Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display CHAPTER...
-
Upload
issac-platter -
Category
Documents
-
view
244 -
download
0
Transcript of Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display CHAPTER...

Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display
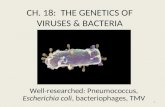
CHAPTER 14
GENE REGULATION IN BACTERIA AND BACTERIOPHAGES

Introduction
The term gene regulation means that the level of gene expression can vary under different conditions
Genes that have constant levels of expression are termed constitutive sometimes called “housekeeping genes”
The benefit of regulating genes is that encoded proteins will be produced only when required

Most regulation of gene expression is at transcriptional level rate of RNA synthesis increased or decreased
Transcriptional regulation involves actions of two types of regulatory proteins Repressors Bind to DNA & inhibit transcription Activators Bind to DNA & increase transcription
Negative control refers to transcriptional regulation by repressor proteins
Positive control to regulation by activator proteins
Transcriptional Regulation

Small effector molecules affect transcription regulation bind to regulatory proteins not to DNA directly
effector molecule may increase transcription inducers
Bind activators & cause activator to bind DNA Bind repressors & prevent repressor from binding DNA
Genes regulated this way are inducible
effector molecule may inhibit transcription Corepressors
bind repressors & cause repressor to bind DNA Inhibitors
bind activators & prevent activator from binding DNA Genes regulated this way are repressible
Transcriptional Regulation

Regulatory proteins have two binding sites
One for a small effector molecule
The other for DNA


gene regulation gods
François Jacob & André Lwoff – 1953 CSH SymposiumJacques Monod – Paris 1961

Diauxic Growth Curve Demonstrated Adaptation to Lac Metabolism

14-13Figure 14.3
Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display
The lac Operon

Regulatory Sequences of the Lac Operon

Negative - repressor protein - LacI Positive - activator protein – CAP or CRP
Induction of Lac operon requires 2 events Release of repression
lactose binds to the lac repressor causing the repressor to release operator site in DNA
Activation cAMP binds CAP protein, cAMP-CAP dimerizes & binds CAP site
in DNA
Insures that operon is on only if lactose is present glucose is low
The Lac Operon Is Regulated both Positively & Negatively

14-15Figure 14.4
Constitutive expression
RNA pol cannot initiate transcription
The lac operon is now repressed

Lac repressor protein (violet) forms a tetramer which binds to two operator sites (red) located 93 bp apart in the DNA causing a loop to form in the DNA. As a
result expression of the lac operon is turned off. This model
also shows the CAP protein (dark blue) binding to the CAP site in the promoter (dark blue
DNA). The -10 & -35 sequences of the promoter are indicated in
green.

14-16Figure 14.4
Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display
The conformation of the repressor is now altered
Repressor can no longer bind to operator
TranslationThe lac operon is now induced

The cycle of lac operon induction & repressionFigure 14.5
Repressor does not completely inhibit transcription
small amounts of the enzymes are made

1950s, Jacob & Monod, & Arthur Pardee, identified mutant bacteria with abnormal lactose adaptation
defect in lacI gene designated lacI– I = induction mutant caused constitutive expression of lac operon
(ie in absence of lactose)
The lacI– mutations mapped very close to the lac operon
The lacI Gene Encodes a Repressor Protein

Jacob, Monod & Pardee hypothesized 2 ways for lacI to function
Used genetic approach to test hypotheses
This hypothesis predicts that lacI works in trans manner
This hypothesis predicts that lacI works in a cis manner

Used F’ plasmids carrying part of lac operon Put into mutant bacteria by conjugation Bacteria that get F’ have 2 copies of lacI
gene merodipoloids
PaJaMo Experiment

2 lacI genes in a merodiploid are alleles lacI– on the chromosome lacI+ on the F’ factor
Genes on F’ plasmid are trans to bacterial chromosome If hypothesis 1 is correct
repressor produced from F’ plasmid can regulate the lac operon on the bacterial chromosome
If hypothesis 2 is correct binding site on F’ plasmid cannot affect lac operon on the
bacterial chromosome, because they are not physically adjacent
PaJaMo Experiment

14-23Figure 14.7
PaJoMo Experiment

Figure 14.7 14-24

Figure 14.7 14-25

Results
Lactose addition has no effect because operon is already on Induction is restored in merodiploid.
Now lactose addition is required to turn operon on

From Jacob & Monod, 1961, J Mol Biol 3:318
Wildtype
Induction mutants

Analysis of Lac Operon Mutants
-
F’I-O+Z+Y+
I+O+Z-Y+
lacI

From Jacob & Monod, 1961, J Mol Biol 3:318

Analysis of Lac Operon Mutants
-
-
Mutation is cis
• In merodiploid, LacZ constitutive, but LacY inducible
• OC only controls transcription of DNA on which OC is located
• O (operator) is cis-regulatory element

Interpreting the Data
The interaction between regulatory proteins & DNA sequences have led to two definitions Trans-effect & trans-acting factor
Genetic regulation that can occur even though DNA segments are not physically adjacent
Mediated by genes that encode DNA-binding regulatory proteins Example: The action of the lac repressor on the lac operon
Cis-effect & cis-acting element A DNA sequence adjacent to the gene(s) it regulates Mediated by sequences that are bound by regulatory proteins Example: The lac operator

Genetic Implications of Trans vs Cis
mutations in trans-acting factors complemented by 2nd wt gene
mutations in cis-acting elements ARE NOT complemented by 2nd wt element
Trans interactions (complementation) indicate mutation in structural gene
Cis interactions indicate mutations in regulatory sequences

From Jacob & Monod, 1961, J Mol Biol 3:318
Wildtype
Induction suppression mutant – Dominant Negative

Dominant Inhibitors or Dominant Negatives
Proteins with multiple functional domains & form multimeric complexes may be altered to prevent one function, but allow the other
When mutants retain ability to form multimeric complexes, dominant inhibition may occur

Analysis of Lac Operon Mutants
Mutation is trans
Dominant-negative
Mutation disrupts ligand binding domain of repressor

Analysis of Lac Operon Mutants
Mutation disrupts DNA binding domain of repressor

catabolite repression
When exposed to both lactose & glucose E. coli uses glucose first, & catabolite repression
prevents the use of lactose When glucose is depleted, catabolite repression is
alleviated, & the lac operon is expressed
The sequential use of two sugars by a bacterium is termed diauxic growth
lac Operon Also Regulated By Activator Protein

Effector molecule in catabolite repression cAMP (cyclic AMP)
cAMP is produced from ATP by adenylyl cyclase
cAMP binds activator protein CAP or CRP (Catabolite Activator Protein) or (cyclic AMP receptor protein)
The lac Operon Is Also Regulated By an Activator Protein

Figure 14.8
States of Lac Regulation
(b) Lactose but no cAMP

Figure 14.8
States of Lac Regulation

Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display
The trp operon (pronounced “trip”) is involved in the biosynthesis of the amino acid tryptophan
The genes trpE, trpD, trpC, trpB & trpA encode enzymes involved in tryptophan biosynthesis
The genes trpR & trpL are involved in regulation trpR Encodes the trp repressor protein
Functions in repression trpL Encodes a short peptide called the Leader peptide
Functions in attenuation
The trp Operon
14-44

Organization of the trp operon & regulation via the trp repressor protein
Figure 14.13

14-47
Organization of the trp operon & regulation via the trp repressor protein
Figure 14.13
Another mechanism of regulation
Med

14-45
Organization of the trp operon & regulation via the trp repressor protein
Figure 14.13
Cannot bind to the operator site
RNA pol can bind to the promoter

Attenuation occurs in bacteria because of the coupling of transcription & translation
During attenuation, transcription actually begins but it is terminated before the entire mRNA is made
A segment of DNA, termed the attenuator, is important in facilitating this termination
In the case of the trp operon, transcription terminates shortly past the trpL region (Figure 14.13c)
Thus attenuation inhibits the further production of tryptophan
The segment of trp operon immediately downstream from the operator site plays a critical role in attenuation
The first gene in the trp operon is trpL It encodes a short peptide termed the Leader peptide

Sequence of the trpL mRNA produced during attenuationFigure 14.14
These two codons provide a way to sense if there is sufficient
tryptophan for translation
The 3-4 stem loop is followed by a sequence
of Uracils
Region 2 is complementary to regions 1 & 3 Region 3 is complementary to regions 2 & 4
Therefore several stem-loops structures are possible
It acts as an intrinsic (-independent) terminator

Therefore, the formation of the 3-4 stem-loop causes RNA pol to terminate transcription at the end of the trpL gene
Conditions that favor the formation of the 3-4 stem-loop rely on the translation of the trpL mRNA
There are three possible scenarios 1. High levels of tryptophan 2. Medium levels of tryptophan – high trp-tRNA 3. Low levels of tryptophan – med-low trp-tRNA

Organization of the trp operon & regulation via the trp repressor protein
Figure 14.13
Repression occurs

Possible stem-loop structures formed from trpL mRNA under different conditions of translation
Figure 14.15
Sufficient amounts of tRNAtrp
Translation of the trpL mRNA progresses until stop codon
Region 2 cannot base pair with any other region
3-4 stem-loop forms
Transcription terminates
RNA polymerase pauses
Med
Attenuation occurs

Possible stem-loop structures formed from trpL mRNA under different conditions of translation
Figure 14.15
Insufficient amounts of tRNAtrp
Region 1 is blocked
3-4 stem-loop does not form
RNA pol transcribes rest of operon
Transcription occurs

The study of many operons revealed a general trend concerning inducible versus repressible regulation
Operons involved in catabolism (ie. breakdown of a substance) are typically inducible
The substance to be broken down (or a related compound) acts as the inducer
Operons involved in anabolism (ie. biosynthesis of a substance) are typically repressible
The inhibitor or corepressor is the small molecule that is the product of the operon
Inducible vs Repressible Regulation















![Forest and rangeland soil biodiversity [Chapter 5] · 2020. 9. 30. · soil viruses as bacteriophages, in part because bacteria are numerous in soils and because bacteriophages have](https://static.fdocuments.us/doc/165x107/60ea3e6fcbdf0d1cd33ab4a9/forest-and-rangeland-soil-biodiversity-chapter-5-2020-9-30-soil-viruses-as.jpg)

