA - - - - - I G T A N D V A S - G A I P R L S C - - - G I L E E G L - M E T L K A V T SS - - - - - V G T I N D V A A - G A K P L L A A - - - S I L E E G F - L A D L K R I E SA I G T . N D . A A G A . P L S . G I L E E G L L E T L V S
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
160 170 180 190 200M Q Q - - - A Q V V G V Q I T G D T K V - E R G K G D G - - F I N T S - I G I I E S L NH P D K E I V F A L G F E T A P S T A L T L Q A A S E N T N F S M F S H V L V I P A Q AL G Q - - - A Q N C G V E I T G D T K V - D R G K G D G - - F I N T S - I G S L D H Q TM A E - - - T R A A G I A I T G D T K V - Q R G A V D K - - F I N T A - M G A I P A I HM A G - - - A R E A G V P I T G D T K V - E Q G K G D G - - F I T T T - V G V V P A I LM . A . . G V I T G D T K V . R G K G D G F I N T S . G . I P A .
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
210 220 230 240 250I Q P K A I A G D T I L V S - - - - D L G H G I A I M A R - - E G L A E S P I E S D - AL L N N P D L L D G F I G P H V S M V I G E P Y E F I A Q Y H K P I V S G F E P L D F QI H P N Q V Q G D R L I L S - - - - D L G H G M A I M A R - - Q G L E E T T I E S D - AWG A Q T L T G D V L L V S - - - - T L G H G A T I L N R - - E Q L G D G E L V S D - AI D G A G A R G D A I L L S - - - - T M G H G V A I L S R - - E S L E D T E I R S D - AI . . G D . . L . S L G H G . A I . A R E L . . . I S D A
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
260 270 280 290 300P L V E P V Q L L K A G I E H - C L - R D T R G G - - L A V L N E L A A A G Y Q F K H -S I WM L L Q L V E N R C E E N Q Y N R L Q K G G N Q I L A A M H K V A V R E K F A R GP V H R E V Q L L S A G I P H - C L - R D T R G G - - L S A V N E I A T S G V T M A R -V L T P L I - T L R D I P G K - A L - R D T R G G - - V A V V H E F A A C G C G I E S -A L H D L V A M L A V V P G R - V L - R D T R G G - - L T T L N E I S Q S G V G M V D -
L L V Q L L . . . L R D T R G G L . . . N E . A A G . .
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
310 320 330 340 350G N Q I P V I A V R G A C E F G F D - P V I A N E G R F A I L P E A R A E G L A - L Q HL D E I P - D G L K I R E E A Q F D A E L F T I P N L K A D H K A C K G E I L K G V K PE T L I P V E E V Q A A C E L G F D - P L V A N E G R F A I V P P E A Q K T V E - I Q TE A A L P V K A V R G V C E L G L D - A L F A N E G K L I A V E R N A E Q V L A - A H SE A A I P V L Q V D A A C E L G L D - P L V A N E G K L A I C A A A D D A L L A - A R GE . I P V . V . . A C E L G F D P L . A N E G . A I . . L A . .
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
360 370 380 390 400Y N A - - - - - - - N A R Q G L V T T Q N G E Q H L A I P V V V E N D G V T R I L E L SWQ C K V F G A C T P E T P G T C M V S - - - - - - S E A C A A Y Y K G R F S T T L K QF H P - - - - - - - Q A T A G T V T - - - G K S A Q T L L V S L E S S G A P R L L D I SH P L G K - - - - - D A A L G E V V E - - - - - - - - R G V R L A G L G V K R T L D P HH P L G R - - - - - E A R R G E V I E D - - - - - - G R F V Q M R T K G G M R V V D L S. . A G V . V . G . R . L D . S
HypE-7002HypE-7120HypE-6803HypE-E. coliHypE-R. eutropha
410 420 430 440 450G E Q L P R IA A E K P K V I S SG E Q L P R IA E P L P R IG E Q L P R IG E Q L P R I
372
Figure B24. ClustalW alignment of HoxE amino acid sequences. HoxE amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 6301, Anabaena sp. PCC 7120 and E. coli. The consensus
amino acid sequence is shown underneath. Gray high-lighted regions indicate identical
amino acids while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. Percent identity and
percent conserved amino acids determined by the ClustalW search program (Thompson
et al., 1994b).
HoxE ClustalW Amino Acid Alignment
HoxE-7002HoxE-6803HoxE-6301HoxE-7120NuoE-E. coli
10 20 30 40 50L S F E T I T M A I S H A V P - A D K R F R V L E V A M K R N Q Y R Q D T L I E I L H K A Q E V
M T V A T D R Q T V P P S A A H P S G D K R F K V L D A T M K R N Q F N Q D A L I E I L H K A Q E IM A T S E T T P S V D P R R R R L E L A I K R Q A A Q A D A L I E I L H E A Q S L
M T T T T P P H P H P S G D K R L K M L D A A I K R H Q Y Q Q D A L I E I L H K A Q E LM H E N Q Q P Q T E A F E L S A A E R E A I E H E M H H Y E D P R A A S I E A L K I V Q K Q
T S A P S . D K R . . L E . A M K R . Q . . Q D A L I E I L H K A Q E .
HoxE-7002HoxE-6803HoxE-6301HoxE-7120NuoE-E. coli
60 70 80 90 100F G Y L E D E V L E Y V A R G L K L P L S R V Y G V A T F Y H L F S L K P K G K H T C V V C L G T AF G Y L E E D V L L Y V A R G L K L P L S R V F G V A T F Y H L F S L K P S G K H T C V V C L G T AY G Y L D R E L L Q WV A E Q L A L P R S K V Y G V A S F Y H L F Q L N P S G R H R C H V C L G T AF G Y L E N D L L L Y I A H S L K L P P S R V Y G V A T F Y H L F S L A P Q G V H S C V V C T G T AR G WV P D G A I H A I A D V L G I P A S D V E G V A T F Y S Q I F R Q P V G R H V I R Y C D S V VF G Y L E . . . L Y V A . . L K L P . S R V Y G V A T F Y H L F S L P G . H . C V V C L G T A
HoxE-7002HoxE-6803HoxE-6301HoxE-7120NuoE-E. coli
110 120 130 140 150C Y V K G S Q E L L D K I D E T L H I K P G E T T P D D Q I S L V T A R C I G A C G I A P A V V Y DC Y V K G A G D L L K T L D Q E V H L K P G E T T E D G Q M S L V T A R C I G A C G I A P A V V Y DC Y V K G S Q A I L D C L I A E L G I R E G E T T N D G S V S L G T V R C V G A C G I A P V V V Y DC Y V K G S S A I L A D L E K A T R I H A G E T T A D G Q L S L L T A R C L G A C G I A P A V V F DC H I N G Y Q G I Q A A L E K K L N I K P G Q T T F D G R F T L L P T C C L G N C D K G P N M M I DC Y V K G S Q . I L L . L . I K P G E T T D G Q . S L . T A R C . G A C G I A P A V V Y D
HoxE-7002HoxE-6803HoxE-6301HoxE-7120NuoE-E. coli
160 170 180 190 200D E V C G K Q N A D H L M A R L R Q L S E G SG K V L G K Q N D E A V L A A I Q P WL S N SG D I Q G R Q E S E A V WQ Q V Q A WQ Q E A HG K V L G N Q T P E S V N E R V Q G WLE D T H A H L T P E A I P E L L E R Y KG . V G . Q E A V . Q W
373
Figure B25. ClustalW alignment of HoxF amino acid sequences. HoxF amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, Synechococcus sp. PCC 6301, Anabaena variabilis, E.
coli, and R. eutropha. The consensus amino acid sequence is shown underneath. Gray
high-lighted regions indicate identical amino acids while lighter shading indicates
conserved amino acids. Dashes represent insertions/deletions included to maximize the
sequence similarity. Percent identity and percent conserved amino acids determined by
the ClustalW search program (Thompson et al., 1994).
374
HoxF ClustalW Amino Acid Alignment
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
10 20 30 40 50M T D L A E L F E I A E A E Q D S H - - - - - - - - - - - - - - - - - - - -
M D I K E L K E I A T K S R E K Q - - - - - - - - - - - - - - - - - - - -M E L T E L L D I G R Q E R S Q Q - - - - - - - - - - - - - - - - - - - -M D WE D L G R L A N E E L T C Q - - - - - - - - - - - - - - - - - - - -M E L N E L L D I G R Q E R S Q Q - - - - - - - - - - - - - - - - - - - -
M M T V A A E I R - - - - - - - - - - - - - - - - - - - - - - -M S R I T T I L E R Y R S D R T R L I D I L WD V Q H E Y G H I P D A V L P Q L G A G L K L S P L
M D . E L . . I A E R Q
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
60 70 80 90 100- - - - - - - - - - - - - T K - - - I Q I R C C T A A G C M S S G S L A V K E E L E K Q I K E K N- - - - - - - - - - - - - T K - - - I R I R C C S A A G C L S S E G E T V K K N L T T A I A A A G- - - - - - - - - - - - - K P - - - V Q I R C C T A A G C L S A N S Q A V Q Q Q L E Q A V K A E G- - - - - - - - - - - - - K P - - - I R L R C C T A T G C R A N G A E A V F K A V Q Q T I A D Q N- - - - - - - - - - - - - K P - - - V Q I R C C T A A G C L S A N S Q A V K Q Q L E E A V K A E G- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - G S N P E K L L A P V L S A F WD E -D R E T A S F Y H F F L D K P S G K Y R I Y L C N S V I A K I N G Y Q A V R E A L E R E T G I R F
K P . I R C C T A A G C S A V . L E A . .
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
110 120 130 140 150L D R - - - - L E V V P V G C M K L C G F A P L V D V S D E T - - - - - - - C F Q Q V M P E V A PL E K - - - - V E V C G V G C M K F C G R G P L V A V D D R N Q - - - - - - L Y E F V T P D Q V GL G E - - - - V Q V S G V G C M R L C C Q G P L V E V E G S G E E K T K Q R L Y E K V T P E D A SL D R - - - - C E A V S V G C L G L C G A G P L V Q C D P S D R - - - - - - L Y S D I R P D Q A AL D G - - - - V Q V A G V G C M R L C C Q G P L V E V E G S G E E E T T Q K L Y G K V R S E D A S- - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - -G T D P N G M F G L F D T P C I G L S D Q E P A M L I D K - - - - - - - - V V F T R L R P G K I TL . . V V G C M . L C G P L V V . L Y V P . A
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
160 170 180 190 200E V D V A L G - - E T P S D K L E I C D R Q A P F F T L Q K P V V L E N S G K I D P E R I E A Y ID V K K L Q K P D A V A E T G L I S G D P H H P F Y A L Q R N I A L E N S G R I D P E S I D E Y IA I G A L K G - - - - K E A Q L S V V N L E Q P F F T Y Q A P I V L E N S G K I D P E R I Q A Y ID V A A A Q G - - - - A A M D L P E V D Q A Q P F F S Q Q L K I V N R H S G L I N P D R L E S Y LV V G T L R G - - - - K A A Q L S V V D L K Q P F F T Y Q A P I V L E N S G K I D P E R I Q A Y I- - - - - - - - - - - - - - - - - - - D - - - - - - - - - - - - - - - - - - - - E S WT L D V Y RD I A Q L K Q - - - - - - - G R S P A E I A N P A G L P S Q D I A Y V D A M V E S N V R T K G P V. V . L G L . D . P F F . Q I V L E N S G . I D P E R I . . Y I
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
210 220 230 240 250A G G Y R S L H Q V L E - - - D L T P M A V V E E I T Q S G L R G R G G G G Y P T G L K WA T V AA G G Y E Q L H K V V Y - - - E M T P E E V I V E M N K S G L R G R G G G G Y P T G L K WA T V AA Q G Y Q G L Y Q V L R - - - E M T P A A V V D S V S R S G L R G R G G A G Y P T G L K WA T V AA G G Y R A L M H T I F - - - D L T P T E V V E I I R L S G L R G R G G G G Y P T G L K WA T V AA Q G Y Q A L Y Q V L R - - - E M T P A G V V D S V N R S G L R G R G G A G Y P T G L K WA T V AR E G Y E G L R K A L - - - - A M A P D D L I A Y V K E S G L R G R G G A G F P T G M K WQ F I PF R G R T D L R S L L D Q C L L L K P E Q V I E T I V D S R L R G R G G A G F S T G L K WR L C RA G Y . L . V L . M T P V V . . S G L R G R G G A G Y P T G L K WA T V A
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
260 270 280 290 300K P G D Q K Y V V C N A D E G D P G A F M D R A V L E S D P H R V L E G M A I A G Y A V G A N H GK P G Q Q K Y V I C N A D E G D P G A F M D R S V L E S D P H R I L E G M A I A A Y A V G A N H GK K G E R K F V I C N A D E G D P G A F M D R S V L E S D P H R V L E G M A I A A Y A V G A S Q GK P S D R K F V V C N G D E G D P G A F M D R S V L E S D P H Q V I E G M A I A A Y A V G A N F GK K G E R K F V I C N A D E G D P G A F M D R S V L E S D P H R V L E G M A I A A Y A V G A S Q GQ D G K P H Y L V V N A D E S E P G T C K D I P L L F A N P H S L I E G I V I A C Y A I R S S H AD E S E Q K Y V I C N A D E G E P G T F K D R V L L T R A P K K V F V G M V I A A Y A I G C R K GK G . K Y V I C N A D E G D P G A F M D R S V L E S D P H R V L E G M A I A A Y A V G A . G
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
310 320 330 340 350Y Y I R A E Y P L A I Q R L E K A I K Q A K S K G L L G S Q I F N - S P F N F T I D I R I G A G AY Y V R A E Y P L A I Q R L Q K A I Q Q A K R Y G L M G T Q I F D - S P I D F K I D I R V G A G AY Y V R A E Y P I A I K R L H T A I H Q A Q R L G L L G S N I F E - S P F D F K I D I R I G A G AY Y V R A E Y P L A I A R L N Q A I R Q A R R R G L L G N S V L D - S R F S F D L E V R I G A G AY Y V R A E Y P I A I K R L Q T A I H Q A Q R L G L L G S N I F E - S P F D F K I D I R I G A G AF Y L R G E V V P V L R R L H E A V R E A Y A A G F L G E N I L G - S G L D L T L T V H A G A G AI Y L R G E Y F Y L K D Y L E R Q L Q E L R E D G L L G R A I G G R A G F D F D I R I Q M G A G AY Y V R A E Y P . A I R L A I . Q A . R G L L G . . I F . S P F D F I D I R I G A G A
375
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
360 370 380 390 400F V C G E E T A L I A S I E G G R G T P R P R P P Y P A Q S G L WG H P T L I N N V E T Y A N I V PF V C G E E T A L I A S V E G K R G T P R P R P P Y P A Q S G L WQ S P T L I N N V E T Y A N V V PY V C G E E T A L M A S I E G K R G V P H P R P P Y P A E S G L WG Y P T L I N N V E T F A N I A PF V C G E E T A L I H S I Q G E R G V P R V R P P Y P A E S G L WG H P T L I N N V E T F A N I A PY V C G E E T A L M A S I E G K R G V P H P R P P Y P A E S G L WG Y P T L I N N V E T F A N I A PY I C G E E T A L L D S L E G R R G Q P R L R P P F P A V A G L Y A C P T V V N N V E S I A S V P AY I C G D E S A L I E S C E G K R G T P R V K P P F P V Q Q G Y L G K P T S V N N V E T F A A V S RY V C G E E T A L I A S I E G K R G P R P R P P Y P A S G L WG P T L I N N V E T F A N I . P
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
410 420 430 440 450I I R E G A D WF S S I G T E K S K G T K V F A L T G K V K N N G L I E V P M G T P V R Q I V E Q MI I R E G G D WY G S I G T E K S K G T K V F A L T G K V E N A G L I E V P M G T T V R Q V V E E MI I R K G A D WF A S I G T A K S K G T K V F A L A G K I R N T G L I E V P M G T S L R Q I V A E MI V E Q G A D WF A A I G T P T S K G T K V F A L T G K L R N N G L I E V P M G I P L R S I V D G MI I R K G A D WF A S I G T A K S K G T K V F A L A G K I R N T G L I E V P M G T S L R Q I V E Q MI L N K G K D WF R S M G S E K S P G F T L Y S L S G H V A G P G Q Y E A P L G I T L R Q L L D M SI M E E G A D WF R A M G T P D S A G T R L L S V A G D C S K P G I Y E V E WG V T L N E V L A M VI I R G A D WF . S I G T K S K G T K V F A L . G K . . N G L I E V P M G T . L R Q I V . M
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
460 470 480 490 500G G G I P D S G T V K S V Q T G G P S G G C I P A D Y L D T P I E Y D S L I K L G T M M G S G G M IG G G V P N G G Q V K A V Q T G G P S G G C I P A D K L D T P I E Y D T L L A L G T M M G S G G M IG G G I P D G G V A K A V Q T G G P S G G C I P A S A F D T P V D Y E S L T N L G S M M G S G G M IG - - I P E S - P V K A V Q T G G P S G G C I P L A Q L D T P V D Y D S L I Q L G S M M G S G G M VG G G I P D G G V A K A V Q T G G P S G G C I P A S A F D T P V D Y E S L T N L G S M M G S G G M IG G - M R P G H R L K F WT P G G S S T P M F T D E H L D V P L D Y E G V G A A G S M L G T K A L QG - - - - - A R D A R A V Q I S G P S G E C V S V A K - - - - - D G E R K L A Y E D L S C N G A F TG G G I P . G G . K A V Q T G G P S G G C I P A L D T P . D Y E S L . L G S M M G S G G M I
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
510 520 530 540 550V M D E A T N M V D V A K F Y M E F C Q C E S C G K C I P C R A G T V Q M S G L L S K M L K G Q A EV M D E S T N M V D V A Q F Y M D F C K S E S C G K C I P C R A G T V Q L Y D L L T R F L E G E A TV M D D T T N M V D V A R F F M E F C M D E S C G K C I P C R V G T V Q L H G L L S K I R E G K A SV M D E N T D M V A I A R F Y M E F C R S E S C G K C I P C R A G T V Q L H E L L G K L S S G Q G TV M D D T T N M V D V A R F F M E F C M D E S C G K C I P C R V G T V Q L H G L L S K I R E G K A SC F D E T T C V V R A V T R WT E F Y A H E S C G K C T P C R E G T Y WL V Q L L R D I E A G K G QI F N C K R D L L E I V R D H M Q F F V E E S C G I C V P C R A G N V D L H R K V E WV I A G K A CV M D E . T N M V D V A R F . M E F C E S C G K C I P C R A G T V Q L H L L . K . G K A .
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
560 570 580 590 600P K D I E L L E Q L C H M V K E A S L C G L G Q S A P N P I L S T L R Y F R A E Y D A L V G S N A DQ E D L I K L E N L C H M V K E T S L C G L G M S A P N P V I S T L R Y F R H E Y E E L L K VL A D L E L L E E L C D M V K N T S L C G L G Q S A P N P V F S T L H Y F R D E Y L A L I A V G A VA I D L Q Q L E D L C Y L V K D T S L C G L G M S A P N P I L S T L R WF R Q E Y E S R L I P E R AF A D L E L L E E L C D M V K N T S L C G L G Q S A P N P V F S T L R Y F R D E Y L A L I A EM S D L D K L N D I A D N I N G K S F C A L G D G A A S P I F S S L K Y F R E E Y E E H I T G R G CQ K D L D D M V S WG A L V R R T S R C G L G A T S P K P I L T T L E K F P E I Y Q N K L V R H E G
D L . L E . L C M V K T S L C G L G S A P N P I . S T L R Y F R . E Y . L . .
HoxF-7002HoxF-6803HoxF-7120HoxF-6301HoxF-A. variabilisNuoF-E. coliHoxF-Ralstonia eutropha
610 620 630 640 650
S L N S T DI A L T H
P F D P A K S T A WA D R T E V K AP L L P S F D L D T A L G G Y E K A L K D L E E V T R
376
Figure B26. ClustalW alignment of HoxU amino acid sequences. The HoxU amino acid
sequence from Synechococcus sp. PCC 7002 was aligned with HoxU proteins from the
following organisms using the ClustalW alignment program from MacVector v. 6.5:
Synechococcus sp. PCC 7002, Synechocystis sp. PCC 6803, Synechococcus sp. PCC
6301, Anabaena variabilis, the N-terminus of the NuoG protein from E. coli and R.
eutropha. The consensus amino acid sequence is shown underneath. Gray high-lighted
regions indicate identical amino acids while lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids determined by the
ClustalW search program (Thompson et al., 1994).
377
HoxU ClustalW Amino Acid Alignment
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
10 20 30 40 50M A V K T - - - - - - - - - - - - - - - - - - L T I N D Q L I S A R A G E T I L E A A R D A G I H IM S V V T - - - - - - - - - - - - - - - - - - L T I D D K A I A I E E G A S I L Q A A K E A G V P IM S V V T - - - - - - - - - - - - - - - - - - L Q I D D Q E L A A N V G Q T V L Q V A R E A S I P IM S V K T - - - - - - - - - - - - - - - - - - L T I N D Q L I S A Q E E E T L L Q A A Q E A G I H IM S V K T - - - - - - - - - - - - - - - - - - L T I N D Q L I S A Q E E E T L L Q A A Q E A G I H IM T V T T S T P S G G G A A A V P P E D L V T L T I D G A E I S V P K G T L V I R A A E Q L G I E IM S I Q - - - - - - - - - - - - - - - - - - - I T I D G K T L T T E E G R T L V D V A A E N G V Y IM S V T L T I D D Q . I S A E G T . L Q A A E A G I I
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
60 70 80 90 100P T L C H L E G V S D V G A C R L C L V E I E G S N K L Q P A C V T E V M E G M V V Q T H - - T E KP T L C H L E G I S E A A A C R L C M V E V E G T N K L M P A C V T A V S E E M V V H T N - - T E KP T L C H L Q G V S D V G A C R L C V V E V A G S P K L Q P A C L L T V S E G L V V Q T R - - S P RP T L C H L E G V G D V G A C R L C L V E I T G S N K L L P A C V T K V A E G M E V R T N - - S D RP T L C H L E G V G D V G A C R L C L V E V A G S N K L L P A C V T K V A E G M E V S T N - - S D RP R F C D H P L L D P A G A C R Q C I V E V E G Q R K P M A S C T I T C T D G M V V K T Q L T S P VP T L C Y L K D K P C L G T C R V C S V K V N G N - - V A A A C T V R V S K G L N V E V N - - D P EP T L C H L E G V D V G A C R L C . V E V G S N K L P A C V T V . E G M V V T N S . .
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
110 120 130 140 150L E E Y R R M T V E L L F A E G N H V C A V C V A N G N C E L Q D M A V E V G M D H S R F P - Y Q YL Q N Y R R M T V E L L F S E G N H V C A I C V A N G N C E L Q D M A I T V G M D H S R F K - Y Q FL E R Y R R Q I V E L F F A E G N H V C A I C V A N G N C E L Q D A A I A V G M D H S R Y P - Y R FL Q K Y R R T I V E M L F A E G N H I C S V C V A N N N C E L Q D L A I E M G M D H V R L E - Y H FL Q R Y R R T I V E M L F A E G N H I C S V C V A N N N C E L Q D L A I E M G M D H V R L E - Y H FA E K A Q H G V M E L L L I N H P L D C P V C D K G G E C P L Q N Q A M S H G Q S D S R F E G K K RL V D M R K A L V E F L F A E G N H N C P S C E K S G R C Q L Q A V G Y E V D M M V S R F P - Y R FL . Y R R . V E L L F A E G N H . C V C V A N G N C E L Q D . A I E V G M D H S R F Y . F
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
160 170 180 190 200P K R E V D I S H K Q F G I D H N R C I L C T R C V R - V C D E I E G A H V WD V S N R G G E S K IP K R E V D L S H P M F G I D H N R C I L C T R C V R - V C D E I E G A H V WD V A Y R G A E C K IP K R D V D L S H R F F G L D H N R C I L C T R C V R - V C D E I E G A H V WD V A M R G E H C R IP N R K V D I S H D R F G V D H N R C V L C T R C I R - V C D E I E G A H T WD M A G R G T N S H VP N R K V D I S H D R F G V D H N R C V L C T R C I R - V C D E I E G A H T WD M A G R G T N S H VT Y E K P V P I S T Q V L L D R E R C V L C A R C T R - F S N Q V A G D P M I E L I E R G A L Q Q VP V R V V D H A S E K I WL E R D R C I F C Q R C V E F I R D K A S G R K I F S I S H R G P E S - -P R V D . S H F G . D H N R C I L C T R C V R V C D E I E G A H . WD . A R G S . .
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
210 220 230 240 250V S G L N Q P WG - - - - - - A V D A C T S - - - - - - - - - - - - - - - - - - - - - - - C G K C VV S G L N Q P WG - - - - - - T V D A C T S - - - - - - - - - - - - - - - - - - - - - - - C G K C VV A G M D Q P WG - - - - - - A V D A C T N - - - - - - - - - - - - - - - - - - - - - - - C G K C II T D L S Q P WG - - - - - - T S D T C T S - - - - - - - - - - - - - - - - - - - - - - - C G K C VI T D L S Q P WG - - - - - - T S D T C T S - - - - - - - - - - - - - - - - - - - - - - - C G K C VG T G E G D P F E S Y F S G N T I Q I C P V G A L T S A A Y R F R S R P F D L I S S P S V C E H C SR I E I D A E L A - - - - - - N A M P P E Q - - - - - - - - - - - - - - - - - - - - - - - V K E A V. . G L Q P WG T . D . C T S C G K C V
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
260 270 280 290 300D A C P T G S I F R K G - - - - - - - - - - - A T V G S K L G D R Q K L E F L I T A R - - - - - - -D A C P T G S I F H K G - - - - - - - - - - - E T T A E K I G D R R K V E F L A T A R - - - - - - -D A C P T G A L F H K G - - - - - - - - - - - E T T G E I E R D R D K L A F L A E A R - - - - - - -N A C P T G A I F Y Q G - - - - - - - - - - - S S V G E M K R D R A K L D F L V T A R - - - - - - -N A C P T G A I F Y Q G - - - - - - - - - - - S S V G E M K R D R A K L D F L V T A R - - - - - - -G G C A T R T D H R R G K V M R R L A A N E P E V N E E WI C D K G R F G F R Y A Q Q R D R L T T PA I C P V G T I L E K R - - - - - - - - - - - V G Y D D P I G R R K Y E I Q S V R A R - - - - - - -
A C P T G . I F . K G . G E . D R K L . F L . T A R
HoxU-7002HoxU-6803HoxU-6301HoxU-7120HoxU-A. variabilisNuoG-E. coli (371 N-TERM)HoxU-Ralstonia eutropha
310 320 330 340 350- - - - - - - - - T K G E WT R- - - - - - - - - K E K E WV R- - - - - - - - - G Q R R WT R- - - - - - - - - E K Q Q WN L- - - - - - - - - E K Q Q WN LL V R N A E G E L E P A S WP E- - - - - - - - - A L E G E D K
W .
378
Figure B27. ClustalW alignment of HoxY amino acid sequences. HoxY amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 6301, and R. eutropha. The consensus amino acid
sequence is shown underneath. Gray high-lighted regions indicate identical amino acids
while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. Percent identity and
percent conserved amino acids determined by the ClustalW alignment tool (Thompson et
al., 1994b).
379
HoxY ClustalW Amino Acid Alignment
HoxY-7002HoxY-6803HoxY-6301HoxY-Ralstonia eutropha
10 20 30 40 50M A Q E T Q Q K I R F A T I WL A G C S G C H M S F L D
M A K I R F A T V WL A G C S G C H M S F L DM D K I K L A T V WL A G C S G C H M S F L D
M R A P H K D E I A S H E L P A T P M D P A L A A N R E G K I K V A T I G L C G C WG C T L S F L DK I . A T . WL A G C S G C H M S F L D
HoxY-7002HoxY-6803HoxY-6301HoxY-Ralstonia eutropha
60 70 80 90 100L D E F L I E L I K Y V D V V F S P V G S D V K D Y P K N V D V C L I E G A V A N Q E N L E L L E KM D E WL I D L A Q K V D V V F S P V G S D L K E Y P D N V D V C L V E G A I A N E E N L E L A L EL D E WL I D L A D G V E V V F S P V A C D R K D Y P E G V D L C L I E G A V A N L D N L E L L H QM D E R L L P L L E K V T L L R S - S L T D I K R I P E R C A I G F V E G G V S S E E N I E T L E H
D E . L I . L . V . V V F S P V . . D . K . Y P . V D . C L . E G A V A N E N L E L L
HoxY-7002HoxY-6803HoxY-6301HoxY-Ralstonia eutropha
110 120 130 140 150V R Q N T K L L I A F G D C A V T T N V T G I R N Q K - - G D - A Q T I L E R G Y K E L T E E H - RL R Q K T K V V I S F G D C A V T A N V P G M R N M L K G S D - - - P V L R R A Y I E L G D G T P QV R D R T R I L V S F G D C A I H A N V P G M R N L W- G A E S A A A V L E R G Y L E L A D T T P QF R E N C D I L I S V G A C A V WG G V P A M R N V F E L K D C L A E A Y V N S A T A V P G A K A V. R T . . L I S F G D C A V . N V P G M R N . D . . L R . Y E L .
HoxY-7002HoxY-6803HoxY-6301HoxY-Ralstonia eutropha
160 170 180 190 200L P Q Q I T G G I L P P L L P R V L P I H E V V D I D L F L P G C P P D A D R I K A A I A P L L E GL P D E P - - G I V P P L L D K V I P L H E V I P V D I F M P G C P P D A H R I R A T L E P L L N GL P N E P - - G I V P P L L N R V T P I H E L V A I E H Y L P G C P P P A D R I R S L L Q A L L D QV P F H P - - - D I P R I T T K V Y P C H E V V K M D Y F I P G C P P D G D A I F K V L D D L V N GL P P G I . P P L L . V P . H E V V . D F . P G C P P D A D R I . . L L L G
HoxY-7002HoxY-6803HoxY-6301HoxY-Ralstonia eutropha
210 220 230 240 250K L P E M E G R E M I K F GE H P L M E G R A M I K F GQ T P P H S G R D L L K F GR - P F D L P S S I N R Y D
P G R . K F G
380
Figure B28. ClustalW alignment of Hyp3 amino acid sequences. Hyp3 amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Anabaena variabilis, and
Prochlorothrix hollandica. The consensus amino acid sequence is shown underneath.
Gray high-lighted regions indicate identical amino acids while lighter shading indicates
conserved amino acids. Dashes represent insertions/deletions included to maximize the
sequence similarity. Percent identity and percent conserved amino acids determined by
the ClustalW alignment tool (Thompson et al., 1994b).
381
Hyp3 Amino Acid Alignment
Hyp3-7002Hyp3-A. variabilisHyp3-7120Hyp3-P. hollandica
10 20 30 40 50M L T A A D I M N P N V V T I K G L A T I A S A T Q C M R V N K T R V L I V D R R H V H D A Y G I LM L K A S D V M T K D V A T I R S S A T V A E A V K L M R A R D WR A L I V D R R H E Q D A Y G I I
M T K D V A T I R S S A T V A E A V K L M R A R D WR A L I V D R R H E Q D A Y G I IM T T P V V T I R R L R T I A D A V R L M R N K G A H A L M V E R R N E A D A Y G I VM T V . T I R A T . A . A V . L M R . . R A L I V D R R H E D A Y G I .
Hyp3-7002Hyp3-A. variabilisHyp3-7120Hyp3-P. hollandica
60 70 80 90 100T A T D I V S K V I A Y G R D P R A I R V Y E I M T K P C I F V S P D L A V E Y V A R L F S Q WN LS E S D I V Y K V I A Y G R D P Y K I R V Y E I M S K P C I A V N P D L G L E Y V A R L F A D Y G LS E S D I V Y K V I A Y G K D P N K I R V Y E V M S K P C I A I N P D L G L E Y V A R L F A D Y G LT E T D I V Y Q V T A H G K D P K T V R V F E V M T K P C I T V N P D L E V E Y V A R L F A M T G I. E . D I V Y K V I A Y G . D P I R V Y E . M . K P C I V N P D L . . E Y V A R L F A . G L
Hyp3-7002Hyp3-A. variabilisHyp3-7120Hyp3-P. hollandica
110 120 130 140 150H S A P V M T D K L L G I I T V E D L I S K S D F L E R P K E L L F A A E M Q A A I Q K T K L I C QH R A P V I Q G E L V G I I S L T D I L A Q S D F L E Q P Y T I L L E Q Q L Q D E I K K A R A V C TH R A P V I Q G D L R G I I S L T D I L A Q S D F L E Q P Y T I L L E Q Q L Q D E I K K A R A V C TR R A P V I Q G Q L L G M I S T T D I L V K S N F V E E P K A Q R L E Q L I Q E A I A A A R Q I C AH R A P V I Q G L . G I I S . T D I L . S D F L E P . L L E Q . Q . I K A R . . C
Hyp3-7002Hyp3-A. variabilisHyp3-7120Hyp3-P. hollandica
160 170 180 190 200E K G H D S S D C I Q A WA M V E E L Q A K T A Y Q Q S T K V D K T A L E E Y L E K N P E A I D H LQ K G I N S E E C A A A WD V I E E M Q A E M A H Q R A E K V S K I A F D D Y C D E Y P E A L E AQ T G I N S E E C A A A WD A V E E M Q A E I A H Q R A E K V S K T A F E D Y C D E Y P E A L E A RD E G T T S P G C A A A WD V V E E L Q A E A A H Q E A K G L I K T A F E E Y L E E N P E A L E A R
G S . C A A A WD . V E E Q A E A H Q A K V K T A F E . Y . E P E A L E A
Hyp3-7002Hyp3-A. variabilisHyp3-7120Hyp3-P. hollandica
210 220 230 240 250M V D N WC S G
L Y E LV Y D V
.
382
Figure B29. ClustalW alignment of HoxH amino acid sequences. HoxH amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 6301, Anabaena variabilis, Anabaena sp. PCC 7120, and
R. eutropha. The consensus amino acid sequence is shown underneath. Gray high-
lighted regions indicate identical amino acids while lighter shading indicates conserved
amino acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids determined by the
ClustalW alignment tool (Thompson et al., 1994b).
383
HoxH ClustalW Amino Acid Alignment
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
10 20 30 40 50M S K T I I I D P M T R I E G H A K I S I Y L D D Q G E V H E A R F H V G E F R G F E K F C E G R PM S K T I V I D P V T R I E G H A K I S I F L N D Q G N V D D V R F H V V E Y R G F E K F C E G R PM T R T I V I D P V T R I E G H A K I S V F L D D Q G N A E A A R F H V V E Y R G F E K F C E G R PM S K R I V I D P V T R I E G H A K I S I Y L D D T G Q V S D A R F H V T E F R G F E K F C E G R PM S K R I V I D P V T R I E G H A K I S I Y L D D T G Q V N D A R F H V T E F R G F E K F C E G R PM S R K L V I D P V T R I E G H G K V V V H L D D D N K V V D A K L H V V E F R G F E K F V Q G H PM S K I V I D P V T R I E G H A K I S I . L D D G . V D A R F H V . E F R G F E K F C E G R P
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
60 70 80 90 100F WE M P A L T S R I C G I C P V S H L I A S S K T G D Q L L A V K I P - - - - - - V G A G K L R RM WE M A G I T A R I C G I C P V S H L L C A A K T G D K L L A V Q I P - - - - - - P A G E K L R RF T E M A G I T A R I C G I C P V S H L L A A A K T G D K I L A V Q I P - - - - - - P A A E N L R RL WE M P G I T A R I C G I C P V S H L L A S A K A G D R I L S V T I P - - - - - - P T A T K L R RL WE M P G I T A R I C G I C P V S H L L A S A K A G D R I L S V T I P - - - - - - P T A T K L R RF WE A P M F L Q R I C G I C F V S H H L C G A K A L D D M V G V G L K S G I H V T P T A E K M R R
WE M P G I T A R I C G I C P V S H L L A A K G D . . L . V I P P A K L R R
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
110 120 130 140 150M I N L A Q I T Q S H A L S F F H L S S P D L I F G WD S D P K T R N V F G L I A A D P D L A R G GL M N L G Q I T Q S H A L S F F H L S S P D F L L G WD S D P A T R N V F G L I A A D P D L A R A GL I N L A Q I I Q S H A L S F F H L S S P D F L L G WD S D P A R R N L F G L I A A D P E S A R A GL M N L G Q I L Q S H A L S F F H L T A P D L L L G M D S D P Q K R N I F G L I A A Q P E L A R G GL M N L G Q I L Q S H A L S F F H L S A P D F L L G M D S D P Q K R N I F G L I A A Q P E L A R G GL G H Y A Q M L Q S H T T A Y F Y L I V P E M L F G M D A P P A Q R N V L G L I E A N P D L V K R VL N L . Q I . Q S H A L S F F H L S P D L L G D S D P R N . F G L I A A P . L A R . G
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
160 170 180 190 200I R L R K F G Q D I I K I L G S Q K V H P A WS I P G G V R S P L T K E G Q T Y I K E G - - - - - -I R L R Q F G Q T V I E L L G A K K I H S A WS V P G G V R S P L S E E G R Q WI V D R - - - - - -I R L R Q F G Q Q L I E WL G G R K I H A A WA V P G G V R S L L S Q E G M V WI R D R - - - - - -I R L R Q F G Q E I I E V L G G A K I H P A WA V P G G V R E P L S V E G R T H I Q E R - - - - - -I R L R Q F G Q E I I E V L G G A K I H P A WA V P G G V R E P L S A E G R T H I Q E R - - - - - -V M L R K WG Q E V I K A V F G K K M H G I N S V P G G V N N N L S I A E R D R F L N G E E G L L SI R L R Q F G Q . . I E . L G G K I H A W V P G G V R P L S E G R I . R
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
210 220 230 240 250L P E A K N T V K N A L S L F K K I L D S - H Q E E V T V F G N F P S L Y L G L I G E S G A WE H YL P E A K E T V Y L A L N L F K N M L D R - F Q T E V A E F G K F P S L F M G L V G K N N E WE H YL P A E L A T V R A T L D R F K R L L D R E F R E E T A V F G D F P S L F M G L V A D N G D WE H YI P E A R T I A L D A L D R F K K L L K D - Y E K E V Q T F G N F P S L F M G L V T P D G L WE T YI P E A R T I A L D A L D R F K K L L K D - Y E K E A Q T F G N F P S L F M G L V T P D G L WE T YV D Q V I D Y A Q D G L R L F Y D F H Q K - H R A Q V D S F A D V P A L S M C L V G D D D N V D Y Y. P E A . . A L F K . . L . E V F G F P S L F M G L V . G WE Y
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
260 270 280 290 300G G I L R M I D S H G N I V G D R L D P N N Y H E F L G E K L Q E D S Y L K S P Y Y K L F G - - - -G G S L R F T D S E G N I V A D N L S E D N Y A D F I G E S V E K WS Y L K F P Y Y K S L G Y P - -G G H L R F V D S Q G R I I A D R L R E D D Y A S F L G E A V E P WS Y L K F P Y Y K P WG Y P - -D G Y I R F V D S A G N I I A D K L D P A R Y Q E F I G E A V Q P D S Y L K S P Y Y R P L G Y P D QD G Y I R F V D S A G N I I A D K L D P T R Y Q E F I G E A V Q P D S Y L K S P Y Y R P L G Y P D QH G R L R I I D D D - K H I V R E F D Y H D Y L D H F S E A V E E WS Y M K F P Y L K E L G R - - -
G L R F . D S G N I I A D . L D Y . F . G E A V S Y L K P Y Y K L G Y P
384
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
310 320 330 340 350- - - - - - E T G V Y R V G P L A R L N I C E H F G T E A A D Q E L I E Y R Q R H G R - I V Q A S F- - - - - - D G - I Y R V G P L A R L N V C H H I G T P E A D Q E L E E Y R Q R A G G - V A T S S F- - - - - - E G - M Y R V G P L A R L N V C D R I G T G R G G S R A T G T A R S R G G - T V T S S FH D Q C R I D S G M Y R V G P L A R L N I C S H I G T T L A D R E L R E F R E L T S G - T A K S S FH D Q C R I D S G M Y R V G P L A R L N I C S H I G T T L A D R E L R E F R E L T S G - T A K S S F- - - - - - E Q G S V R V G P L G R M N V T K S L P T P L A Q E A L E R F H A Y T K G R T N N M T L
. G Y R V G P L A R L N . C H I G T . A D E L E . R G T . S S F
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
360 370 380 390 400V Y H H A R L I E I L G S L E R I E R M I D D P D L F S N - - R L Q A E A G V N Q T E A V G V S E AF Y H Y A R L V E I L A C L E A I E L L M A D P D I L S K - - N C R A K A E I N C T E A V G V S E AL Y H Y A R L V E I L A S L E K I A E M V E D P N L Q T G - - F L R S Q A G V N C L E A I G V S E AF Y H Y A R L I E I L A C I E H I E M L L D D P D I L S N - - R L R S E A G V N Q L E A V G V S E AF Y H Y A R L I E I L A C I E H I E I L L N D P D I L S T - - R L R S E A G V N Q L E G V G V S E AH T N WA R A I E I L H A A E V V K E L L H D P D L Q K D Q L V L T P P P N A WT G E G V G V V E A
Y H Y A R L I E I L A . E . I E L . D P D . S L R A G V N . E A V G V S E A
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
410 420 430 440 450P R G T L F H H Y Q V D E N G L L K K I N L V I A T G Q N N F A I N R T V T Q I A K H Y I H G E - TP R G T L F H H Y K I D E D G L I K K V N L I I A T G N N N L A M N K T V A Q I A K H Y I R N H - DP R G T L F H H Y R V D S H G K I E R V N L I I A T G Q N N L A M N R T V T Q I A Q H Y I R H G - EP R G T L F H H Y Q V D E N G L L Q K V N L I I A T G Q N N L A M N R T V A Q I A R H F I Q G T - EP R G T L F H H Y Q V D E N G L L Q K V N L I I A T G Q N N L A M N R T V A Q I A R H F I Q G T - EP R G T L L H H Y R A D E R G N I T F A N L V V A T T Q N H Q V M N R T V R S V A E D Y L G G H G EP R G T L F H H Y V D E G L . K V N L I I A T G Q N N L A M N R T V Q I A . H Y I G E
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
460 470 480 490 500V A E G I L N R V E A G V R N Y D P C L S C S T H A A G Q M P M I L Q L I S P D G S I V K E I R R DV Q E G F L N R V E A G I R C Y D P C L S C S T H A A G Q M P L M I D L V N P Q G E L I K S I Q R DV Q E S F L N R V E A G I R C F D P C L S C S T H T A G Q M P L K I E I F D S R G E L Y Q C L C R DI P E G M L N R V E A G I R A F D P C L S C S T H A A G Q M P L H I Q L V A A N G N I V N Q V WR EI L E G M L N R V E A G I R A F D P C L S C S T H A A G Q M P L H I Q L V A A D G N I V N Q V WR EI T E G M M N A I E V G I R A Y D P C L S C A T H A L G Q M P L V V S V F D A A G R L I D E R A R. E G L N R V E A G I R . D P C L S C S T H A A G Q M P L I L . G . . . R .
HoxH-7002HoxH-6803HoxH-6301HoxH A. variabilisHoxH-7120HoxH-Ralstonia eutropha
510 520 530 540 550S
LK L G V
385
Figure B30. ClustalW alignment of HoxW amino acid sequences. HoxW amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, and R. eutropha. The consensus amino acid sequence is
shown underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. The percent identity and percent conserved amino
acids were determined with the ClustalW alignment tool (Thompson et al., 1994b).
HoxW ClustalW Amino Acid Alignment
HoxW-7002HoxW-6803HoxW-7120HoxW-Ralstonia eutropha
10 20 30 40 50M K Q T L M I G Y G N T L R S D D G A G Q K V A E A F F D Q - - E N I T A I A T H Q L T P
M P G Q S T K S T L I I G Y G N T L R G D D G V G R Y L A E E I A Q Q N WP H C G V I S T H Q L T PM K K T V M V I G Y G N D L R S D D G I G Q R I A N E V A S WR L P S V E S L A V H Q L T P
M K E Q E - I D R I A T M I Y E A P L G E Y I G R D G A A I L A E H A A E A R L L K G D EK I G Y G N T L R D D G . G . A . . A . . H Q L T P
HoxW-7002HoxW-6803HoxW-7120HoxW-Ralstonia eutropha
60 70 80 90 100- E L V E D L V Q V E Q V Y F I D A A P - - - - - - - - I E T V T I K P I Q R R D N - E H N F G H F- E L A E A I A A V D R V I F I D A Q L Q E S A N E P S V E V V A L K T L E P N E L - S G D L G H R- D L A D S L A S V D L A I F I D A C L P V H G - - - - - F D V K V Q P L F A A G D - I D S N V H TF L Y R R G D V T S S F Y I V T D G R L A L V R E K T N E R T A P I V H V L E K G D L V G E L G - F
. L . . . . V . . I F I D A L V . . G H
HoxW-7002HoxW-6803HoxW-7120HoxW-Ralstonia eutropha
110 120 130 140 150I D - - P K S L - - L N L - A Q E I Y H Y A P D A Y L V L I - - - P A Q D F K L G E N - Y S E I T QG N - - P R E L - - L T L - A K I L Y G V E V K A WWV L I - - - P A F T F D Y G E K - L S P L T AG D - - P R S L - - L A L - T K A I Y G N C P T A WWV T I - - - P G A N F E I G D R - F S R T A EI D Q T P H S L S V R A L G D A A V L S F S A E S I K P L I T E H P E L I F N F M R A V I K R V H H. D P . S L L L . . Y A . V L I P . F G . S .
HoxW-7002HoxW-6803HoxW-7120HoxW-Ralstonia eutropha
160 170 180 190 200K A I E T A I H L - - L Q E R L T P C M KR A Q A E A L A Q - - I R P L V L G E RT G K A I A L V K - - I I Q I L D K V N N L WF E V G A V AV V V T V G E H E R E L Q E Y I S T G G R G R - - - G
. A . . .
386
Figure B40. ClustalW alignment of HypA amino acid sequences. HypA amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 6301, Aquifex aeolicus and Bradyrhizobium japonicum.
The consensus amino acid sequence is shown underneath. Gray high-lighted regions
indicate identical amino acids while lighter shading indicates conserved amino acids.
Dashes represent insertions/deletions included to maximize the sequence similarity.
Percent identity and percent conserved amino acids determined by the ClustalW
alignment tool (Thompson et al., 1994b).
HypA ClustalW Amino Acid Alignment
HypA-7002HypA-6803HypA-6301HypA-Aquifex aeolicusHypA-Bradyrhizobium japonicum
10 20 30 40 50M H E A I M T E T V I A N A A A E Q N A T K I G L T M R I G I S G V V P E L S F A F E AM H E S L M E Q T L I A I A Q A E H G A S Q I R L T L R V G Q S G V V A D L R F A F E VM H E S L A T A L V T A L WQ A V A E A Q Q I S L K L R L G WA G V D A E L R F A F S LM H E S I V Q S L L L I E D Y A R N N A K S V K V V V S I G L S G V E P H L E M A F N TM H E A L C E G I I I V E E E A R R A F A K V V V C L E I G L S H V A P E L Q F C F E AM H E S L . . . I A A A I L L R I G . S G V . P E L F A F E .
HypA-7002HypA-6803HypA-6301HypA-Aquifex aeolicusHypA-Bradyrhizobium japonicum
60 70 80 90 00V A G T L A E Q A Q I I E T V P V C Y C A Q C R P F T P P D F Y E C P L C S L S Q H I LV R Q T M A A E A R E I E E I P V C R C Q H C E N F Q P E D I Y R C P H C Q I S Q T V MV Q Q T I A A S A Q V I E S V P A F R C Q T C Q Q T P P P - L A A C S H C S D R WQ L QF K E T V A E K A E I M E I E K L I K C M D C K E S E K E E N M L C P G C S L N T Q I IV A A T I A Q G A K E I V E T P G A WC M A C K S V E I K Q Y E P C P S C G Y Q L Q V TV T . A A . I E . P . . C C . P . C P C S . Q .
HypA-7002HypA-6803HypA-6301HypA-Aquifex aeolicusHypA-Bradyrhizobium japonicum
110 120 130 140 50S G K V E L K S L E ID G K L E L A S L E SQ G R L Q L Q S M E VS G Q M F L K S L E E V DG G E M R V R E L E D
G . . L . S L E
387
Figure B41. ClustalW alignment of HypB amino acid sequences. HypB amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Synechococcus sp. PCC 6301, Bradyrhizobium japonicum, and Azotobacter
vinelandii. The consensus amino acid sequence is shown underneath. Gray high-lighted
regions indicate identical amino acids while lighter shading indicates conserved amino
acids. Dashes represent insertions/deletions included to maximize the sequence
similarity. Percent identity and percent conserved amino acids determined by the
ClustalW alignment tool (Thompson et al., 1994b).
388
HypB ClustalW Formatted Alignments
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
10 20 30 40 50M C G N C G C N - - A V E K P V E I H T H A H D H - - - - - - - S H G H H D H H H P H D H E H H H HM C Q N C G C S - - A V G T V A H S H H H H G D G - - - - - - - N F A H S H D D H D Q Q E H H H H HM C G V C G C Q - - S P D P V V I A T P E P - - - - - - - - - - - - - - - - - - - - - - - - - - - -M C V T C G C S D E S E D K I T N L E T G E M V H N - - - - - - - H Q C I Q H T L A D G T V I T H SM C T V C G C S D - G K A S I E H A H D H H H D H G H D H D H G H D G H H H H H H G H D Q D H H H HM C . C G C S . . . . . H H D H . H H H H H H H
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
60 70 80 90 100H N H D G - - - - - - - - - - N H S - - - - - - - - - - H A I D I N Q S I F A K N D R L A E R N R GG N Y S K - - - - - - - - - - S P S Q Q T V T I E P D R Q S I A I G Q G I L S K N D R L A E R N R G- - - - - - - - - - - - - - - - - - - - - - - - - - - - R Q I S L A E A I L H R N D H A A A H N R EH S H E Q - - - - - - - - - - E P S Q - - I P A K I H N T T I S L E Q E I L G K N N L L A A Q N R GH D H A H G D A G L L D C G A N P A G Q K I T G M S S D R I I Q V E R D I L G K N D R L A A D N R AH H P S . . . I . Q I L . K N D R L A A . N R G
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
110 120 130 140 150Y F L A K D L F V I N V V S S P G S G K T A L L E R T I E Q L K N K L N S A V I V G D L K T D N D AY F Q A K G L L V M N F L S S P G A G K T A L I E K M V G D R Q K D H P T A V I V G D L A T D N D AH F Q R A G V L A L N L L S S P G S G K T A L L V R S F R E L P P T L R P A A I V G D L A T D R D AWF K G R N I L S L N L M S S P G A G K T T L L T Q T I N D L K H Q L P I T V I E G D Q E T I N D AR F R A D E V L A F N L V S S P G A G K T S L L V R A V S E L K D S F A I G V I E G D Q Q T S N D A. F A . . L . . N L . S S P G A G K T A L L R . . . L K L A V I V G D L T D N D A
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
160 170 180 190 200Q R L R K N N I P V A Q I T T G T L C H L D A E M V L N A A R K L A L D G V H L L I I E N V G N L VQ R L R S A G A I A I Q V T T G N I C H L E A E M V A K A A Q K L D L D N I D Q L I I E N V G N L VQ R L Q A T G A P A D Q I E P A E L C H L E A D L V H R A C H Q L D L S A I D I L F I E N V G N L VE K I K E T G C K V V Q I N T G T G C H L D A S M I E R G L Q Q L N P P I N S V L M I E N V G N L VE R I R A T G V P A I Q V N T G K G C H L D A A M V G E A Y D R L P WL N G G L L F I E N V G N L VQ R L R T G . P A . Q I T G . C H L D A . M V . . A . . L L . . . L I E N V G N L V
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
210 220 230 240 250C P A A Y D L G E H K R I V L L S T T E G E D K P L K Y P T M F K S A D V V I I N K I D I A E V V GC P T T Y D L G E D L R V V L F S V T E G E D K P L K Y P A T F K S A Q V I L V T K Q D I A A A V DC P A A F D L G E E R R V L L L S V T E G E D K P L K Y P S A F S R A D L V L I T K V D L A E A V EC P A L F D L G E Q A K V V I L S V T E G E D K P I K Y P H I F R A S E I M I L T K I D L L P Y V NC P A A F D L G E A C K I V V F S T T E G E D K P L K Y P D M F A A S S L M L I N K I D L A S V L DC P A A F D L G E R V V L L S V T E G E D K P L K Y P F . A . . . L I T K I D L A . V .
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
260 270 280 290 300F D R D S A L E N V K K M C P Q A Q I F E L S A R T G E G M E S WL N Y L Q S Q Y Q D I A F Q A V IF D A E L A WQ N L R Q V A P Q A Q I F A V S A R T G K G L Q S WY E Y L D Q WQ L Q H Y S P L V DF D Q A A A I A N L R A V N P T A T I L P V S S R G G Q G WS D WL D WL Q V Q R S H L L A P V K AF D V Q R C V K Y A K Q V N P N I Q I F Q V - - - - - - - - - - - - - F L Q L R V L R VF D L A R T I E Y A R R V N P K I E V L T L S A R T G E G F A A F Y A WI R K R M A A T T P A A M TF D . A . N . R V N P . A Q I F V S A R T G G W . L Q . . .
HypB-7002HypB-6803HypB-6301HypB-7120HypB-B. japonicum
310 320 330 340 350T VP A L A
A A E.
389
Figure B42. ClustalW alignment of HypF amino acid sequences. HypF amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Aquifex aeolicus and R. sphaeroides. The consensus amino acid sequence is
shown underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW alignment tool (Thompson et al., 1994b).
390
HypF ClustalW Amino Acid Alignment
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
10 20 30 40 50L L V K R R L R L E I Q G T V Q G V G F R P F V Y Q L A T A L N L F G WV N N S T A G V T I E V E G
M L K T V A I Q V Q G R V Q G V G F R P F V Y T L A Q E M G L N G WV N N S T Q G A T V V I T AM P F K R L K L H L K G A V Q G V G F R P F V Y R I A K E L G L K G F V I N D S K G V Y I E V E G
M Q A WR I R V R G Q V Q G V G F R P F V WQ L A R A R G L R G V V L N D A E G V L I R V A G. . . . . G V Q G V G F R P F V Y L A G L G . V N . G V I V G
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
60 70 80 90 100G R S P L N L F L E K L Q A E L P P N A K I D A L K Y Q Y L E L I G Y N N F E I H A S Q T - G E K ID E K A I A D F T E R L T K T L P P P G L I E Q L A V E Q L P L E S F T N F T I R P S S D - G P K TE E E R L K K F L F K L N R E K P P L A R I Y S Q E I Q F L E P V N Y E D F V I R K S E E K G E K ED - - - L G D F A A A L R D Q A P P L A R V D A V E V T A A V C D D L P E G F Q I A A S G A A G A E. L F . . L . P P A . I . . . L . F I . S G K
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
110 120 130 140 150A I V L P D L A T C S E C I A E I F D P Q N R R Y Q Y P F T N C T H C G P R Y S I I E T L P Y D R SA S I L P D L S T C S A C L T E L F D P S D R R Y L Y P F I N C T H C G P R Y T I I E A L P Y D R CV L V L P D I A T C E D C L R E L F T P E D R R Y M Y P F I N C T N C G P R F T I I E R L P Y D R KT R V T P D A A T C P D C L A E I R G - E G R R R G Y A F T N C T H C G P R F S I L Q S L P Y D R A. V L P D . A T C . C L E . F P R R Y Y P F N C T H C G P R . . I I E L P Y D R
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
160 170 180 190 200L T S M A D F S M C A D C Q R E Y E D P G D R R F H A Q P N A C P I C G P K L E F L S H C Q G Q E NR T T M A R F R Q C T D C E R E Y K Q P G D R R F H A Q P N A C P R C G P Q L A F WN R Q G Q V I AN T T M K V F E M C P E C K R E Y E N P L D R R F H A Q P N A C P K C G P WV S L Y K D - G K L I AR T T M A P F A M C P A C R A E Y E D P A D R R F H A Q P I A C P D C G P R L WL E A G G A E L P G
T T M A F M C . C R E Y E P . D R R F H A Q P N A C P C G P L . . .
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
210 220 230 240 250T N Q S P L E A A I Q Y I R D G K I V A L K G L G G F Q L L V D A R N N K A V Q Q L R G R K Q R P DE A N E A L N F A V D N L K V G N I I A I K G L G G F H L C C D A T D F E A V E K L R L R K H R P DE K N E A L E L L I E E I K I G K I V A V K G V G G F H L I C N A T N E E S V R T L R K R K R R S E- - - D A I G L A A A R L K A G E I L A V K G L G G F H L A C D A T N A D A V D L L R A R K R R P A
. . A L . A . . K . G I . A . K G L G G F H L . C D A T N . A V L R . R K . R P .
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
260 270 280 290 300K P F A V MY P D L P S I K N D C F L S Q L E E A F L T S Q A S P T V L L Q K K K E F N L A E N V AK P L A V MY G N L G Q I V E H Y Q P N N L E V E L L Q S A A A P I V L L N K K K Q L I L V E N I AK P F A V MF K S L E Q V E A Y A N P T E L E K A L L I S P E R P I V L I Q K K K - - E L A P S V SK P F A L MA R - E E D L A R I V A V S P A A L A A L R D P A A P I V L M P A R G - - S L P E T L AK P F A V M. L . . L E A . L S A P I V L . . K K K L . E . A
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
310 320 330 340 350P H N P N L G V M L P Y T P L H H L L L K S L N F P V I A T S G N R S D E P I C I D E T E A L E R LP G N P R V G V M L A Y T P L H H L L L K K L K K P M V A T S G N L A G E Q I C I D N I D A L T R LP G L K R V G A F L P Y S P L H H L I L N S L D F P V V A T S A N I S E E P I I K D N K E A L E K LP G M A E L G V M L P Y T P L H H L L L D A F G G V L V M T S G N L S G A P Q V I G N D E A R E K LP G . G V M L P Y T P L H H L L L L P . V A T S G N . S E P I I D N E A L E . L
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
360 370 380 390 400K N I A D G F L I H N R R I L R P V D D S V V R V M N N T P I I L R R S R G Y A P E P L T L K - - RQ N I A D G F L V H D R P I V C P V D D S V V Q I V A G K P L F L R R A R G Y A P Q P I T L P - - KG E L A D L I L V H N R D I K R R C D D S V V K V V S G V P T P I R R S R G Y A P L P V E V P - - FS A F A D A F L M H D R A I A R R L D D S V V R V D P - - P M V L R R A R G Q V P G T L P L P P G F
. A D . F L . H R I . R . D D S V V . V P L R R R G Y A P P . L P
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
410 420 430 440 450T L S K N V L A M G A H L K N T V A I A Q K N R L F L S Q H I G D L S N Q L T L Q A M T K T L Q K LP T Q K K L L A M G G H Y K N T V A I A K Q N Q A Y V S Q H L G D L N S A P T Y Q N F E E A I A H LE L P K R V L A V G G M L K N T F A L G F K N Q V I L S Q H I G D I E N L N T L K V F E E S V F D LE T A P Q I V A Y G G Q M K A A L C L I K T G Q A L L G H H L G E L D E A L T WE A F L Q A D A D Y
K . L A G G K N T . A . . N Q . L S Q H . G D L . T . F . L
391
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
460 470 480 490 500S Q I Y D F Q P D I I A C D L H P D Y L S T Q Y A K N L A Q K L N I S L I S V Q H H H A H I Y A C MS Q L Y D F S P Q E I V A D L H P D Y F S H Q Y A E N Q A L P V T F - - - - V Q H H Y A H I L A V MM E L Y E F E P D V V V C D M H P R Y E T T R WA E E F S R K R G I P L I K V Q H H Y A H M L S C MA A L F D H R P Q A V A V D L H P D F R A S R H G A A R A G R L G V P L I A V Q H H H A H L A A C L
L Y D F P . . . D L H P D Y . . . A A . . . L I V Q H H A H . . A C M
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
510 520 530 540 550A E H - Q L E S P L L G - - - - V A WD G T G Y G E D G T I WG G E F F WV T K Q S C E R I A H F KA E H G V M E E S V L G - - - - I A WD G T G Y G M D G T I WG G E F L K I T Q G T WQ R I A H L QA E N - G I K E K V L G - - - - I A WD G T G Y G E D G T L WG G E F L V A D Y T S Y E R A F N F KG E N - - L WP K D G G K V A V I V L D G L G L G P D G T V WG G E L L L G D Y K G F E R V A WL KA E . . L G I A WD G T G Y G D G T . WG G E F L . . . E R . A K
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
560 570 580 590 600P F P L P G G D R A S R E P R R S A L G L L S Q L Y D L Q V L K K L N L P T I Q A F S A T E L E L LP F H L L G N Q Q A I K Y P H R I A L A L L WP T F G D D F S A D S - L G N WL N F N N G F K N K IP V K L I G G E K A V K E P R R V A L S L L F D I F G E E A L N L D - L L P V K S F S E R E L K N LP A P L I G G D R A Q I E P WR N A L V R L D A A G L S D L A D R L - - - - - - - F P A A P R D L AP L . G G . . A . E P . R A L . L L . . . . . . L F .
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
610 620 630 640 650L S M L K K D I N - - - - - - - - - - - - T P F T S S V G R L F D G V A S L L D L R - Q R E S F E GN S R L N Q D L N N K N L R Q L WQ R G Q A P L T S S M G R L F D G I A T L I G L I - N E V T F E GY L A WK K G I N - - - - - - - - - - - - S P L S S S V G R L F D A L A S L L N L K - Q I L S Y E GR Q L A A K G I N - - - - - - - - - - - - A P L S S S A G R L F D A V A A C L G I C P M R Q S Y E G
. K I N P L . S S . G R L F D . . A . L L L . S . E G
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
660 670 680 690 700Q A A M A L E F S I D G L Q I P D F Y Q F Q Y T K N D S I L E I D S R G I F Q G I I Q D L Q N D L PQ A A I A L E A Q I M P N L T E E Y Y P L T L N N K E K K L A V D WR P L I K A I T T E D R - - S KQ G A M M V E D L Y D P - L V K D N Y P Y E I R G K E - - - - V D L R K A F L E V L K E K D - - - KE A A M R L E S L A A D T G P V P D L P C V G G - - - - - - A I D P A P L F Q L L A A G E R - - - PQ A A M L E . . . Y P . . . D R . F . . . .
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
710 720 730 740 750K N F I A A K F H N T L V E I I F D I Y L H T L K L G F N S R K N I V L A G G C F Q N K Y L L E Q TT N L I A T K F H N S L V N L I I T I A Q Q - - - - - - Q G I E K V A L G G G C F Q N C Y L L A S TS - L A A S R F I N T L A K V C E D I A L M - - - - - - V G I E R V C L S G G V M Q N D P L V T K ID - R V A H A L H A S L A Q A F A A E A R R - - L I E A G Q A E A V A L T G G C F Q N S R L A T M T
. A . F H N . L . . I A . E V . L G G C F Q N L . T
HypF-7002HypF-6803HypF-Aquifex aeolicusHypF-Rhodobacter capsulatus
760 770 780 790 800I Q K L E S S G A N I Y Y P Q K F P P N D G A I A L G Q V M V V T G Q A II T A L K K A G F S P L WP R E L P P N D G A I C M G Q L L A K I Q A R Q Y I CK E L L E K E K F K V Y T H Q K V P P N D G G I A L G Q A V F G L S L VR N F L A D Q G - - I L T Q G R I P A N D G G L A L G Q A L V A A A K L E S N
L G . . . P P N D G . I A L G Q . . . . . .
392
Figure B43. ClustalW alignment of HypC amino acid sequences. HypC amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, E. coli, Synechocystis sp.
PCC 6803, and R. eutropha. The consensus amino acid sequence is shown underneath.
Gray high-lighted regions indicate identical amino acids while lighter shading indicates
conserved amino acids. Dashes represent insertions/deletions included to maximize the
sequence similarity. Percent identity and percent conserved amino acids determined by
the ClustalW alignment tool (Thompson et al., 1994b).
HypC ClustalW Amino Acid Alignment
HypC-7002HypC-E. coliHypC-6803HypC-Ralstonia eutropha
10 20 30 40 50M C L A V P G K I L E I V G D - D P L F K M G R V S F S G V V R E V S L A Y V P E A - - - - - - Q VM C I G V P G Q I R T I D G N - - - - - - Q A K V D V C G I Q R D V D L T L V G S C D E N G Q P R VM C L A L P G Q V V S L M P N S D P L L L T G K V S F G G I I K T I S L A Y V P E V - - - - - - K VM C L A I P A R L V E L Q A D - - - - - Q Q G V V D L S G V R K T I S L A L M A D A - - - - - - V VM C L A . P G . . . . G . V G . . . S L A V . . V
HypC-7002HypC-E. coliHypC-6803HypC-Ralstonia eutropha
60 70 80 90 100G D Y A V V H A G F A L S V L D E V A A T E T L A T L A E M E S F A G GG Q WV L V H V G F A M S V I N E A E A R D T L D A L Q N M F D V E P D V G A L L Y G E E KG D Y V I V H V G F A I S I V D E E A A Q E T L I D L A E M G VG D Y V I V H V G Y A I G K I D P E E A E R T L R L F A E L E R V Q P P A S E P M H G M N I H Q E PG D Y V . V H V G F A . S . . D E A . T L L A E M
HypC-7002HypC-E. coliHypC-6803HypC-Ralstonia eutropha
110 120 130 140 150
A
393
Figure B44. ClustalW Alignment of HypD Amino Acid Sequences. HypD amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, Anabaena sp. PCC 7120, E. coli, and R. eutropha. The consensus amino acid
sequence is shown underneath. Gray high-lighted regions indicate identical amino acids
while lighter shading indicates conserved amino acids. Dashes represent
insertions/deletions included to maximize the sequence similarity. Percent identity and
percent conserved amino acids determined by the ClustalW alignment tool (Thompson et
al., 1994b).
394
HypD ClustalW Amino Acid Alignment
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
10 20 30 40 50M K F V D E F R D P A A V Q K Y V Q A I A A L V T R P - - - - WT - - - I M E I C G G Q T H S I V KM K Y V D E Y R D A Q A V A H Y R Q A I A R E I T K P - - - - WT - - - L M E I C G G Q T H S I V KM K Y V D E F R E P E K A E A L R R E I E K L S Q Q L - - - - D K H I K I M E V C G G H T H S I F KM R F V D E Y R A P E Q V M Q L I E H L R E R A S H L S Y T A E R P L R I M E V C G G H T H A I F KM K Y I E E F R D G E L A Q R I A A H V R A E A R P G - - - - Q R - Y N F M E F C G G H T H A I S RM K Y V D E F R D P E . V . . . I . . . . . I M E . C G G H T H S I K
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
60 70 80 90 100Y G I D Q L L P P E I T L I H G P G C P V C V T P A E L I D Q A I A L A Q L P D V V L C S F G D M LY G L D A L L P K N L T L I H G P G C P V C V T P M E L I D Q A L WL A K Q P E I I F C S F G D M LY G I E E I L P A N I E L I H G P G C P V C V M P K G R L D D A I A I S Q N P N V I L T T F G D T MF G L D Q L L P E N V E F I H G P G C P V C V L P M G R I D T C V E I A S H P E V I F C T F G D A MY G V T E L L P E N V R M I H G P G C P V C V L P I G R I D L A L H L A L E R D A I V C T Y G D T MY G . D L L P N . L I H G P G C P V C V P G R I D A . L A P . V I . C T F G D M
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
110 120 130 140 150R V P G T R - L D L L S V K A Q G A A V K M V Y S P L D A L K M A Q E N P D K Q V I F F A V G F E TR V P G S G - A D L L S I K A Q G G D V R I V Y S P L D C L A I A R E N P N R E V V F F G V G F E TR V P G S K - T T L L Q A K A Q G A D I R M V Y S P L D S L Q I A R N H P D K E I V F F A L G F E TR V P G K Q - G S L L Q A K A R G A D V R I V Y S P M D A L K L A Q E N P T R K V V F F G L G F E TR V P A S G G M S L I R A K A H G A D I R M V Y S A A D A L K I A Q R H P Q R E V V F L A I G F E TR V P G S . . L L A K A Q G A D V R M V Y S P L D A L K I A Q E N P R E V V F F A . G F E T
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
160 170 180 190 200T A P T T A M A V Y Q A A K L G L T N F S L L V A H V S V P P A I E A I L S A P - - - - - - D R T IT A P A T A M T L H Q A R A Q G I S N F S L L C A H V L V P P A M E A L L G N P - - - - - - N S L VT A P S T A L T I L Q A A S E N I T N F S M F S N H V L V I P A L Q A L L N N P - - - - - - D L Q LT M P T T A I T L Q Q A K A R D V Q N F Y F F C Q H I T L I P T L R S L L E Q P - - - - - - D N G IT T P P T A L I I R E A K A R Q V D N F S V L C C H V L T P S A I T H I L E S P E V R D Y G T V P IT A P . T A . T . Q A . A . . N F S . L C H V L V P P A . A L L . P D I
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
210 220 230 240 250Q G F L L A G H V C T V M G Y Q E Y E A I A K N Y Q I P L I V T G F E P L D I V Q G I Y L C V K Q LQ G F L A A G H V C T V T G E R A Y Q H I A E K Y Q V P I V I T G F E P V D I M Q G I F A C V R Q LD G F I G P G H V S M V I G T E P Y E F I A Q Q Y H K P I V V S G F E P L D I F Q S I WM L L Q Q LD A F L A P G H V S M V I G T D A Y N F I A S D F H R P L V V A G F E P L D L L Q G V V M L V Q Q KD G F V G P A H V S I V I G T R P Y E H F S R E Y G K P V V I A G F E P L D V M Q A I L M L V R Q VD G F L . P G H V S V I G T Y E I A Y . P . V V . G F E P L D I Q G I . M L V . Q L
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
260 270 280 290 300E E G R S H I E N Q Y R R V V Q A A G N A T A Q Q L V T E I F E I V P R - T WR G I G E I S Q S G LE S G Q F T C N N Q Y R R S V Q P Q G N A H A Q K I I D Q V F E P V D R - H WR G L G L I P A S G LV E N R C E V E N Q Y N R L V Q K G G N Q I A L A A M H K V F A V R E K F A WR G L D E I P D S G LI A A H S K V E N Q Y R R V V P D A G N L L A Q Q A I A D V F C V N G D S E WR G L G V I E S S G VN S G R A E V E N E F V R A V T R D G N E S A Q A M V S E V F E L R P S F E WR G L G E V P Y S A L
G R V E N Q Y R R . V Q . G N . A Q . . . V F E . . WR G L G E I P S G L
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
310 320 330 340 350G L R E K Y A V F D A S R K F K L D L T H F Q P - Q A S S S C I S G E I L R G R K K P K Q C P A F GG L R P A F A P WD A A V K F A N L L Q T M A P T M G E T V C I S G E I L Q G Q R K P S D C P A F GK I R E E Y A Q F D A E L K F T I P N L K V A D - - - H K A C K C G E I L K G V L K P WQ C K V F GH L T P D Y Q R F D A E A H F R P A P Q Q V C D - - - D P R A R C G E V L T G K C K P H Q C P L F GR I R A Q F A R F D A E Q R F D L R Y R P V P D - - - N K A C E C G A I L R G V K K P T D C K L F A. L R Y A F D A E . K F . V D . C C G E I L . G . K P Q C P . F G
HypD-7002HypD-6803HypD-7120HypD-E. coliHypD-Ralstonia eutropha
360 370 380 390 400T T C T P E R P L G A P M V S S E G A C A A Y Y R Y GT I C T P E Q P L G A P M V S S E G A C A A Y Y R Y R Q Q L P E P V G A A R VT A C T P E T P I G T C M V S S E G A C A A Y Y K Y G R F S T T L Q K Q A A E K P K V T I S SN T C N P Q T A F G A L M V S S E G A C A A WY Q Y R Q Q E S E AT V C T P E N P M G S C M V S S E G A C A A H Y S Y G R F K D I P L V A AT . C T P E P . G A M V S S E G A C A A Y Y . Y G A
395
Figure B45. ClustalW alignment of CtaCI amino acid sequences. CtaCI amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
396
CtaCI ClustalW Amino Acid Alignment
CtaCI-7002CtaCI-6803CtaCI-7120
10 20 30 40 50M K I V T N Q R H F Q T V C F N V V T S S Y S K K A P T F P V N I P N S I T T L I A G I A I T L I S
M K I P G S V I T L L I G V V I T V V SM Q Q I P V S L WT L I A G I V V G V I S
. I P . S . T L I A G I V I T V I S
CtaCI-7002CtaCI-6803CtaCI-7120
60 70 80 90 100L WY G Q N H G L L P V A A S A D A N D V D E L F N L M M T I A T G L F I L I E G V L V I C L I R FL WY G Q N H G L M P V A A S A D A E K V D G I F N Y M M T I A T G L F L L V E G V L V Y C L I R FL WI G Q N H N L L P I Q A S E Q A P L V D G F F N I M F T I A V A L F L V V E G T I L I F L F K YL WY G Q N H G L L P V A A S A D A V D G . F N . M M T I A T G L F L L V E G V L V I C L I R F
CtaCI-7002CtaCI-6803CtaCI-7120
110 120 130 140 150R R K Q G D L T D G P A I E G N V P L E I V WT A I P T V I V F I L A I Y S F E I Y N K M G G L D PR R R K D D Q T D G P P I E G N V P L E I L WT A I P T V I V F T L A V Y S F E V Y N N L G G L D PR R R R G D N T D G V P V E G N V P L E I F WT A I P S I I V I C L G I Y S V D V F N Q M G G L E PR R R . G D . T D G P P I E G N V P L E I . WT A I P T V I V F L A I Y S F E V Y N . M G G L D P
CtaCI-7002CtaCI-6803CtaCI-7120
160 170 180 190 200M V - - S G G G M T M A H H H Q H N P N T M D N M V A M E P D S K I A I G I G K S M S A N D - E D PT I S R D N A G Q Q M A H N H M G H M G S M G N M V A M A G D G D V A L G I G L D S E E Q G - V N PG T - H P H A S A H V A H S S G T A L A A T L N D T S T S A I N - P G I G I G A S P T T A G K T A D
. . A G M A H H . . M . N M V A M . D . A I G I G . S . . G P
CtaCI-7002CtaCI-6803CtaCI-7120
210 220 230 240 250L V V D V N G L Q Y A WI F T Y P D T G I V S G D L H V P V D R P I Q L N M K A A D V I H A F WL PL M V D V K G I Q Y A WI F T Y P E T G I I S G E L H A P I D R P V Q L N M E A G D V I H A F WI PL V V N V T G M Q F A WX F D Y P D N G V S A G E L H V P V G A D V Q L N L S A Q D V I H S F WV PL V V D V G . Q Y A WI F T Y P D T G I . S G E L H V P V D R P V Q L N M A . D V I H A F W. P
CtaCI-7002CtaCI-6803CtaCI-7120
260 270 280 290 300E F R I K Q D V M P G Q V S Q L S F V A N R E G T Y P V I C A E L C G S Y H G G M K T T M T V E T AQ L R L K Q D V I P G R G S T L V F N A S T P G Q Y P V I C A E L C G A Y H G G M K S V F Y A H T PQ F R L K Q D A I P G V P T E L R F V A T K P G T Y P V V C A E L C G G Y H G S M R T Q V I V H T PQ F R L K Q D V I P G . S L F V A . . P G T Y P V I C A E L C G . Y H G G M K T V H T P
CtaCI-7002CtaCI-6803CtaCI-7120
310 320 330 340 350E G Y D Q WV Q S R T V A L Q D G E G Q P L P V D S T A L T D A E F L Q A Y A E E M G I I E N T L EE E Y D D WV A A N A P A P T E S M A M T L P K A T T A M T P N E Y L A P Y A K E M G V Q T E A L AE E F D S WL A E N Q V A Q Q Q N L H Q A V A V N P A N L S T S E F L A P H T Q D L G I S A A T L EE E Y D WV A N V A Q . . Q L P V . T A L T E F L A P Y A E M G I T L E
CtaCI-7002CtaCI-6803CtaCI-7120
360 370 380 390 400Q I P H H P L G M M S M A QQ L K D Q T S P V G D L LT L H T T S V NQ L . . . .
397
Figure B46. ClustalW alignment of CtaDI amino acid sequences. CtaDI amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
398
CtaDI ClustalW Amino Acid Alignment
CtaDI-7002CtaDI-6803CtaDI-7120
10 20 30 40 50M S D A T I H H A G D - - - - R - - - - - - - - - - - - - K WT D Y F T F C T D H K V I G I
M T I A A E N L T A N H P R R - - - - - - - - - - - - - - - - - - K WT D Y F T F C V D H K V I G IM T R V E F P P H I P P D D N Q P K N L A V G H G L T L P A WK WR D Y F T F N V D H K V I G I
. . T . H H P D . K WT D Y F T F C V D H K V I G I
CtaDI-7002CtaDI-6803CtaDI-7120
60 70 80 90 100Q Y L V T A F L F Y F I G G A L A E I V R T E L A T P D P D F V S P E V Y N Q M F T M H G T I M I FQ Y L V T S F L F F F I G G S F A E A M R T E L A T P S P D F V Q P E M Y N Q L M T L H G T I M I FQ Y L V T A F L F Y L I G G L M A I A I R T E L A T P D A D F I D P N L Y N A F M T N H G T I M I FQ Y L V T A F L F Y F I G G . A E A . R T E L A T P D P D F V P E . Y N Q M T H G T I M I F
CtaDI-7002CtaDI-6803CtaDI-7120
110 120 130 140 150L WI V P - A G A A F A N Y L I P L M I G A D D M A F P R L N A V A F WM Q P V G G I L L I L S F FL WI V P - A G A A F A N Y L I P L M V G T E D M A F P R L N A V A F WL T P P G G I L L I S S F FL WI V P S A I G G F G N Y L I P L M I G A R D M A F P K L N A I A F WL N P P A G L L L L L S F IL WI V P A G A A F A N Y L I P L M I G A . D M A F P R L N A V A F WL . P P G G I L L I L S F F
CtaDI-7002CtaDI-6803CtaDI-7120
160 170 180 190 200V G A P Q A G WT S Y P P L S L I S G K WG E E L WI L C L L I L G T A S I L G A I N F V T T I F KV G A P Q A G WT S Y P P L S L L S G K WG E E L WI L S L L L V G T S S I L G A I N F V T T I L KF G G S Q S G WT A Y P P L S L V T A P T A Q T L WI L A I V L V G T S S I L G S V N F V V T I L MV G A P Q A G WT S Y P P L S L . S G K WG E E L WI L L L L V G T S S I L G A I N F V T T I L K
CtaDI-7002CtaDI-6803CtaDI-7120
210 220 230 240 250M R A P D M D I H S M P L F C WA M L A T S A L I L L S T P V L A A A L I L L S F D L M A G T A F FM R I K D M D L H S M P L F C WA M L A T S S L I L L S T P V L A S A L I L L S F D L I A G T S F FM K V P S M K WD Q L P L F C WA I L A T S V L A L L S T P V L A A G L V L L L F D L N F G T S F FM R . P D M D . H S M P L F C WA M L A T S . L I L L S T P V L A A A L I L L S F D L A G T S F F
CtaDI-7002CtaDI-6803CtaDI-7120
260 270 280 290 300N P T G G G D P I V Y Q H L F WF Y S H P A V Y I M V L P F F G V I S E I L P V H S R K P I F G Y RN P V G G G D P V V Y Q H L F WF Y S H P A V Y I M I L P F F G V I S E V I P V H A R K P I F G Y RK P D A G G N V V I Y Q H L F WF Y S H P A V Y L M I L P I F G I M S E V I P V H A R K P I F G Y KN P G G G D P V V Y Q H L F WF Y S H P A V Y I M I L P F F G V I S E V I P V H A R K P I F G Y R
CtaDI-7002CtaDI-6803CtaDI-7120
310 320 330 340 350A I A Y S G L A I S F L G L I V WA H H M F T S G T P G WL R M F F M A T T M L V A V P T G I K I FA I A Y S S L A I S F L G L I V WA H H M F T S G T P G WL R M F F M A T T M L I A V P T G I K I FA I A Y S S V A I C V V G L F V WV H H M F T S G T P G WM R M F F T I S T L I V A V P T G V K I FA I A Y S S L A I S F L G L I V WA H H M F T S G T P G WL R M F F M A T T M L V A V P T G I K I F
399
CtaDI-7002CtaDI-6803CtaDI-7120
360 370 380 390 400S WC A T L WG G K L Q L N S A L L F A I G F L S S F L I G G L T G V M L A A A P F D I H V H D T YS WC G T L WG G K I Q L N S A M L F A F G F L S S F M I G G L T G V M V A S V P F D I H V H D T YG WV A T L WG G K I R F T S A M L F A I G L L S M F V M G G L S G V T M G T A P F D V H V H D T YS WC A T L WG G K I Q L N S A M L F A I G F L S S F . I G G L T G V M . A . A P F D I H V H D T Y
CtaDI-7002CtaDI-6803CtaDI-7120
410 420 430 440 450F V V G H F H Y V L F G G S V F A L F G A V Y H WF P K M T G K M Y N E T WG K I H F A M T F I G FF V V G H F H Y V L F G G S A F A L F S G V Y H WF P K M T G R M V N E P L G R L H F I L T F I G MY V V A H F H Y V L F G G S V F G I Y A G I Y H WF P K M T G R K L G E G WG R I H F A L T L V G TF V V G H F H Y V L F G G S V F A L F . G V Y H WF P K M T G R M . N E WG R I H F A L T F I G
CtaDI-7002CtaDI-6803CtaDI-7120
460 470 480 490 500N M T F L P M H Y L G L Q G M N R R I A L Y D P Q F Q P L N Q V C T L G S Y I L A L S T L P F L V SN L T F M P M H E L G L M G M N R R I A L Y D V E F Q P L N V L S T I G A Y V L A A S T I P F V I NN L T F L P M H K L G L Q G M P R R V A M Y D P Q F V D L N V L C T I G A F I L G L S V I P F A I NN L T F L P M H L G L Q G M N R R I A L Y D P Q F Q P L N V L C T I G A Y I L A L S T I P F . I N
CtaDI-7002CtaDI-6803CtaDI-7120
510 520 530 540 550I V L G L V N G K A A G R N P WR A L T L E WQ T T S P P S I E N F D E P P V L WA G P Y E Y G I DV F WS L F K G E K A A R N P WR A L T L E WQ T A S P P I I E N F E E E P V L WC G P Y D F G I DV I WS WS K G E L A G D N P WE A L S L E WT T S S P P L V E N WE V L P V V T H G P Y D Y G H SV . WS L K G E . A G R N P WR A L T L E WQ T . S P P . I E N F E E P V L W G P Y D Y G I D
CtaDI-7002CtaDI-6803CtaDI-7120
560 570 580 590 600G E P R D - E D S I E E M L A E V A E M ST E L M D D E E T V Q T L I A D A A G SL E A A P - E V S V S T. E . D E . S V T . A . . A
400
Figure B47. ClustalW alignment of CtaEI amino acid sequences. CtaEI amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
CtaEI ClustalW Amino Acid Alignment
CtaEI 7002CtaEI-6803CtaEI-7120
10 20 30 40 50MQ S S T T N T A I A T D Y Q T P P T - - - - - E
MT S P MG P T A R D F WQ N S R N S A T I Q R L I V S P MQ T T A L S S D L N S T Y T P G E A H GMT S V T A H E A H G - - - - - - - - - - - - - GM S T T A . . . D . . T G
CtaEI 7002CtaEI-6803CtaEI-7120
60 70 80 90 100H H G H P D Y R MF G L V MF L V A E S MI F L G L F A A F L I Y K A T MP G F - - - - T K E L E LH H G H P D L R MF G V V L F L V A E S A I F L G L F T A Y L I Y R S V MP A WP P E G T P E L E LH E A H P D L R V WG L L T F L I S E S L MF G G F F A T Y L F F K G T T E V WP P E - G T E V E LH H G H P D L R MF G L V F L V A E S . I F L G L F A A Y L I Y K . T MP . WP P E T E L E L
CtaEI 7002CtaEI-6803CtaEI-7120
110 120 130 140 150L V P T V N T I I L V S S S F V MH K G Q S A I K N D D V K G L Q L WF G I T A L MG A V F L G G QL L P G V N S I I L I S S S F V MH K G Q A A I R N N D N A G L Q K WF G I T A A MG I I F L A G QF V P A I N T A I L L S S S V V I H F G D MA I K K G N V WG MR I WY F L T A I MG A V F L A G QL V P . V N T I I L . S S S F V MH K G Q A I K N D V G L Q . WF G I T A . MG A V F L A G Q
CtaEI 7002CtaEI-6803CtaEI-7120
160 170 180 190 200V Y E Y A H ME F G L T E H L F G S C F Y V L T G F H G L H V T A G L L F T L A V L WR S R E A G HMY E Y F H L E MG L T T N L F A S C F Y V L T G F H G L H V T F G L L L I L S V L WR S R Q P G HV Y E Y Q N L G Y G L T A N V F A N C F Y I MT G F H G L H V F I G L L L I L G V L WR S R R S G HV Y E Y H L E . G L T N L F A S C F Y V L T G F H G L H V T . G L L L I L . V L WR S R G H
CtaEI 7002CtaEI-6803CtaEI-7120
210 220 230 240 250Y S G Q A H F G V E A A E L Y WH F V D V V WN Y P V R P C LY S R T S H F G V E A A E L Y WH F V D V V WI V L F I L V Y L LY S E T K H T G I E MA E I Y WH F V D I I WI V L F T L V Y L L N L LY S T H F G V E A A E L Y WH F V D V V WI V L F L V Y L L
401
Figure B48. ClustalW alignment of CtaCII amino acid sequences. CtaCII amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
402
CtaCII ClustalW Amino Acid Alignment
CtaCII-7002CtaCII-6803CtaCII-7120
10 20 30 40 50M K L K T I I S L S A I A L L L G V F S Y WV G Q WS Y S WL P P Q A S V E S Q L V D R L F S T L VM S R K N L I L L A V Y I V F T V G A S L WL G Q R A Y Q WL P P A A A Q E A Q P V G D L F S F L V
N I V T L V I G A I A V T I T S F WI G K L A Y T WL P P Q A A A E S I L I D D L F S F L VM K N I I . L . . . A . . . . . S . W. G Q A Y . WL P P Q A A . E S Q L V D D L F S F L V
CtaCII-7002CtaCII-6803CtaCII-7120
60 70 80 90 100A I G T F I L F G V A G T M T Y S I L F H R A G R Y D T S D G P P I E G N V T L E I V WT T I P F LS L G S V V F L G V A G A M A Y S V I F H R F S L Q N P Q - G A P I R G N A R L E I F WT V V P I IT M G A F I F L G V T S T L F Y S L L F H R A A E N D L S D G P H I E G N V T L E V V WQ A I P I L. . G . F I F L G V A G T M Y S . L F H R A . . D S D G P P I E G N V T L E I V WT . I P I L
CtaCII-7002CtaCII-6803CtaCII-7120
110 120 130 140 150I V I Y L A Y F S Y Q T Y R E M N I Q A P G - - - - - - H M H Q E A L V T N V S T G K T T M A M P VL V T WI A WY S Y V I Y Q R M N V L G P L P V V E V P Q L L G E K A I A A D A P A E L A M A Q G SL V V WI A T Y S Y Q I Y E Q M G I Q G R T - - - A L V H L H N P M E M E S A Y A A T E D G L V E PL V . WI A . Y S Y Q I Y M N I Q G P . . H L H . E . . . A M A
CtaCII-7002CtaCII-6803CtaCII-7120
160 170 180 190 200T N - - - - V E V T A K Q WA WI F H Y P E K N V T S T E L H L P V N Q R A H F I L R S P D V I H GN I N P E R I G V E V K Q WL WT F T Y P N G G V T S H E L H L P L D R R V T L N M T S K D V L H GE E - - - K I D V I A K Q WA WV F H Y P E K N V T S T E L H L P S D R R V K L A L H S E D V L H G
. I . V A K Q WA W. F H Y P E K N V T S T E L H L P . D R R V . L . L . S D V L H G
CtaCII-7002CtaCII-6803CtaCII-7120
210 220 230 240 250F F I P A F R V K Q D V I P F E D T D F E F T P I K T G K Y R I R D S Q F S G T Y F A A M Q A D V VF Y V P N F R I K Q D I V P N R E I E F S F T P N R L G E Y K L H D S Q F S G T Y F A V M T A P V VF Y I P A F R L K Q D I I P N H N I D F E F T P I R E G K Y H L T D S Q Y S G T Y F A T M Q A N V VF Y I P A F R . K Q D I I P N . . I D F E F T P I R G K Y . L . D S Q F S G T Y F A . M Q A V V
CtaCII-7002CtaCII-6803CtaCII-7120
260 270 280 290 300V E S Q E D Y Q T WL N Q A A R Q P P T P A P N Q A V S E Y A R R Q Q K E D R A A WK T V P P A P PV Q S L S D Y Q A WL E S Q K S L T P G E L P N P A L D E F K Q T P T T P L K S G WP T V P P G T RV E S P E E Y H K WL A K I A T H K P G T A Y N Q A S A E Y A Q S I T Q Q V K T G WK T V A PV E S E D Y Q WL . A . P G A P N Q A . E Y A Q . T . K . G WK T V P P .
CtaCII-7002CtaCII-6803CtaCII-7120
310 320 330 340 350P V V N Y H PQ
403
Figure B49. ClustalW alignment of CtaDII amino acid sequences. CtaDII amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
404
CtaDII ClustalW Amino Acid Alignment
CtaDII-7002CtaDII-6803CtaDII-7120
10 20 30 40 50MT Q A P P E A L T N E G Q P L T N L P - - N WR T F F S F S T D H K V I G L Q Y I V T S F F F F LMT N Q P T A S V P F Q R P - - - - - - - - - WH D Y L K F S T D H K V I G I Q Y L L MS F C F F LMT N I P I E G V Q L P E G K P H H P S P G G WK E Y F S F S H D H K V I G I Q Y L V T S F I F F LMT N . P E . V W. . Y F S F S T D H K V I G I Q Y L V T S F F F L
CtaDII-7002CtaDII-6803CtaDII-7120
60 70 80 90 100V A G I F A MI MR G E L I T P E P D L V D R T V Y N A L F T MH G S I ML F G WT F P V L A G F AV A G L L A MI I R A E L L T P Q L D V V D R S L Y N G L F T L H G T I MI F L WI F P A N V G L AV G G I F A MI L R G E L I T P E S D L I D R T V Y N G MF T MH G T V ML F L WT F P S L V G L AV A G I F A MI . R G E L I T P E D L V D R T V Y N G L F T MH G T I ML F L WT F P . L V G L A
CtaDII-7002CtaDII-6803CtaDII-7120
110 120 130 140 150N Y L V P L MI G A Q D MA F P R L N A I A F WMV P V F G S L L V L S F L A P G G P A Q A G WWSN Y L I P L MI G A R D V A F P V L N A I A F WL MP V V G V L L I G S F F L P T G T A Q A G WWSN Y L V P L MI G A R D MA F P R L N A A A F WMV P V V G I L L MT S F F V P G G P A Q S G WWAN Y L V P L MI G A R D MA F P R L N A I A F WMV P V V G . L L . . S F F . P G G P A Q A G WWS
CtaDII-7002CtaDII-6803CtaDII-7120
160 170 180 190 200Y P P V S I Q N P A G V L L N G E F L WL V S V A L S G I S S I L G A V N I V T T I V Q MR C P G MY P P V S I Q N P S G N F I N G E F L WL L A V A T S G I S S I MG G V N F V T T V WK L R A P G LY P P V S T Q N P T G N L I N G Q V I WL L A V A I S G V S S I MG A V N F V T T I V K MR A P G MY P P V S I Q N P . G N L I N G E F L WL L A V A . S G I S S I MG A V N F V T T I V K MR A P G M
CtaDII-7002CtaDII-6803CtaDII-7120
210 220 230 240 250G WF K T P A F V WT V L A A Q I I Q L F G L P A L T A G A V ML L F D L T V G T T F F D A S Q G GT L L K MP V Y V WT I L S A Q I L Q L F C L P A L T G G A V ML L F D L S F G T T F F D P F N Q GG F F R MP L F V WA V F S A Q I I Q L F G L P A L T A G A V ML L L D I T V G T S F F D P S K G GG . F K MP . F V WT V L S A Q I I Q L F G L P A L T A G A V ML L F D L T V G T T F F D P S . G G
CtaDII-7002CtaDII-6803CtaDII-7120
260 270 280 290 300S P V L Y Q H F F WF Y S H P A V Y V MV L P V F G F F S E L F P V Y A R K P L F G Y K V V V V S SN P I I Y Q H L F WF Y S H P A V Y V MA L P A F G V F S E I L P V F A R K P L F G Y K T V A I S SN P V MF Q H Y F WF Y S H P A V Y V I I L P I F G I F S E I F P V Y S R K P L F G Y K V V A I S SN P V . Y Q H . F WF Y S H P A V Y V M. L P . F G . F S E I F P V Y A R K P L F G Y K V V A I S S
CtaDII-7002CtaDII-6803CtaDII-7120
310 320 330 340 350MI I V V L S A V V WV H H MF A S G T P P WMR ML F MF S T ML I S V P T G I K V F A WV A T LF L I A I Q G T F V WV H H MF T S A T P N WMR MF F MA S T ML I A V P T G I K V L A WT A T VML I A V V S A I V WV H H L Y V S G T P A WMR MF F ML T T ML V S V P T G I K V F A WV A T IML I A V . S A . V WV H H MF . S G T P WMR MF F M. S T ML I S V P T G I K V F A WV A T .
CtaDII-7002CtaDII-6803CtaDII-7120
360 370 380 390 400WG G K L R L D T P ML F A MG G L I N F V F A G I T G I ML A S V P V D I H V N N T Y F V V G H FWR G S L R L K T P ML F C L G G I L MF L F A G I T G I ML A S A P F D L H V N N T Y F V V G H FWG G K I R L N T P ML F A L G G L I L F V F A G I V G I ML S S V P V D V H V N N T Y F V V G H FWG G K L R L T P ML F A L G G L I F V F A G I T G I ML A S V P V D . H V N N T Y F V V G H F
CtaDII-7002CtaDII-6803CtaDII-7120
410 420 430 440 450H Y V I Y G A I V F G I Y G A V Y H WF P K MT G K MY Y E G L G K L H F WL T MI G T T L N F L PH Y V V F G T V T MA I Y G A I Y F WF P K MT G R MY N E A WG K L H F A L T F I G A N L N F F PH Y V L F G T V T MG MY A A I Y H WF P K MT G R MY Y E G WG K L H F WL T F I G T N L N F F PH Y V . F G T V T MG I Y G A I Y H WF P K MT G R MY Y E G WG K L H F WL T F I G T N L N F F P
CtaDII-7002CtaDII-6803CtaDII-7120
460 470 480 490 500MH P V G L MG MP R R V A S Y D P E F A F WN V I A S I G G F I L G MS T I P F L L N MI A S WIMH P I G L Q G ML R R I S S Y D P E Y T A WN V V A S L G A F L L G MS T L P F I A N MV A S A FMH P L G L Q G ML R R V S S Y A P E Y E G WN I V A S L G A F L L G MS T L P F I F N MV V S WMMH P . G L Q G ML R R V S S Y D P E Y . WN V V A S L G A F L L G MS T L P F I . N MV A S W
CtaDII-7002CtaDII-6803CtaDII-7120
510 520 530 540 550N G D R A P A N P WR A I G L E WL V P S P P E H E N F E E L P I V I A E P Y G Y G K S E P L T E NQ G R R V G N N P WN S L G L E WT T P S P P P E E N F E V I P T I T V E P Y G Y D R P V D L T T EH G E K A P D N P WR A I G L E WL V A S P P P V E N F E E I P V V I S E P Y G Y G K S E P L T A E. G . R A P N P WR A I G L E WL V P S P P P E N F E E I P . V I . E P Y G Y G K S E P L T E
CtaDII-7002CtaDII-6803CtaDII-7120
560 570 580 590 600L G GV T T
.
405
Figure B50. ClustalW alignment of CtaEII amino acid sequences. CtaEII amino acid
sequences of the following organisms were aligned using the ClustalW alignment
program from MacVector v. 6.5: Synechococcus sp. PCC 7002, Synechocystis sp. PCC
6803, and Anabaena sp. PCC 7120. The consensus amino acid sequence is shown
underneath. Gray high-lighted regions indicate identical amino acids while lighter
shading indicates conserved amino acids. Dashes represent insertions/deletions included
to maximize the sequence similarity. Percent identity and percent conserved amino acids
determined by the ClustalW search program (Thompson et al., 1994b).
CtaEII ClustalW Amino Acid Alignment
CtaEII-7002CtaEII-6803CtaEII-7120
10 20 30 40 50M T A I N E T P I S A T S H G H G E E D H R L F G F I V F L L S E S V I F I S F F V G Y I V Y K L SM E S G N H L P H V E P T E E Q - E P D N L G F G F P V F L M S E S V V F I S F F V T Y T I L R L T
G H E I S H E H G H D E E G N K M F G F I V F L L S E S V I F L S F F A G Y I V Y K T TM . N . P . . H G H E E D N . . F G F I V F L L S E S V I F I S F F V G Y I V Y K L T
CtaEII-7002CtaEII-6803CtaEII-7120
60 70 80 90 100P T D WL P P G V E G L E I H D P A I N T V V L V S S S G V I Y L A E R F L H K E N L WG F R F F WN K P WF P P G V D G L D V T R G A I N T M V L V T S S G A I I L A E K A L H R G E M K L F R L L WT P N WL P V G V E G L E V R D P A I N T V V L V A S S F V I Y F A E L A L K R Q N L R L F R I F L
WL P P G V E G L E V . D P A I N T V V L V . S S G V I Y L A E . A L H R N L . L F R . F W
CtaEII-7002CtaEII-6803CtaEII-7120
110 120 130 140 150L L T M A M G S Y F L Y G Q A V E WQ S L E F E F T S G V Y G G I F Y L L T G F H G L H V L T G V LL A T I S L G I V F L F G Q A A E WA G M P F G L D A G S A G G T F F L L T G F H G L H V F T G V CS A T M A M G S Y F L V G Q A I E WS H L E F G F T S G V Y G G M F Y L L T G F H G L H V F T G I LL A T M A M G S Y F L . G Q A . E W L E F G F T S G V Y G G F Y L L T G F H G L H V F T G V L
CtaEII-7002CtaEII-6803CtaEII-7120
160 170 180 190 200L Q G V M L G R S F L P N N Y A G G Q Y G V E A T S WF WH F V D V I WI I L F G L I Y L WQL L L Y M Y WR S L Q P H N F D R G H E G V T A I A L F WH F V D V I WI I L F I L L Y L WP A NL Q F I I L V R S L I P G N Y D T G H F G V N A T S L F WH F V D VL Q . . M L . R S L . P N Y D G H . G V A T S L F WH F V D V I WI I L F . L . Y L W
CHRISTOPHER T. NOMURA
EDUCATION1988-1989 Laney Junior College, Oakland1989-1994 B. A., Biology with Honors, University of California, Santa Cruz1994-2001 Ph.D., Biochemistry and Molecular Biology, The Pennsylvania
State University
HONORS/PROFESSIONAL TRAINING
1994-1996 NIH Biotechnology Predoctoral Fellow, Dept. of Biochemistry and Molecular Biology, The Pennsylvania State University
1994-1996 Graham Endowed Graduate Fellowship, Dept. of Biochemistry and MolecularBiology, The Pennsylvania State University
1995 Teaching Assistant, Dept. of Biochemistry and Molecular Biology, ThePennsylvania State University
1995 NSF Predoctoral Fellow, Honorable Mention
1994 MIRT Fellow, Laboratorio de Mamiferos Marinos (Dr. Claudino Campagna) and University of California-Santa Cruz (Dr. C. Leo Ortiz)
1993 MARC Undergraduate Fellow, University of California-Santa Cruz, (Dr. C. LeoOrtiz)
1992-1993 MBRS Fellow, University of California-Santa Cruz, (Dr. Barry Bowman)
PUBLICATIONS1. Nomura, C. T. and D. A. Bryant. 1997. Characterization of cytochrome c6 from
Synechococcus sp. PCC 7002, p. 269-274. In G.P. Peschek, W. Löffelhardt, and G.Schmetterer (ed.), The Phototrophic Prokaryotes. Kluwer Academic/PlenumPublishers, New York, New York.
2. Huckauf, J., Nomura, C. T., Forchhammer, K., and M. Hagemann. 2000. Stressresponses of Synechocystis sp. strain PCC 6803 mutants impaired in genes encodingputative alternative sigma factors. Manuscript submitted to Microbiology.
3. Nomura, C. T., Persson, S., Inoue-Sakamoto, K., Sakamoto, T., Shen, G., and D. A.Bryant. 2001. A role for cytochrome oxidase in high-light and oxidative stressresponse in the cyanobacterium Synechococcus sp. PCC 7002. Manuscript inpreparation.
4. Nomura, C. T., Persson, S., and D. A. Bryant. 2001. Cloning and sequence analysisof electron transport protein genes from Synechococcus sp. PCC 7002: A comparativestudy of electron transport proteins from cyanobacteria and chloroplasts. Manuscriptin preparation.