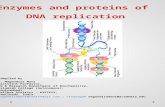
Dna replication and enzymes involved in dna replication
-
Upload
narayan-prahlad -
Category
Science
-
view
141 -
download
2
Transcript of Dna replication and enzymes involved in dna replication

DNA REPLICATIONDNA REPLICATION
Guided By-Dr.Cletus D’SouzaDept. Of Biochemistry
Presented By-Narayan Prahlad VVIth SemesterDept.of Molecular Biology

ContentsContents
Introduction Chemistry of DNA Synthesis Extension of 3’-OH of Primer Enzymes involved in replication Mechanism of Replication Proof Reading Conclusion Reference Acknowledgement

INTRODUCTIONINTRODUCTION

Chemistry of DNA SynthesisChemistry of DNA Synthesis
• Requires 4 Deoxynucleoside Triphosphates (dGTP,dCTP,dATP,dTTP).• Have three Phosphate groups-attached via 5’-OH & 2’ deoxyribose.
• Second essential substrate- Primer:Template Junction• Primer- Should have exposed 3’-OH.

Extension of 3’-OH of PrimerExtension of 3’-OH of Primer
• New chain grows-extension of 3’ end of primer-Phosphodiester bond is formed by SN2 mechanism.

EnzymesEnzymes
DNA Polymerase-• Mainly catalyses DNA synthesis.• Multimolecular forms of this enzyme- DNA Pol-I, DNA Pol-II, DNA
Pol-III DNA Pol-I: Polymerase Activity. 5’ to 3’ Exonuclease Activity-Used to remove primer and replace with DNA. DNA Pol-III: Core replication enzyme. Holoenzyme Has proof reading capabilities(3’ to 5’ exonuclease activity).

Polymerase Polymerase
clamp loader
Sliding clamp
3'-5' exonuclease

DNA Pol-III
• The fingers and the thumb are composed-helices. • The incoming dNTP is in red (for the base and the deoxyribose) and yellow (for the triphosphate moiety). • The template strand of the DNA is shown in dark gray, and the primer strand is shown in light gray.

Continued..Continued..
Helicase-• Hexameric assembly of 6 identical subunits.• Uses ATP (Hydrolysis)-unwind DNA.
Sliding Clamp-• Part of DNA Pol-III (β-Subunit).• Encircles and slides along DNA-tethering it to
template.

Continued..Continued.. Single Strand DNA Binding Protein (SSB)-• Tertamer of identical subunits.• Binds to ssDNA-preventing it from recoiling.
DNA Primase-• Synthesis of Oligoribonucleotide(Primer) -extended by DNA Pol-III.
Ligase-• Seals the gaps between Okazaki fragments-forming a
continuous strand.

Mechanism of ReplicationMechanism of Replication
STEP-1 -UnwindingUnwinding of DNA by Helicase of DNA by Helicase

• Step-2- Binding of SSBs to ssDNABinding of SSBs to ssDNA

• Step-3
Binding of TopoisomeraseBinding of Topoisomerase

• Step-4Action of DNA Pol-IIIAction of DNA Pol-III

Sliding ClampSliding Clamp

Thrombone ModelThrombone Model
• In order to explain synthesis on lagging strand-Thrombone model was proposed.

Synthesis of DNA on Lagging Synthesis of DNA on Lagging StrandStrand



Action of DNA Pol-I & LigaseAction of DNA Pol-I & Ligase
• DNA Pol-I replaces RNA fragments with DNA and fills the gaps.
• Ligase joins the fragments –Phosphodiester bonds and forms a continuous strand.

Proof ReadingProof Reading
• DNA Pol-3.
• When a mismatched nucleotide comes-rate of addition of nucleotide by polymerase slows down.
• Exonuclease catalytic site in DNA Pol-3.
• Error rate- 1 in 1010 bp added, occasionally- 1 in 105 .


ConclusionConclusion
• DNA replication is a very important phase in cell cycle.• Replication is a must so as to ensure the equal distribution of
genetic material to daughter cells.• Thus all enzymes have to coordinate well in order for
replication to occur successfully.

ReferencesReferences
• Molecular Biology of Gene by Watson;7th edition; Cold Spring Harbour Laboratory Press, Cold Spring Harbour New York.
• Biochemistry 5th edition by Lubert Stryer;W.H Freeman Company.
• Molecular Cell Biology by Lodish;6th edition;W.H Freeman and Company;New York.
• http://sites.fas.harvard.edu/~biotext/animations/replication1.swf

AcknowledgementAcknowledgement
• I would like thank my guide Dr.Cletus D’Souza for his valuable guidance.
• I would also like to thank our course coordinator Dr.N.S Devaki for providing me this opportunity to present this seminar.
Thank You all for your patient listening.