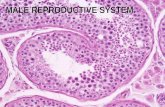
Cross section of rat testis Showing Seminiferous Tubules and Interstitium
description
Transcript of Cross section of rat testis Showing Seminiferous Tubules and Interstitium

Kent Christensen, Univ. Michigan
Cross section of rat testisCross section of rat testisShowing Seminiferous Showing Seminiferous
Tubules and Interstitium Tubules and Interstitium
Cross section of rat testisCross section of rat testisShowing Seminiferous Showing Seminiferous
Tubules and Interstitium Tubules and Interstitium

Functional and Anatomical Functional and Anatomical Compartments of the TestisCompartments of the TestisFunctional and Anatomical Functional and Anatomical
Compartments of the TestisCompartments of the Testis

Scott Miller, Univ Utah
Interstitium of rat testis Interstitium of rat testis showing endothelium, showing endothelium, Leydig cells (L), and Leydig cells (L), and
macrophages (arrow). macrophages (arrow). Note close association of Note close association of
macrophages and macrophages and Leydig cells. Leydig cells.
Interstitium of rat testis Interstitium of rat testis showing endothelium, showing endothelium, Leydig cells (L), and Leydig cells (L), and
macrophages (arrow). macrophages (arrow). Note close association of Note close association of
macrophages and macrophages and Leydig cells. Leydig cells.

Scott Miller, Univ. Utah
Close association of Close association of Leydig cell and Leydig cell and
macrophage, lower panel macrophage, lower panel shows close up of shows close up of
“digitation” of Leydig cell “digitation” of Leydig cell process extending onto process extending onto macrophage surface.macrophage surface.
Close association of Close association of Leydig cell and Leydig cell and
macrophage, lower panel macrophage, lower panel shows close up of shows close up of
“digitation” of Leydig cell “digitation” of Leydig cell process extending onto process extending onto macrophage surface.macrophage surface.

Cytokines, ROS
?
Macrophage-Leydig cell interactionsMacrophage-Leydig cell interactionsMacrophage-Leydig cell interactionsMacrophage-Leydig cell interactions

cholesterol
Extracellularlipoprotein
Cholesterolpool
LH
ATP
cAMPPKA+
Pregnenolone
Progesterone
Androstenedione
TESTOSTERONE
m
3HSD
P450c17
17HSD
acetate
++
Tra
nsc
rip
tion

• Of all the steroidogenic enzymes, P450c17 is the most sensitive to repression
• Most cytokines tested inhibit c17 transcription: – IL-1, IL-2, IL-6, TNF, TGF, INF, INF
• Inflammatory mediators: PGF2, ceramide, vasopressin, PKC agonists
• Environmental disruptors such as dioxin, pthalates, PAHs, etc. are inhibitory
• Androgen-mediated feedback repression
P450c17 is sensitive to P450c17 is sensitive to transcriptional repression transcriptional repression
P450c17 is sensitive to P450c17 is sensitive to transcriptional repression transcriptional repression

IL-1, TNF and PMA vs. IL-1, TNF and PMA vs. Testosterone productionTestosterone productionIL-1, TNF and PMA vs. IL-1, TNF and PMA vs. Testosterone productionTestosterone production
0
500
1000
1500
20002500
3000
3500
4000
4500
con
cAM
P
cAM
P+IL1
cAM
P+TNF
cAM
P+PMA
ng
/10
6 LC
/24
h

IL-1, TNFIL-1, TNF and PMA vs. and PMA vs. steroidogenic mRNA expressionsteroidogenic mRNA expression
IL-1, TNFIL-1, TNF and PMA vs. and PMA vs. steroidogenic mRNA expressionsteroidogenic mRNA expression
cAMP+IL-1
+TNF+PMA
P450c17
P450scc

cAMP responsive regions of the cAMP responsive regions of the CypCyp1717 promoterpromoter
cAMP responsive regions of the cAMP responsive regions of the CypCyp1717 promoterpromoter
0
20
40
60
80
100
-2500 -1021 -346 -245
Rel
ativ
e C
AT
Act
ivity
Control
cAMP

TNF inhibits Cyp17 promoter activityTNF inhibits Cyp17 promoter activityTNF inhibits Cyp17 promoter activityTNF inhibits Cyp17 promoter activity

Calphostin reverses TNF inhibition of Calphostin reverses TNF inhibition of Cyp17 promoterCyp17 promoter
Calphostin reverses TNF inhibition of Calphostin reverses TNF inhibition of Cyp17 promoterCyp17 promoter

TNF and PMA stimulate translocation of PKC from cytoplasm to membrane
TNF and PMA stimulate translocation of PKC from cytoplasm to membrane
control
PMA TNF
No antibody

Putative transfactor binding sitesPutative transfactor binding sitesPutative transfactor binding sitesPutative transfactor binding sites

-436 mouse GTGACCTTAT GCAAACTAAC CCTAAAAGAC CTCTCTCTCC TCAACTATCA GATAATAAGA GTGACCTTAT GCCGACTAAC CTTTGAAGAT CTCTTTCTCC TCAACTGTCA GATAGTAAGA -447 rat
-376 mouse CTGAAGTCTC TTTGACAGCT TTGGCTAGCT GCAACCTGAT GACATTAATT ATTAACTGTG CTGCAGTCTC T--------- ---------- GAAACCCGAT GGCAGTAATT ATTAACCGTA -387 rat
-316 mouse CAGCACTTTT GACATTACAG CACGCACTCT GAAACCTTGA TCTTAATCTG ATAGCATTTG TAGCACTT T GACATTACA CACAGACTCT AAAACCTTGA TCTCACTCTG ATAGCATTTG -346 rat
-59 mouse CACGTCTTCAAGGTGA CTCGACGTCAAGGTGA -73 rat
Comparison of cAMP-responsive sequences in mouse and rat
Binding sites (ATF2/cjun-like, Steroidogenic factor 1, StF-IT-2, and StF-IT-1/COUP-TF1) are shown in bold color in the sequence for the species in which it was identified. Those that are conserved between species are underlined. Sequence differences are shown in blue for mouse and pink for rat.

Characterization of the Cyp17 Promoter Characterization of the Cyp17 Promoter Revealed a Region Between -245 and -346 Revealed a Region Between -245 and -346
Responsible for the Minimal cAMP Responsible for the Minimal cAMP Responsiveness of the GeneResponsiveness of the Gene
Characterization of the Cyp17 Promoter Characterization of the Cyp17 Promoter Revealed a Region Between -245 and -346 Revealed a Region Between -245 and -346
Responsible for the Minimal cAMP Responsible for the Minimal cAMP Responsiveness of the GeneResponsiveness of the Gene
0
20
40
60
80
100
-2500 -1021 -346 -245
Rel
ativ
e C
AT
Act
ivity
Control
cAMP

Site-directed mutagenesis of Site-directed mutagenesis of Cyp17Cyp17 CRR CRR
(-346 to –245)(-346 to –245)
Site-directed mutagenesis of Site-directed mutagenesis of Cyp17Cyp17 CRR CRR
(-346 to –245)(-346 to –245)
•Oligos were designed to place an XhoI once every ten base pairs within the 100 base pair CRR.
•This resulted in changing as few as three (mutant 6) to as many as six (mutant 1 and 7) of every ten nucleotides.
•Mutagenesis was performed with Altered Sites (Promega) and all mutants were verified by sequencing.

Cyclic-AMP induction of CRR mutantsCyclic-AMP induction of CRR mutantsCyclic-AMP induction of CRR mutantsCyclic-AMP induction of CRR mutants

cAMP Induction of CRR mutantscAMP Induction of CRR mutantscAMP Induction of CRR mutantscAMP Induction of CRR mutants
0
50
100
150
200
250
Wild
Type
Mutan
t 1
Mutan
t 2
Mutan
t 3
Mutan
t 4
Mutan
t 5
Mutan
t 6
Mutan
t 7
Mutan
t 8
Mutan
t 9
% o
f Wild
Typ
e In
duct
ion
by c
AM
P

Putative sites revealed by Putative sites revealed by mutantsmutants
Putative sites revealed by Putative sites revealed by mutantsmutants
gcaacctgat gacattaatt attaactgtg cagcactttt gacattacag
CTCGAGtgat CTcGAGaatt CtCGaGtgtg cTCGaGtttt CTcGAGacag
mut 1 mut 2 mut 3 mut 4 mut 5
cacgcactct gaaaccttga tcttaatctg atagcatttg cctctgggag
cTcgAGctct CTCGAGttga CTCGaGtctg CtCgAGtttg cACGAgggag
mut 6 mut 7 mut 8 mut 9 mut 10
ATF2
AhR/Arnt (core sequence)
SF-1

-440 -250
ATF2 mutants 2,5,9
C/EBP– upstream site
AhR/ARNT mutant 6
SF-1 mutant 7
ARE
Putative regulatory motifs revealed by mutagenesis
?

The Minimal cAMP Responsive Region of The Minimal cAMP Responsive Region of the the Cyp17Cyp17 Promoter (CRR): Promoter (CRR):
The Minimal cAMP Responsive Region of The Minimal cAMP Responsive Region of the the Cyp17Cyp17 Promoter (CRR): Promoter (CRR):
-346 -245
-346 TGATGACAT TAATTATTAA CTGTGCAGCA
CTTTTGACAT TACAGCACGC ACTCTGAAAC
CTTGATCTTA ATCTAGCATT TGCCTCTGGG
AGGATCCATA GCG -245
Putative ATF-2 binding site

Binding of Nuclear Proteins to the CRR Probe is Binding of Nuclear Proteins to the CRR Probe is Augmented by Treatment of MA-10 Cells with cAMPAugmented by Treatment of MA-10 Cells with cAMP
Binding of Nuclear Proteins to the CRR Probe is Binding of Nuclear Proteins to the CRR Probe is Augmented by Treatment of MA-10 Cells with cAMPAugmented by Treatment of MA-10 Cells with cAMP

Nuclear Proteins from Primary Leydig Cells Nuclear Proteins from Primary Leydig Cells Form Two Complexes with the CRR ProbeForm Two Complexes with the CRR Probe
Nuclear Proteins from Primary Leydig Cells Nuclear Proteins from Primary Leydig Cells Form Two Complexes with the CRR ProbeForm Two Complexes with the CRR Probe

The Upstream ATF-2 C/EBPThe Upstream ATF-2 C/EBP Binding Site Binding SiteThe Upstream ATF-2 C/EBPThe Upstream ATF-2 C/EBP Binding Site Binding Site
-450 TTGTGTGACC TTATGCAAAC TAACCCA -423
-450 -245

Nuclear Proteins from Control and cAMP-Nuclear Proteins from Control and cAMP-Treated MA-10 cells Bind to the Upstream Treated MA-10 cells Bind to the Upstream
ATF-2 C/EBPATF-2 C/EBP Probe Probe
Nuclear Proteins from Control and cAMP-Nuclear Proteins from Control and cAMP-Treated MA-10 cells Bind to the Upstream Treated MA-10 cells Bind to the Upstream
ATF-2 C/EBPATF-2 C/EBP Probe Probe

Incubation of the Upstream ATF-2 C/EBPIncubation of the Upstream ATF-2 C/EBP probe probe with Nuclear Proteins Isolated from Primary with Nuclear Proteins Isolated from Primary Leydig Cells Results in Complex FormationLeydig Cells Results in Complex Formation
Incubation of the Upstream ATF-2 C/EBPIncubation of the Upstream ATF-2 C/EBP probe probe with Nuclear Proteins Isolated from Primary with Nuclear Proteins Isolated from Primary Leydig Cells Results in Complex FormationLeydig Cells Results in Complex Formation

Formation of the Higher Order Complex Formed Formation of the Higher Order Complex Formed by the CRR Complex is Decreased by Addition of by the CRR Complex is Decreased by Addition of Unlabeled ATF-2 or C/EBPUnlabeled ATF-2 or C/EBP Competitor Oligos Competitor Oligos
Formation of the Higher Order Complex Formed Formation of the Higher Order Complex Formed by the CRR Complex is Decreased by Addition of by the CRR Complex is Decreased by Addition of Unlabeled ATF-2 or C/EBPUnlabeled ATF-2 or C/EBP Competitor Oligos Competitor Oligos

Binding of Nuclear Proteins to the Upstream Binding of Nuclear Proteins to the Upstream ATF-2 C/EBPATF-2 C/EBP Probe can be Inhibited by Probe can be Inhibited by
Addition of ATF-2 or C/EBPAddition of ATF-2 or C/EBP Competitor Oligos Competitor Oligos
Binding of Nuclear Proteins to the Upstream Binding of Nuclear Proteins to the Upstream ATF-2 C/EBPATF-2 C/EBP Probe can be Inhibited by Probe can be Inhibited by
Addition of ATF-2 or C/EBPAddition of ATF-2 or C/EBP Competitor Oligos Competitor Oligos

Overexpression of C/EBPOverexpression of C/EBP Induces Induces Transcription of the -491/-255 Transcription of the -491/-255 CypCyp 17 reporter 17 reporter
Overexpression of C/EBPOverexpression of C/EBP Induces Induces Transcription of the -491/-255 Transcription of the -491/-255 CypCyp 17 reporter 17 reporter
0
1
2
3
4
5
6
7
Cyp 17 Cyp 17 + ATF-2 Cyp 17 +C/EBPb
Cyp 17 + ATF-2+ C/EBPb
Fol
d In
duct
ion
Control
+ cAMP
**
*

Western Analysis of ATF-2 Expression
0.000.200.400.600.801.001.201.40
12 hSFM
1 hcAMP
2 hcAMP
4 hcAMP
8 hcAMP
12 hcAMP
Fol
d In
duct
ion
ATF-2 Expression in MA-10 Cells is Not ATF-2 Expression in MA-10 Cells is Not Affected by cAMP TreatmentAffected by cAMP Treatment
ATF-2 Expression in MA-10 Cells is Not ATF-2 Expression in MA-10 Cells is Not Affected by cAMP TreatmentAffected by cAMP Treatment

p38
p34
Western Analysis of C/EBP Expression
0
2
4
6
8
10
12 hSFM
1 hcAMP
2 hcAMP
4 hcAMP
8 hcAMP
12 hcAMP
Fol
d In
duct
ion
**
C/EBPC/EBP Expression is Significantly Expression is Significantly Increased in MA-10 Cells with cAMPIncreased in MA-10 Cells with cAMPC/EBPC/EBP Expression is Significantly Expression is Significantly
Increased in MA-10 Cells with cAMPIncreased in MA-10 Cells with cAMP

Summary of Cyp17 studySummary of Cyp17 studySummary of Cyp17 studySummary of Cyp17 study
• TNF-mediated inhibition of transcription involves activation of PKC
• ATF2 and C/EBP participate cooperatively in cAMP-induction of transcription
• ATF2 is constitutively expressed
• C/EBP expression is induced by cAMP

HypothesesHypothesesHypothesesHypotheses
• ATF2 and C/EBP interact as heterodimers binding to the “ATF2” sites in the promoter
• The stoichiometry of C/EBP and ATF2 interaction is critical to driving transcription
• Repressors may act by inhibiting C/EBP expression or through post-translational modifications that inhibit its activity
• C/EBP phosphorylation by PKC may block it from interacting with ATF2

Effect of LPS on steroidogenic mRNA levelsEffect of LPS on steroidogenic mRNA levels Effect of LPS on steroidogenic mRNA levelsEffect of LPS on steroidogenic mRNA levels
P450scc
P450c17
3-HSD
actin
LPS - + - + - + - + - +
time 2h 4h 6h 8h 24h

0
2
4
6
8
10
12
14
LPS vs. serum testosterone: 2-24 hoursLPS vs. serum testosterone: 2-24 hoursLPS vs. serum testosterone: 2-24 hoursLPS vs. serum testosterone: 2-24 hours
Tes
tost
ero
ne
(ng
/ml)
control
LPS
Time post LPS
24 h2 h 4 h 8 h6 h

Steroidogenic Acute Steroidogenic Acute Regulatory Protein: StARRegulatory Protein: StAR
Steroidogenic Acute Steroidogenic Acute Regulatory Protein: StARRegulatory Protein: StAR
• Essential for steroid hormone biosynthesis• Cyclic-AMP dependent expression• Facilitates cholesterol transfer across inner-
mitochondrial (aqueous) space• Translated as a 37 kDa precursor protein that
is processed to the 30 kDa mature form as it translocates into the mitochondria
• Cholesterol transport activity depends on intact m

StAR facilitates cholesterol transferStAR facilitates cholesterol transfer StAR facilitates cholesterol transferStAR facilitates cholesterol transfer

Pulsatile nature of cholesterol Pulsatile nature of cholesterol flux into the mitochondriaflux into the mitochondria
Pulsatile nature of cholesterol Pulsatile nature of cholesterol flux into the mitochondriaflux into the mitochondria

StAR ProcessingStAR ProcessingStAR ProcessingStAR Processing
signal peptides
37 kDa
Outer mitochondrial membrane
Inner- mitochondrial membrane
critical regioncholesterol transfer
matrix
Cytosol
37
3230
Inner-mitochondrial forms
N'
32 kDaN'
30 kDaN'

sccAdx-red
adx
3 H SD
M itochondria lm atrix
C ytosolchol
cholchol
S tARN ’ C ’

sccAdx-red
adx
3 H SD
M itochondria lm atrix
C ytosol
chol
cholchol
S tARN ’ C ’
PBR

M ito c ho nd ria lm a trix
C yto so l
TO MTIM
PBRVD
AC
AN
T
HKC KC p hD

MTS1-37
ITS38-47
pCMV-StAR
StAR-stop
Tom20OMTS
StAR/CCHL
CCHLIMSS
StAR/Tom20
StAR -N47
StAR N-terminal localization expression clones
StAR -ITS
MTS1-37
TAA

What mediates the acute LPS What mediates the acute LPS inhibition?inhibition?
What mediates the acute LPS What mediates the acute LPS inhibition?inhibition?
• Tested numerous inflammatory mediators in Leydig cells in vitro-- none mimicked the acute LPS “effect”– cytokines (TNF, IL-1, IL-6, IFN, TGF)– prostaglandins (PGF2, PGE) – catecholamines (norepi, isoproteranol) – ceramide (C2, C8)– Most nitric oxide donors (Sin-1, SNAP, SNP, Nor-3)– Calcium inophore (A23187)

LPS vs. StAR protein LPS vs. StAR protein expression: 2 hr after injectionexpression: 2 hr after injection
LPS vs. StAR protein LPS vs. StAR protein expression: 2 hr after injectionexpression: 2 hr after injection
30 kDa
37 kDa
conLPS

Carbonyl cyanide Carbonyl cyanide mm--chlorophenylhydrazone (cccp)chlorophenylhydrazone (cccp)
Carbonyl cyanide Carbonyl cyanide mm--chlorophenylhydrazone (cccp)chlorophenylhydrazone (cccp)
• Carbonyl cyanide m-chlorophenyl-hydrazone (cccp): potent uncoupler of oxidative phosphorylation; protonophore, mitochondrial disrupter.
• Causes transient disruption of m

Mitochondrial respiration, OX-PHOS and Mitochondrial respiration, OX-PHOS and mm
Mitochondrial respiration, OX-PHOS and Mitochondrial respiration, OX-PHOS and mm
H+
e-
m

CCCP Vs. Progesterone in MA10s
0100200300400500
con cAMP cAMP +cccp
R22 R22 +cccp
ng
/ml

Effect of CCCP on StAR proteinEffect of CCCP on StAR proteinEffect of CCCP on StAR proteinEffect of CCCP on StAR protein
Control cAMP cAMP + cccp cccp
37 kDa
30 kDa

con cA cA+cccp
StAR
cyclophilin
3.4 kB
1.6 kB
2.9 kB
Effect of CCCP on StAR mRNAEffect of CCCP on StAR mRNAEffect of CCCP on StAR mRNAEffect of CCCP on StAR mRNA

Effect of CCCP on StAR synthesisEffect of CCCP on StAR synthesisEffect of CCCP on StAR synthesisEffect of CCCP on StAR synthesis
Control cAMP cccp cAMP + cccp
37kDa
30kDa

Effect of CCCP on StAR Effect of CCCP on StAR synthesissynthesis
Effect of CCCP on StAR Effect of CCCP on StAR synthesissynthesis
0
10
20
30
40
50
60
70
80C
orr
ecte
d d
ensi
ty
37 kDa30 kDa

Effect of cccp on StAR decayEffect of cccp on StAR decayEffect of cccp on StAR decayEffect of cccp on StAR decay
0
500000
1000000
1500000
2000000
2500000
3000000
3500000
0 60 120 180 240 300 360 420 480
minutes
cccp
control

Time course of StAR decayTime course of StAR decayTime course of StAR decayTime course of StAR decay

Time course of StAR decayTime course of StAR decayTime course of StAR decayTime course of StAR decay
0
20000
40000
60000
80000
100000
0 15 30 45 60 75 90 105 120
minutes
den
sity
30 kDa
32 kDa
37 kDa

0
100
200
300
400
500
600
700
800
900
con cAMP +Oligom +arsen +CCCP
ng
/ml
Effect of mitochondrial agents Effect of mitochondrial agents on progesterone productionon progesterone production
Effect of mitochondrial agents Effect of mitochondrial agents on progesterone productionon progesterone production

Effect of mitochondrial agents Effect of mitochondrial agents on StAR protein expressionon StAR protein expression
Effect of mitochondrial agents Effect of mitochondrial agents on StAR protein expressionon StAR protein expression
37 kDa
Control
cAMP+ oligomycin
+ arsenate
+ CCCP
30 kDa

3.2 kB
1.6 kB
StAR
cyclophilin
Effect of mitochondrial agents Effect of mitochondrial agents on StAR mRNA expressionon StAR mRNA expression
Effect of mitochondrial agents Effect of mitochondrial agents on StAR mRNA expressionon StAR mRNA expression
ConcAM
P+ oligm.
+ aresn.
+ CCCP

Tetramethylrhodamine Tetramethylrhodamine Ethyl Ester (TMRE)Ethyl Ester (TMRE)
Tetramethylrhodamine Tetramethylrhodamine Ethyl Ester (TMRE)Ethyl Ester (TMRE)
• Tetramethylrhodamine
Ethyl Ester (TMRE): Uptake is dependent on m. Rapidly and reversibly taken up by allowing dynamic measurement of membrane potential by fluorescent microscopy and flow cytometry.

controlcontrolcontrolcontrol CCCP-treatedCCCP-treated CCCP-treatedCCCP-treated
CCCP disruptsCCCP disrupts mm in MA10sin MA10sCCCP disruptsCCCP disrupts mm in MA10sin MA10s

• Testicular Macrophages are known to produce ROS when activated
• ROS are produced rapidly after exposure to LPS
• Many potential sources of ROS in testicular interstitium
Do reactive oxygen species Do reactive oxygen species (ROS) mediated the acute (ROS) mediated the acute inhbitory effects of LPS?inhbitory effects of LPS?
Do reactive oxygen species Do reactive oxygen species (ROS) mediated the acute (ROS) mediated the acute inhbitory effects of LPS?inhbitory effects of LPS?

ROS causes increase in 37 kDa StAR in ROS causes increase in 37 kDa StAR in Leydig cells in vitroLeydig cells in vitro
ROS causes increase in 37 kDa StAR in ROS causes increase in 37 kDa StAR in Leydig cells in vitroLeydig cells in vitro
Con cAMP +10 +25 +100 +250
cAMP + H2O2 (M)
0
20
40
60
80
100
120
ratio
{37
/30
x 10
00}
30/32 kDa
37 kDa
Con cAMP +10 +25 +100 +250
44%

Effect of HEffect of H22OO22 on StAR mRNA on StAR mRNAEffect of HEffect of H22OO22 on StAR mRNA on StAR mRNA
Northern Blot
StAR mRNA
Contr. cAMP. 100 200 250 500
Cyclophilin mRNA

TMRE staining of MA-10 cells TMRE staining of MA-10 cells exposed to H2O2exposed to H2O2
TMRE staining of MA-10 cells TMRE staining of MA-10 cells exposed to H2O2exposed to H2O2
ControlControlControlControl 100100M HM H22OO22100100M HM H22OO22

TMRE staining of MA-10 cells TMRE staining of MA-10 cells exposed to H2O2—time lapseexposed to H2O2—time lapse
TMRE staining of MA-10 cells TMRE staining of MA-10 cells exposed to H2O2—time lapseexposed to H2O2—time lapse

B
C
PFC Increases DHR-123 FluorescencePFC Increases DHR-123 FluorescencePFC Increases DHR-123 FluorescencePFC Increases DHR-123 Fluorescence
+PFC Brightfield +PFC Merge+PFC Fluorescence
Control Brightfield Control Fluorescence Control Merge
PFC: 4-phenyl-3-furoxancarbonitrile

Lipid peroxides
0
0.1
0.2
0.3
0.4
0.5
0.6
0.7
control LPS
MD
A +
HN
E (
uM
/10
e6
LC
)
LPS inhibits Leydig cells in vivo via ROSLPS inhibits Leydig cells in vivo via ROSLPS inhibits Leydig cells in vivo via ROSLPS inhibits Leydig cells in vivo via ROS
Increased lipid Increased lipid peroxidation and peroxidation and depolarization of Leydig depolarization of Leydig cell mitochondria support cell mitochondria support involvement of ROS in involvement of ROS in LPS action in vivoLPS action in vivo
Increased lipid Increased lipid peroxidation and peroxidation and depolarization of Leydig depolarization of Leydig cell mitochondria support cell mitochondria support involvement of ROS in involvement of ROS in LPS action in vivoLPS action in vivo

chol
chol
cholc ho le ste ro lp o o l
PKA
ROS
?
c ha p e ro nin
m ito c ho nd ria
c yto so l
H+
m

ROSROSROSROS
InflammationInflammationLPS, sepsisLPS, sepsis
Ischemia/Ischemia/reperfusionreperfusionIschemia/Ischemia/
reperfusionreperfusion
AgingAgingAgingAging
AlcoholAlcoholAlcoholAlcohol
XenobioticsXenobioticsPAHs, PPsPAHs, PPs
XenobioticsXenobioticsPAHs, PPsPAHs, PPs
AdenosineAdenosineAdenosineAdenosine
MitochondriaMitochondria
NucleusNucleus
ArsenateArsenateArsenateArsenate
CytokinesCytokinesCytokinesCytokines
NONO°°NONO°°
UVaUVaUVaUVa
DiabetesDiabetesDiabetesDiabetes

Steroidogenic machinerySteroidogenic machinerySteroidogenic machinerySteroidogenic machinery

Sites of immune inhibitionSites of immune inhibitionSites of immune inhibitionSites of immune inhibition
ROS

But what does it all But what does it all mean, anyway?mean, anyway?
But what does it all But what does it all mean, anyway?mean, anyway?

ConclusionsConclusionsConclusionsConclusions
• Inflammation and infection may contribute to, or cause decreased male reproductive function
• There is a push-pull system between the immune and endocrine systems– During times of sickness the immune system
suppresses the reproductive system (testosterone behavior vs. sickness behavior)
– During times of normal health testosterone suppresses the immune response

Importance of Immune-Importance of Immune-endocrine interactionsendocrine interactions
Importance of Immune-Importance of Immune-endocrine interactionsendocrine interactions
• Females are more susceptible to autoimmune diseases than males
• Estradiol and prolactin are both immuno-stimulatory
• Testosterone is immuno-inhibitory • Castration results in marked increase in thymic
cell proliferation• Higher concentration of androgen receptors in
thymus than all other tissues except prostate

Antechinus StuartiiAntechinus Stuartiivictim of his own testosteronevictim of his own testosterone
Antechinus StuartiiAntechinus Stuartiivictim of his own testosteronevictim of his own testosterone

NIH: HD25271 HD35544NIH: HD25271 HD35544NIH: HD25271 HD35544NIH: HD25271 HD35544
John Allen John Allen Paul JanusPaul JanusFred LeporeFred Lepore
Beth NardulliBeth NardulliSalil GindeSalil GindeJohn Choi John Choi Thorsten DiemerThorsten Diemer
John Allen John Allen Paul JanusPaul JanusFred LeporeFred Lepore
Beth NardulliBeth NardulliSalil GindeSalil GindeJohn Choi John Choi Thorsten DiemerThorsten Diemer
Hales LabHales LabHales LabHales Lab
Bruce Bosmann Bruce Bosmann Barbara ClarkBarbara ClarkJim FergusonJim FergusonLarry Jamison Larry Jamison Jean-Guy LeHoux Jean-Guy LeHoux Artur MayerhofferArtur MayerhofferMark McLean Mark McLean Yossi Orly Yossi Orly Anita Payne Anita Payne Richard PestellRichard PestellCatherine Rivier Catherine Rivier Focko RommertsFocko RommertsDouglas StoccoDouglas Stocco
Bruce Bosmann Bruce Bosmann Barbara ClarkBarbara ClarkJim FergusonJim FergusonLarry Jamison Larry Jamison Jean-Guy LeHoux Jean-Guy LeHoux Artur MayerhofferArtur MayerhofferMark McLean Mark McLean Yossi Orly Yossi Orly Anita Payne Anita Payne Richard PestellRichard PestellCatherine Rivier Catherine Rivier Focko RommertsFocko RommertsDouglas StoccoDouglas Stocco
collaboratorscollaboratorscollaboratorscollaborators
Karen Held HalesKaren Held HalesKaren Held HalesKaren Held Hales